Abstract
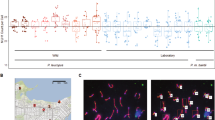
One hundred fourteen progeny from an interspecific backcross between laboratory mice and M. spretus were typed for six markers spanning most of mouse Chromosome (Chr) 16. Additional maps of 9–10 markers of this chromosome were derived from analysis of over 500 progeny from four backcrosses between inbred laboratory strains and members of the Mus musculus group, M.m. musculus and M.m. molossinus (subspecies). The results of these analyses confirmed the gene order: (CEN)-Prm-1/Prm-2-Igl-1-Smst-Mtv-6-Gap43-Pit-1(dw)-D21S16h-App-Sod-1-Ets-2-Mx. Maps produced from these five crosses were of similar lengths, but recombination in several regions was affected by sex of the F1 parent or by the combination of strains used in the cross. As reported previously, recombination frequencies were elevated significantly at the distal end of the chromosome in a cross using F1 males. The male map showed significant compression in the interval Smst to Gap43. Both male and female intersubspecific maps were expanded near the proximal and distal ends of the chromosome relative to the interspecific cross. The spretus cross was compressed in the proximal interval, Prm-1-Igl-1-Smst, and was slightly expanded in the Smst-Gap43 interval, relative to intersubspecific crosses using F1 females. Female intersubspecific maps were expanded about 50% near the distal end of the chromosome when compared to the interspecific cross. The expansion or compression of maps using different strain or sex combinations has implications for the efficient production of high resolution recombinational maps of the mouse genome.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Avner, P., Amar, L., Arnaud, D., Hanauer, A. and Cambrou, J.: Detailed ordering of markers localizing to the Xq26-Xqter region of the human X chromosome by the use of an interspecific Mus spretus mouse cross. Proc Natl Acad Sci USA 84: 1629–1633, 1987.
Bishop, D.T.: The information content of phase-known matings for ordering genetic loci. Genet Epidemiol 2: 349–361, 1985.
Callahan, R., Gallahan, D. and Kozak, C.: Two genetically transmitted BALB/c mouse mammary tumor virus genomes located on chromosomes 12 and 16. J Virol 49: 1005–1008, 1984.
Camper, S.A., Katz, R.W., Saunders, T.L. and Reeves, R.H.: The Pit-1 transcription factor gene is a candidate for the murine Snell dwarf mutation. Genomics 8: 586–590, 1990.
Chamberlain, J.S., Grant, S.G., Reeves, A.A., Mullins, L.J., Stephenson, D.A., Hoffman, E.P., Monaco, A.P., Kunkel, L.M., Caskey, C.T. and Chapman, V.M.: Regional localization of the murine Duchenne muscular dystrophy gene on the mouse X chromosome. Somat Cell Mol Genet 13: 671–678, 1987.
Cheng, S.V., Nadeau, J.H., Tanzi, R.E., Watkins, P.C., Jagadesh, J., Taylor, B.A., Haines, J.L., Sacchi, N. and Gusella, J.F.: Comparative mapping of DNA markers from the familial Alzheimer disease and Down syndrome regions of chromosome 21 to mouse chromosomes 16 and 17. Proc Natl Acad Sci USA 85: 6032–6036, 1988.
Crosby, J.L., Bleakley, R.C. and Nadeau, J.H.: A complex of serine protease genes expressed preferentially in cytotoxic T-lymphocytes is closely linked to the T-cell receptor α- and β-chain genes on mouse chromosome 14. Genomics 6: 252–259, 1990.
Donis-Keller, H., Green, P., Helms, C., Cartinhour, S., Weiffenbach, B., Stephens, K., Kieth, T.P., Bowden, B.W., Smith, D.R., Lander, E.S., Botstein, D., Akots, G., Rediker, K.S., Gravius, T., Brown, V.A., Rising, M.B., Parker, C., Powers, J.A., Watt, D.E., Kauffman, E.R., Bricker, A., Phipps, P., Muller-Kahle, H., Fulton, T.R., Ng, S., Schumm. J., Braman, J.C., Knowlton, R.G., Barker, D.E., Crooks, S.M., Lincoln, S.E., Daly, M.J. and Abrahamson, J.: A genetic linkage map of the human genome. Cell 51: 319–337, 1987.
Feinberg, A. and Vogelstein, B.: A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 137: 266–267, 1984.
Hammer, M.F., Schimenti, J. and Silver, L.M.: Evolution of mouse chromosome 17 and the origin of inversions associated with t haplotypes. Proc Natl Acad Sci USA 86: 3261–3265, 1989.
Irving, N.G., Harvey, J.A. and Brown, S.D.M.: The multipoint genetic mapping of mouse chromosome 16. Genomics, in press, 1991.
Johnson, P.A., Peschon, J.J., Yelick, P.C., Palmiter, R.D. and Hecht, N.B.: Sequence homologies in the protamine-1 and -2 genes in the mouse. Biochim Biophys Acta 950: 45–53, 1988.
Kosik, K.S., Orecchio, L.D., Bruns, G.A.P., MacDonald, G.P., Cox, D.R., and Neve, R.L.: Human Gap-43: Its deduced amino acid sequence and chromosomal localization in mouse and human. Neuron 1: 127–132, 1988.
