Abstract
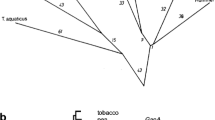
We have cloned and sequenced the single-copy nuclear gene (GapC) encoding the complete 335-amino acid cytosolic glyceraldehyde-3-phosphate dehydrogenase (GAPC) from the red alga Gracilaria verrucosa. The proline residue which contributes to the specificity of NAD+ binding in other GAPC-like proteins is present. Putative regulatory regions, including GC-rich regions, a GATA element, and 11-base T- and T/G-clusters, but excluding TATA- and CCAAT-boxes, were identified upstream. Two types of GapC cDNAs differing in polyadenylation site were characterized. An 80-bp phase-two spliceosomal intron was identified in a novel position interrupting the highly conserved cofactor-coding region I. The G. verrucosa GAPC was easily aligned with other known GAPC-type sequences. Inferred phylogenetic trees place red algae among the eukaryote crown taxa, although with modest bootstrap support and without stable resolution among related GAPC lineages.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bachmann B, Lüke W, Hunsmann G (1990) Improvement of PCR-amplified DNA sequencing with the aid of detergents. Nucleic Acids Res 18:1309
Bestor TH (1990) DNA methylation: evolution of a bacterial immune function into a regulator of gene expression and genome structure in higher eukaryotes. Phil Trans R Soc Lond B 326:179–188
Bhattacharya D, Elwood HJ, Goff LJ, Sogin ML (1990) Phylogeny of Gracilaria lemaneiformis (Rhodophyta) based on sequence analysis of its small-subunit ribosomal RNA coding region. J Phycol 26:181–186
Bird A (1992) The essentials of DNA methylation. Cell 70:5–8
Bird CJ, Rice EL, Murphy CA, Ragan MA (1992) Phylogenetic relationships in the Gracilariales (Rhodophyta) as determined by 18 rDNA sequences. Phycologia 31:510–522
Boyen C, Leblanc C, Bonnard G, Grienenberger J-M, Kloareg B (1994) Nucleotide sequence of the cox3 gene from Chondrus crispus: evidence that UGA encodes tryptophan and evolutionary implications. Nucleic Acids Res 22:1400–1403
Brinkmann H, Martinez P, Quigley F, Martin W, Cerff R (1987) Endosymbiotic origin and codon bias of the nuclear gene for chloroplast glyceraldehyde-3-phosphate dehydrogenase from maize. J Mol Evol 26:320–328
Brinkmann H, Cerff R, Salomon M, Soll J (1989) Cloning and sequence analysis of cDNAs encoding the cytosolic precursors of subunits GapA and GapB of chloroplast glyceraldehyde-3-phosphate dehydrogenase from pea and spinach. Plant Mol Biol 13:81–94
Cavalier-Smith T (1985) Selfish DNA and the origin of introns. Nature 315:283–284
Cavalier-Smith T, Allsopp MTEP, Chao EE (1994) Chimeric conundra: are nucleomorphs and chromists monophyletic or polyphyletic? Proc Natl Acad Sci USA 91:11368–11372
Cerff R, Bohnert HJ, Ragan M, Sachs MM (1994) Plant-wide nomenclature of nuclear genes encoding cytosolic and chloroplast glyceraldehyde-3-phosphate dehydrogenases. Pl Mol Biol Rep 12:S36-S37
Corbier C, Clermont S, Billard P, Skarzunski T, Branlant C, Wonacott A, Branlant G (1990) Probing the coenzyme specificity of glyceraldehyde-3-phosphate dehydrogenases by site-directed mutagenesis. Biochemistry 29:7101–7106
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16:10881–10890
Craik CS, Rutter WJ, Fletterick R (1983) Splice junctions:association with variation in protein structure. Science 220:1125–1129
Darnell JE (1978) Implications of RNA·RNA splicing in evolution of eukaryotic cells. Science 202:1257–1260.
