Summary
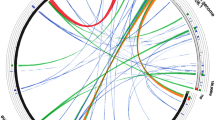
Physical maps of liverwort Marchantia polymorpha chloroplast DNA were constructed with restriction endonucleases, BamHI, SmaI, KpnI, and XhoI. The molecular size, calculated from the sum of the restriction fragments, is 121.0±1.0 kilobase (kb) pairs. This value is in good agreement with that of 118.7±2.0 kb pairs obtained from contour length measurements of the chloroplast DNA electron micrographs. The physical map indicates that the chloroplast DNA contains two copies of inverted repeats. The length of the inverted repeat is at most 11.7 kb pairs. Southern hybridization analysis indicates that two sets of the chloroplast ribosomal RNA genes are located in the inverted repeat regions. The site of the ribulose bisphosphate (RuBP) carboxylase (large subunit) gene was also determined on a BamHI fragment, Ba5, by using 32P-labeled DNA fragments containing the tobacco chloroplast RuBP carboxylase gene as a probe.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Atchison BA, Whitfeld PR, Bottomley W (1976) Comparison of chloroplast DNAs by specific fragmentation with Eco RI endonuclease. Mol Gen Genet 148:263–269
Belliard G, Pelletier G, Vedel F, Quetier F (1978) Morphological characteristics and chloroplast DNA distribution in different cytoplasmic parasexual hybrids of Nicotiana tabacum. Mol Gen Genet 165:231–237
Bedbrook JR, Bogorad L (1976) Endonuclease recognition sites mapped on Zea mays chloroplast DNA. Proc Natl Acad Sci USA 73:4309–4313
Bedbrook JR, Kolodner R, Bogorad L (1977) Zea mays chloroplast ribosomal RNA genes are part of a 22,000 base pair inverted repeat. Cell 11:739–749
Bedbrook JR, Link G, Coen DM, Bogorad L, Rich A (1978) Maize plastid gene expressed during photoregulated development. Proc Natl Acad Sci USA 75:3060–3064
Bogorad L (1979) The chloroplast, its genome and possibilities for genetically manipulating plants. In: Setlow JK, Hollaender A (eds) Genetic engineering, vol I. Plenum Press, New York, p 181
Bovenberg WA, Kool AJ, Nijkamp HJJ (1981) Isolation, characterization and restriction endonuclease mapping of the Petunia hybrida chloroplast DNA. Nucl Acids Res 9:503–517
Chu NM, Oishi KK, Tewari KK (1981) Physical mapping of the pea chloroplast DNA and localization of the ribosomal RNA genes. Plasmid 6:279–292
Coen DM, Bedbrook JR, Bogorad L, Rich A (1977) Maize chloroplast DNA fragment encoding the large subunit of ribulosebis-phosphate carboxylase. Proc Natl Acad Sci USA 74:5487–5491
Denhardt DT (1966) A membrane-filter technique for the detection of complementary DNA. Biochem Biophys Res Commun 23:641–646
Driesel AJ, Crouse EJ, Gordon K, Bohnert HJ, Herrmann RG (1979) Fractionation and identification of spinach chloroplast transfer RNAs and mapping of their genes on the restriction map of chloroplast DNA. Gene 6:285–306
Driesel AJ, Speirs J, Bohnert HJ (1980) Spinach chloroplast mRNA for a 32,000 dalton polypeptide. Size and localization on the physical map of the chloroplast DNA. Biochim Biophys Acta 610:297–310
Van Ee JH, Vos YJ, Planta RJ (1980) Physical map of chloroplast DNA of Spirodela oligorrbiza; analysis by the restriction endonucleases PstI, XhoI and SacI. Gene 12:191–200
Gelvin S, Heizmann P, Howell SH (1977) Identification and cloning of the chloroplast gene coding for the large subunit of ribulose-1,5-bisphosphate carboxylase from Chlamydomonas reinhardii. Proc Natl Acad Sci USA 74:3193–3197
Gordon KHJ, Crouse EJ, Bohnert HJ, Herrmann RG (1981) Restriction endonuclease cleavage site map of chloroplast DNA from Oenothera parviflora (Euoenothera plastome IV). Theor Appl Genet 59:281–296
Gordon KHJ, Crouse EJ, Bohnert HJ, Herrmann RG (1982) Physical mapping of differences in chloroplast DNA of the five wild-type plastomes in Oenothera subsection Euoenothera. Theor Appl Genet 61:373–384
Gray PW, Hallick RB (1977) Restriction endonuclease map of Euglena gracilis chloroplast DNA. Biochemistry 16:1665–1671
Gray PW, Hallick RB (1978) Physical mapping of the Euglena gracilis chloroplast DNA and ribosomal RNA gene region. Biochemistry 17:284–289
Herrmann RG, Whitfeld PR, Bottomley W (1980a) Construction of a Sall/PstI restriction map of spinach chloroplast DNA using low-gelling-temperature-agarose electrophoresis. Gene 8:179–191
