Summary
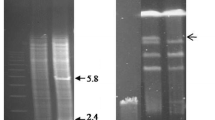
Covalently closed circular DNA was isolated from a strain of Streptomyces coelicolor ATCC 10147 and from a strain of Streptomyces coelicolor subspecies flavus ATCC 19894, using two different methods. The two plasmids were of uniform monomer size: 8.9 kb for pS 10147, the plasmid from S. coelicolor ATCC 10147, and around 125 kb for the plasmid from S. coelicolor ATCC 19894.
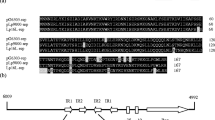
A restriction enzyme map was constructed for pS 10147, using seven enzymes. Four of the enzymes, (BamHI, Bgl,II, PvuII, and XhoI) cut pS 10147 once while PstI made two cuts. The GC content of this plasmid was calculated to be 72%. The possible utilisation of pS 10147 as a cloning vector in Streptomyces is discussed.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bendich A (1957) Methods for characterization of nucleic acids by base composition. In: Colowick SP, Kaplan NO (eds) Methods in Enzymology Vol. 3. Academic Press, New York, p 715–723
Bibb MJ, Freeman RF, Hopwood DA (1977) Physical and genetical characterization of a second sex factor SCP2 for Streptomyces coelicolor A3 (2). Mol Gen Genet 154:155–166
Bibb MJ, Schottel JL, Cohen SN (1980) A DNA cloning system for interspecies gene transfer in antibiotic-producing Streptomyces. Nature 284:526–531
Gerbaud C, Fournier P, Blanc H, Aigle M, Heslot H, Guérineau M (1979) High frequency of yeast transformation by plasmids carrying part or entire 2 μm yeast plasmid. Gene 5:233–253
Hayakawa T, Otaki N, Yonehara H, Tanaka T, Sakaguchi K (1979) Isolation and characterization of plasmids from Streptomyces. J Antibiot Tokyo 32:1348–1350
Hopwood DA (1978) Extrachromosomally determined antibiotic production. Annu Rev Microbiol 32:373–392
Inman RB, Schnös M (1970) Partial denaturation of thymine and 5-Bromouracil-containing λ DNA in alkali. J Mol Biol 49:93–98
Kutzner HJ, Waksman SA (1959) Streptomyces coelicolor Müller and Streptomyces violaceoruber Waksman and Curtis, two distinctly different organisms. J Bacteriol 78:528–538
Malik VS (1977) Preparative method for the isolation of supercoiled DNA from a chloramphenicol-producing streptomycete. J Antibiot Tokyo 30:897–899
Malik VS, Reusser F (1979) Restriction enzyme map for Streptomycete plasmid pUC3. Plasmid 2:627–631
Meselson M, Stahl FW, Vinograd J (1957) Equilibrium sedimentation of macromolecules in density gradients. Proc Natl Acad Sci USA 43:581–585
Monson AM, Bradley SG, Enquist LW, Cruces G (1969) Genetic homologies among Streptomyces violaceoruber strains. J Bacteriol 99:702–706
Nojiri C, Watabe H, Katsumata K, Yamada Y, Murakami T, Kumata Y (1980) Isolation and characterization of plasmids from parent and variant strains of Streptomyces ribosidificus. J Antibiot Tokyo 33:118–121
Okanishi M, Ohta T, Umezawa H (1970) Possible control of formation of aerial mycelium and antibiotic production in Streptomyces by episomic factors. J Antibiot Tokyo 23:45–47
Okanishi M, Suzuki K, Umezawa H (1974) Formation and reversion of Streptomycete protoplasts: cultural condition and morphological study. J Gen Microbiol 80:389–400
Okanishi M (1977) Involvement of plasmids in the production of secondary metabolites. Amino Acid Nucl Acid 35:15–30
Okanishi M (1978) Plasmids and antibiotic synthesis in Streptomyces In: Sebek OK, Laskin AI (eds). Genetics of industrial microorganisms. A.S.M. Washington, p 134–140
Philippsen P, Kramer RA, Davis RN (1978) Cloning of the yeast ribosomal DNA repeat unit in SstI and HindIII lambda vectors using genetic and physical size selections. J Mol Biol 123:371–386
Pridham TG, Anderson P, Foley C, Lindenfelser LA, Hesseltine CW, Benedict RC (1957) A selection of media for maintenance and taxonomic study of Streptomyces. Antibiotics annual 1956–1957, 947–953
Rush MG, Warner RC (1970) Alkali denaturation of covalently closed circular duplex deoxyribonucleic acid. J Biol Chem 245:2704–2708
Sanger F, Coulson AR, Friedman T, Air GM, Barrell BG, Brown NL, Fiddes JC, Hutchison III CA, Slocombe PM, Smith M (1978) The nucleotide sequence of bacteriophage ϕX 174. J Mol Biol 125:225–246
Schildkraut CL, Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from its buoyant density in CsCl. J Mol Biol 4:430–443
Schrempf H, Bujard H, Hopwood DA, Goebel W (1975) Isolation of covalently closed circular deoxyribonucleic acid from Streptomyces coelicolor A3 (2). J Bacteriol 121:416–421
Schroeder JL, Blattner FR (1978) Least-squares method for restriction mapping. Gene 4:167–174
Sutcliffe JG (1978) Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harbor Symp. Quant Biol 31:173.179
Thompson CJ, Ward JM, Hopwood DA (1980) DNA cloning in Streptomyces: resistance genes from antibiotic-producing species. Nature 286:525–527
Vinograd J, Leibowitz J, Watson R (1968) Early and late helix-coil transitions in closed circular DNA. The number of superhelical turns in Polyoma DNA. J Mol Biol 33:173–197
Wakizaka A, Kurosaka K, Okuhara E (1979) Rapid separation of DNA constituents, bases, nucleosides and nucleotides, under the same chromatographic conditions using high-performance liquid chromatography with a reversed-phase column. J Chromatogr 162:319–326
Yagisawa M, Huang TSR, Davies JE (1978) Possible involvement of plasmids in biosynthesis of neomycin. J Antibiot Tokyo 31:809–813
Zähner H, Ettlinger L (1957) Zur Systematik der Actinomyceten. 3 Die Verwertung verschiedener Kohlenstoffquellen als Hilfsmittel der Artbestimmung innerhald der Gattung Streptomyces. Arch Mikrobiol 26:307–328
Author information
Authors and Affiliations
Additional information
Communicated by W. Gajewski
Rights and permissions
About this article
Cite this article
Pernodet, JL., Guerineau, M. Isolation and physical characterization of streptomycete plasmids. Molec. Gen. Genet. 182, 53–59 (1981). https://doi.org/10.1007/BF00422766
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00422766