Doolittle RF, Feng DF, Anderson KL, Alberro MR (1990) A naturally occurring horizontal gene transfer from a eukaryote to a prokaryote. J Mol Evol 31:383–388
Doolittle WF (1978) Genes in pieces: were they ever together? Nature 272:581–582
Douglas SE, Murphy CA, Spencer DF, Gray MW (1991) Cryptomonad algae are evolutionary chimaeras of two phylogenetically-distinct unicellular eukaryotes. Nature 350:148–151
Emanuel JR (1991) Simple and efficient system for synthesis of non-radioactive nucleic acid-hybridization probes using PCR. Nucleic Acids Res 19:2790
Ercolani L, Florence B, Denaro M, Alexander M (1988) Isolation and complete sequence of a functional human glyceraldehyde-3-phosphate dehydrogenase gene. J Biol Chem 263:15335–15341
Feisentein J (1989) PHYLIP — phylogeny inference package (version 3.2). Cladistics 5:164–166
Fothergill-Gilmore LA, Michels PAM (1993) Evolution of glycolysis. Prog Biophys Mol Biol 59:105–235
Gilbert W (1978) Why genes in pieces? Nature 271:501
Hardie DG, Coggins JR (1986) Multidomain proteins — structure and evolution. Elsevier Science Publishers (Biomedical Division), Amsterdam
Harmsen MC, Schuren FHJ, Moukhaa SM, van Zuilen CM, Punt PJ, Wessels JGH (1992) Sequence analysis of the glyceraldehyde-3-phosphate dehydrogenase genes from the basidiomycetes Schizophyllum commune, Phanerochaete chrysosporium and Agaricus bisporus. Curr Genet 22:447–454
Harris JI, Waters M (1976) Glyceraldehyde-3-phosphate dehydrogenase. In: Boyer PD (ed) The enzymes, vol 13. Academic Press, New York, pp 1–14
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Systematic Biol 42:182–192
Hori H, Osawa S (1987) Origin and evolution of organisms as deduced from 5S ribosomal RNA sequences. Mol Biol Evol 4:445–472
Iwabe N, Kuma KI, Hasami M, Osawa S, Miyata T (1989) Evolutionary relationship of archaebacteria, eubacteria, and eukaryotes inferred from phylogenetic trees of duplicated genes. Proc Natl Acad Sci USA 86:9355–9359
Kersanach R, Brinkmann H, Liaud M-F, Zhang DX, Martin W, Cerff R (1994) Five identical intron positions in ancient duplicated genes of eubacterial origin. Nature 367:387–389
Lam E, Chua NH (1989) ASF-2: a factor that binds to the cauliflower mosaic virus 35S promoter and a conserved GATA motif in Cab promoters. Plant Cell 1:1147–1156
Lawrence JG, Hartl DL, Ochman H (1991) Molecular considerations in the evolution of bacterial genes. J Mol Evol 33:241–250
Liaud M-F, Valentin C, Martin W, Bouget F-Y, Kloareg B, Cerff R (1994) The evolutionary origin of red algae as deduced from the nuclear genes encoding cytosolic chloroplast glyceraldehyde-3-phosphate dehydrogenases from Chondrus crispus. J Mol Evol 38:319–327
Logsdon Jr JM, Palmer JD (1994) Origin of introns — early or late? Nature 369:526
Markos A, Miretsky A, Müller M (1993) A glyceraldehyde-3-phosphate dehydrogenase with eubacterial features in the amitochondriate eukaryote, Trichomonas vaginalis. J Mol Evol 37:631–643
Martin W, Cerff R (1986) Prokaryotic features of a nucleus-encoded enzyme. cDNA sequence for chloroplast and cytosolic glyceraldehyde-3-phosphate dehydrogenase from mustard (Sinapis alba). Eur J Biochem 159:323–331
Martin W, Brinkmann H, Savonna C, Cerff R (1993) Evidence for a chimeric nature of nuclear genomes: eubacterial origin of eukaryotic glyceraldehyde-3-phosphate dehydrogenase genes. Proc Natl Acad Sci USA 90:8692–8696
Martinez P, Martin W, Cerff R (1989) Structure, evolution and anaerobic regulation of a nuclear gene encoding cytosolic glyceraldehyde-3-phosphate dehydrogenase from maize. J Mol Biol 208:551–565
Michels PAM, Marchand M, Kohl L, Allert S, Wierenga RK, Opperdoes FR (1991) The cytosolic and glycosomal isoenzymes of glyceraldehyde-3-phosphate dehydrogenase in Tripanosoma brucei have a distant evolutionary relationship. Eur J Biochem 198:421–428