Herrmann RG, Palta HK, Kowallik KV (1980b) Chloroplast DNA from three Archegoniates. Planta 148:319–327
Jurgenson JE, Bourque DP (1980) Mapping of rRNA genes in and inverted repeat in Nicotiana tabacum chloroplast DNA. Nucl Acids Res 8:3505–3516
Koller B, Delius H (1980) Vicia faba chloroplast DNA has only one set of ribosomal RNA genes as shown by partial denaturation mapping and R-loop analysis. Mol Gen Genet 178:261–269
Link G, Bogorad L (1980) Sizes, locations, and directions of transcription of two genes on a cloned maize chloroplast DNA sequence. Proc Natl Acad Sci USA 77:1832–1836
Malnoe P, Rochaix J-D, Chua NH, Spahr P-F (1979) Characterization of the gene and messenger RNA of the large subunit of ribulose 1,5-diphosphate carboxylase in Chlamydomonas reinhardii. J Mol Biol 133:417–434
Manning JE, Richards OC (1972) Isolation and molecular weight of circular chloroplast DNA from Euglena gracilis. Biochim Biophys Acta 259:285–296
Ohyama K, Wetter LR, Yamano Y, Fukuzawa H, Komano T (1982) A simple method for isolation of chloroplast DNA from Marchantia polymorpha L. cell suspension cultures. Agric Biol Chem 46:237–242
Ono K, Ohyama K, Gamborg OL (1979) Regeneration of the liverwort Marchantia polymorpha L. from protoplasts isolated from cell suspension culture. Plant Sci Lett 14:225–229
Palmer JD, Thompson WF (1981) Rearrangements in the chloroplast genomes of mung bean and pea. Proc Natl Acad Sci USA 78:5533–5537
Rawson JRY, Kushner SR, Vapnek D, Alton NK, Boerma CL (1978) Chloroplast ribosomal RNA genes in Euglena gracilis exist as three clustered tandem repeats. Gene 3:191–209
Rawson JRY, Clegg MT, Thomas K, Rinehart C, Wood B (1981) A restriction map of the ribosomal RNA genes and the short single-copy DNA sequence of the pearl millet chloroplast genome. Gene 16:11–19
Rigby PWJ, Dieckmann M, Rhodes C, Berg P (1977) Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol 113:237–251
Rochaix JD (1978) Restriction endonuclease map of the chloroplast DNA of Chlamydomonas reinhardii. J Mol Biol 126:597–617
Rochaix JD, Malnoe P (1978) Anatomy of the chloroplast ribosomal DNA of Chlamydomonas reinhardii. Cell 15:661–670
Seyer P, Kowallik KV, Herrmann RG (1981) A physical map of Nicotiana tabacum plastid DNA including the location of structural genes for ribosomal RNAs and the large subunit of ribulose bisphosphate carboxylase/oxygenase. Current Genetics 3:189–204
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
Sugiura M, Kusuda J (1979) Molecular cloning of tobacco chloroplast ribosomal RNA genes. Mol Gen Genet 172:137–141
Vedel F, Quetier F, Bayen M (1976) Specific cleavage of chloroplast DNA from higher plants by Eco RI restriction nuclease. Nature 263:440–442
Vedel F, Quetier F, Dosba F, Doussinault G (1978) Study of wheat phylogeny by Eco RI analysis of chloroplastic and mitochondrial DNAs. Plant Sci Lett 13:97–102
Westhoff P, Nelson N, Bünemann H, Herrmann RG (1981) Localization of genes for coupling factor subunits on the spinach plastic chromosome. Current Genetics 4:109–120
Whitfeld PR, Herrmann RG, Bottomley W (1978) Mapping of the ribosomal RNA genes on spinach chloroplast DNA. Nucl Acids Res 5:1741–1751
Whitfeld PR, Bottomley W (1980) Mapping of the gene for the large subunit of ribulose bisphosphate carboxylase on spinach chloroplast DNA. Biochem International 1:172–178
Yamagishi H, Inokuchi H, Ozeki H (1976) Excision and duplication of su3+-transducing fragments carried by bacteriophage ϕ80. I. Novel structure of ϕ80sus2psu3+ DNA molecule. J Virol 18:1016–1023
Zurawski G, Perrot B, Bottomley W, Whitfeld PR (1981) The structure of the gene for the large subunit of ribulose 1,5-bisphosphate carboxylase from spinach chloroplast DNA. Nucl Acids Res 9:3251–3270
Author information
Authors and Affiliations
Additional information
Communicated by J. Schell
Rights and permissions
About this article
Cite this article
Ohyama, K., Yamano, Y., Fukuzawa, H. et al. Physical mappings of chloroplast DNA from liverwort Marchantia polymorpha L. cell suspension cultures. Molec Gen Genet 189, 1–9 (1983). https://doi.org/10.1007/BF00326047
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00326047