Summary
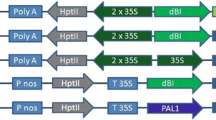
We have previously described substantial variation in the level of expression of two linked genes which were introduced into transgenic petunia plants using Agrobacterium tumefaciens. These genes were (i) nopaline synthase (nos) and (ii) a chimeric chlorophyll a/b binding protein/octopine synthase (cab/ocs) gene. In this report we analyze the relationship between the level of expression of the introduced genes and T-DNA structure and copy number in 40 transgenic petunia plants derived from 26 transformed calli. Multiple shoots were regenerated from 8 of these calli and in only 6 cases were multiple regenerated shoots from each callus genotypically identical to each other. Many genotypes showed no nos gene expression (22/28). Most of the plants (16/22) which lacked nos gene expression did contain nos-encoding DNA with the expected restriction enzyme map. Similarly, amongst the genotypes showing no cab/ocs gene expression, the majority (11/28) did not show any alterations in restriction fragments corresponding to the expected cab/ocs coding sequences (10/11). Approximately half of the plants carried multiple copies of T-DNA in inverted repeats about the left or right T-DNA boundaries. No positive correlation was observed between the copy number of the introduced DNA and the level of expression of the introduced genes. However, plants with high copy number complex insertions composed of multiple inverted repeats in linear arrays usually showed low levels of expression of the introduced genes.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Beachy RN, Chen Z-L, Horsch RB, Rogers SG, Hoffmann NJ, Fraley RT (1985) Accumulation and assembly of soybean betaconglycinin in seeds of transformed petunia plants. EMBO J 4:3047–3054
Depicker A, Stachel S, Dhaese P, Zambryski P, Goodman H (1982) Nopaline synthase: transcript mapping and DNA sequence. J Mol Appl Genet 1:561–573
Jones JDG, Dunsmuir P, Bedbrook JR (1985) High level expression of introduced chimaeric genes in regenerated transformed plants. EMBO J 4:2411–2418
Joos H, Inze D, Caplan A, Sormann M, Van Montagu M, Schell J (1983) Genetic analysis of T-DNA transcripts in nopaline crown galls. Cell 32:1057–1067
Jorgensen R, Snyder C, Jones JDG (1987) T-DNA is organised predominately in inverted repeat structures in plants transformed with Agrobacterium tumefaciens C58 derivatives. Mol Gen Genet 207:471–477
Kloppsteck K (1985) Diurnal and circadian rhythmicity in the expression of light-induced plant nuclear messenger RNAs. Planta 165:502–506
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbour Laboratory, Cold Spring Harbour, New York
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nagy F, Morelli G, Fraley RT, Rogers SG, Chua N-H (1985) Photoregulated expression of a pea rbcS gene in leaves of transgenic plants. EMBO J 4:3063–3068
Southern EM (1975) Detection of specific sequences among DNA fragments separated by electrophoresis. J Mol Biol 98:503–517
Walling L, Drews GN, Goldberg RB (1986) Transcriptional and post-transcriptional regulation of soybean seed protein mRNA levels. Proc Natl Acad Sci USA 83:2123–2127
Zambryski P, Joos H, Genetello C, Leemans J, Van Montagu M, Schell J (1983) Ti plasmid vector for the introduction of DNA into plant cells without alteration of their normal regeneration capacity. EMBO J 2:2143–2150
Author information
Authors and Affiliations
Additional information
Communicated by R.B. Goldberg
Rights and permissions
About this article
Cite this article
Jones, J.D.G., Gilbert, D.E., Grady, K.L. et al. T-DNA structure and gene expression in petunia plants transformed by Agrobacterium tumefaciens C58 derivatives. Mol Gen Genet 207, 478–485 (1987). https://doi.org/10.1007/BF00331618
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00331618