Osiewacz HD, Ridder R (1991) Genome analysis of imperfect fungi: electrophoretic karyotyping and characterization of the nuclear gene coding for glyceraldehyde-3-phosphate dehydrogenase (gpd) of Curvularia lunata. Curr Genet 20:151–155
Patthy L (1987) Intron-dependent evolution: preferred types of exons and introns. FEBS Lett 214:1–7
Pearson WR, Lipman DJ (1988) Improved tools for biological sequence comparison. Proc Natl Acad Sci USA 85:2444–2448
Perasso R, Baroin A, Qu LH, Bachellerie JP, Adoutte A (1989) Origin of the algae. Nature 339:142–144
Punt PJ, Dingemanse MA, Jacobs-Meijsing BJ, Pouwels PH, van den Hondel CA (1988) Isolation and characterization of the glyceraldehyde-3-phosphate dehydrogenase gene of Aspergillus nidulans. Gene 69:49–57
Ragan MA, Gutell RR (1995) Are red algae plants? Bot J Linn Soc (in press)
Ragan MA, Bird CJ, Rice EL, Gutell RR, Murphy CA, Singh RK (1994) A molecular phylogeny of the marine red algae (Rhodophta) based on the nuclear small-subunit rRNA gene. Proc Natl Acad Sci USA 91:7276–7280
Risler JL, Delacroix H, Henaut A (1988) Amino-acid substitution in structurally related proteins (a pattern recognition approach): determination of a new and efficient scoring matrix. J Mol Biol 204:1019–10219
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Schindler U, Cashmore AR (1990) Photoregulated gene expression may involve ubiquitous DNA-binding proteins. EMBO J 9:3415–3427
Shih M-C, Lazar G, Goodman HM (1986) Evidence in favor of the symbiotic origin of chloroplasts: primary structure and evolution of tobacco glyceraldehyde-3-phosphate dehydrogenase. Cell 47:73–80
Shih M-C, Heinrich P, Goodman HC (1991) Cloning and chromosomal mapping of nuclear genes encoding chloroplast and cytosoloic glyceraldehyde-3-phosphate dehydrogenase from Arabidopsis thaliana. Gene 104:133–138
Smith TL (1989) Disparate evolution of yeasts and filamentous fungi indicated by phylogenetic analysis of glyceraldehyde-3-phosphate dehydrogenase genes. Proc Natl Acad Sci USA 86:7063–7066
Smith TL, Leong SA (1990) Isolation and characterization of a Ustilago maydis glyceraldehyde-3-phosphate dehydrogenase-encoding gene. Gene 93:111–117
Stone EM, Rothblum KN, Schwartz RJ (1985) Intron-dependent evolution of the chicken glyceraldehyde phosphate dehydrogenase gene. Nature 313:498–500
Swofford DL (1993) PAUP: phylogenetic analysis using parsimony, Version 3.1. (Computer program) The Illinois National History Survey, Champaign, Illinois
Van Wert SL, Yoder OC (1992) Structure of the Cochliobolus heterostrophus glyceraldehyde-3-phosphate dehydrogenase. Curr Genet 22:29–35
Zhou Y-H, Ragan MA (1993) cDNA cloning and characterization of the nuclear gene encoding chloroplast glyceraldehyde-3-phosphate dehydrogenase from the marine red alga Gracilaria verrucosa. Curr Genet 23:483–489
Zhou Y-H, Ragan MA (1994) Cloning and characterization of the nuclear gene encoding plastid glyceraldehyde-3-phosphate dehydrogenase from the marine red alga Gracilaria verrucosa. Curr Genet 26:79–86
Zhou Y-H, Ragan MA (1995) Nuclear-encoded protein-coding genes of the agarophyte Gracilaria verrucosa (Hudson) Papenfuss. Proc Intl Seaweed Symp 15 (in press)
Zhu T, Schupp JM, Oliphant A, Keim P (1994) Hypomethylated sequences: characterization of the duplicate soyabean genome. Mol Gen Genet 224:638–645
Author information
Authors and Affiliations
Additional information
Communicated by R.W. Lee
GSDB accession L38661 Issued as NRCC no. 38071
Present address Human Genetics Center, University of Texas, P.O. Box 20334, Houston, TX 77225, USA
Rights and permissions
About this article
Cite this article
Zhou, YH., Ragan, M.A. The nuclear gene and cDNAs encoding cytosolic glyceraldehyde-3-phosphate dehydrogenase from the marine red alga Gracilaria verrucosa: cloning, characterization and phylogenetic analysis. Curr Genet 28, 324–332 (1995). https://doi.org/10.1007/BF00326430
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00326430