Li, S., Crenshaw, E.B., Rawson, E.J., Simmons, D.M., Swanson, L.W., and Rosenfeld, M.G.: Dwarf locus mutants lacking three pituitary cell types result from mutations in the POU-domain gene Pit-1. Nature 347: 528–533, 1990.
Miller, S.A., Dykes, D.D. and Polesky, H.F.: A simple salting out procedure for extracting DNA from human nucleated cells. Nucl Acids Res 16: 1215, 1988.
Mock, B.A., D'Hoostelaere, L.A., Matthai, R. and Huppi, K.: A mouse homeo box gene, Hox 1.5, and the morphological locus, Hd, map to within 1 cM on chromosome 6. Genetics 116: 607–612, 1987.
O'Hara, B.F., Bendotti, C., Reeves, R.H., Oster-Granite, M.L., Coyle, J.T. and Gearhart, J.D.: Genetic mapping and analysis of somatostatin expression in Snell dwarf mice. Mol Brain Research 4: 283–292, 1988.
Reeves, R.H.: Use of Lotus 1-2-3 spreadsheet software for managing and analyzing genetic backcross data. BioTechniques 6: 12–14, 1988.
Reeves, R.H., Crowley, M.R., O'Hara, B.F. and Gearhart, J.D.: Sex, strain and species differences affect recombination across an evolutionarily conserved segment of mouse chromosome 16. Genomics 8: 141–148, 1990.
Reeves, R.H., Gallahan, D., O'Hara, B.F., Callahan, R. and Gearhart, J.D.: Genetic mapping of Prm-1, Igl-1, Smst, Mtv-6, Sod-1, and Ets-2, and localization of the Down syndrome region on mouse chromosome 16. Cytogenet Cell Genet 44: 76–81, 1987.
Reeves, R.H., Gearhart, J.D., Hecht, N.B., Yelick, P., Johnson, P. and O'Brien, S.J.: The gene encoding protamine-1 is located on human chromosome 16, and near the proximal end of mouse chromosome 16 where it is tightly linked to the gene encoding protamine-2. J Hered 80: 442–446, 1989.
Robert, R., Barton, P., Minty, A., Daubas, P., Weydert, A., Bonhomme, F., Catalan, J., Chazottes, D., Guénet, J.L. and Buckingham, M.: Investigation of genetic linkage between myosin and actin genes using an interspecific mouse backcross. Nature 314: 181–183, 1985.
Roderick, T.H. and Hillyard, A.L.: Differences in recombination due to sex in mice. Mouse News Lett 85: 87, 1989.
Scott, C.L., Mushinski, J.F., Huppi, K., Weigert, M. and Potter, M.: Amplification of Igl-1 constant genes in populations of wild mice. Nature 300: 757–760, 1982.
Seldin, M.F., Howard, T.A. and D'Eustachio, P.: Comparison of linkage maps of mouse chromosome 12 derived from laboratory strain intraspecific and Mus spretus interspecific backcrosses. Genomics 5: 24–28, 1989.
Seldin, M.F., Morse, H.C., Reeves, J.P., Scribner, C.L., LeBoeuf, R.C., and Steinberg, A.D.: Genetic analysis of “autoimmune” gld mice. I. Identification of a restriction fragment length polymorphism closely linked to the gld mutation within a conserved linkage group. J Exp Med 167: 688–693, 1988.
Seldin, M.F., Martinez, I., Howard, T.A., Naylor, S.I., and Sakaguchi, A.Y.: Localization of mouse melanoma growth stimulatory activity gene (Mgsa) between Afp and Gus on chromosome 5 using interspecific backcross mice. Cytogenet Cell Genet 54: 68–70, 1990.
Snedecore, G.W. and Cochran, W.G.: Statistical Methods, pp. 228–257, Iowa State University Press, Ames IA, 1967.
Staehli, P., Pravtcheva, D., Lundin, L.G., Acklin, M., Ruddle, F., Lindenmann, J., and Haller, O.: Interferon regulated influenza virus resistance gene Mx is localized on mouse chromosome 16. J Virol 58: 967–969, 1986.
Stewart, G.D., Harris, P., Galt, J., and Ferguson-Smith, M.A.: Cloned DNA probes originally mapped to human chromosome 21 and their use in determining the origin of nondisjunction. Nucl Acids Res 13: 4125–4132, 1985.
Watson, D.K., Kozak, C., Smith, M.J., Reeves, R.H., Gearhart, J.D., Nunn, M.F., Nash, W., Fowle, J.R., Duesberg, P., Papas, T.S., and O'Brien, S.J.: Conserved chromosomal positions of dual domains of the ets protooncogene in cats, mice, and humans. Proc Natl Acad Sci USA 83: 1792–1796, 1986.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Reeves, R.H., Crowley, M.R., Moseley, W.S. et al. Comparison of interspecific to intersubspecific backcrosses demonstrates species and sex differences in recombination frequency on mouse Chromosome 16. Mammalian Genome 1, 158–164 (1991). https://doi.org/10.1007/BF00351062
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00351062