Abstract
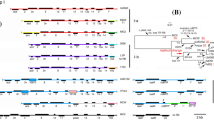
The octopine/cucumopine (o/c) Ti plasmids of the grapevine-associated Agrobacterium vitis strains constitute a family of related DNA molecules. Restriction maps were established of two limited-host-range o/c Ti plasmids, pTiAg57 and pTiAB3, and of the wide-host-range o/c Ti plasmid pTiHml. Together with the previously obtained map of the wide-host-range o/c Ti plasmid pTiTm4, about 1000 kb were mapped with a resolution of 0.2 kb, allowing a detailed comparison of the various structures. One region of the o/c Ti plasmids is highly conserved and differs mainly by the presence or absence of relatively small DNA fragments (0.9–2.7 kb); the other region has been modified more extensively and carries large sequences specific for each Ti plasmid type. The sequence similarity within large conserved regions shows that these plasmids have diverged recently and that their evolution was driven by large-scale genetic events rather than single nucleotide changes. These results have important implications for studies on bacterial evolution.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Arber W (1991) Elements in microbial evolution. J Mol Evol 33:4–12
Bonnard G, Vincent F, Otten L (1989) Sequence and distribution of IS866, a novel T-region associated insertion sequence from Agrobacterium tumefaciens. Plasmid 22:70–81
Buchholz WG, Thomashow MF (1984) Host range encoded by the Agrobacterium tumefaciens tumor-inducing plasmid pTiAg63 can be expanded by modification of its T-DNA oncogene complement. J Bacteriol 160:327–332
Clark-Curtiss JE, Walsh GP (1989) Conservation of genomic sequences among isolates of Mycobacterium leprae. J Bacteriol 171:4844–4851
Cook D, Barlow E, Sequeira L (1989) Genetic diversity of Pseudomonas solanacearum: detection of restriction fragment length polymorphisms with DNA probes that specify virulence and the hypersensitive response. Mol Plant-Microbe Interact 2:113–121
Currier TC, Nester EW (1976) Isolation of covalently closed circular DNA of high molecular weight from bacteria. Anal Biochem 76:431–441
Fournier P, Paulus F, Otten L (1993) IS870 requires a 5′-CTAG-3′ target sequence to generate the stop codon for its large ORF1. J Bacteriol, in press
Gabriel DW, Hunter JE, Kingsley MT, Moller JW, Lazo GR (1988) Clonal population structure of Xanthomonas campestris and genetic diversity among citrus canker strains. Mol PlantMicrobe Interact 1:59–65
Gough J, Murray N (1983) Sequency diversity among related genes for recognition of specific targets in DNA molecules. J Mol Biol 166:1–19
Hoekema A, de Pater BS, Fellinger AJ, Hooykaas PJJ, Schilperoort RA (1984) The limited host range of an Agrobacterium tumefaciens strain extended by a cytokinin gene from a wide host range T-region. EMBO J 3:3043–3047
Huss B, Bonnard G, Otten L (1989) Isolation and functional analysis of a set of auxin genes with low root-inducing activity from an Agrobacterium tumefaciens biotype III strain. Plant Mol Biol 12:271–283
Kado CI (1991) Molecular mechanisms of crown gall tumorigenesis. Crit Rev Plant Sci 10:1–32
Kerr A, Panagopoulos CG (1977) Biotypes of Agrobacterium radiobacter var. tumefaciens and their biological control. Phytopath Z 90:172–179
Knauf VC, Panagopoulos CG, Nester EW (1982) Genetic factors controlling host range of Agrobacterium tumefaciens. Phytopathology 12:1545–1549
Knauf VC, Panagopoulos CG, Nester EW (1983) Comparison of Ti plasmids from three different biotypes of Agrobacterium tumefaciens isolated from grapevines. J Bacteriol 153:1535–1542
Knauf VC, Yanofsky M, Montoya A, Nester EW (1984) Physical and functional map of an Agrobacterium tumefaciens plasmid that confers a narrow host range. J Bacteriol 160:564–568
McFadden JJ, Butcher PD, Chiodini R, Hermon-Taylor J (1987) Crohn's disease-isolated mycobacteria are identical to Mycobacterium paratuberculosis, as determined by DNA probes that distinguish between mycobacterial species. J Clin Microbiol 25:796–801
Nei M, Li W-H (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Ophel K, Kerr A (1990) Agrobacterium vitis sp. nov. for strains of Agrobacterium biovar 3 from grapevines. Int J Syst Bacteriol 40:236–241
Otten L, Canaday J, Gérard J-C, Fournier P, Crouzet P, Paulus F (1992a) Evolution of Agrobacteria and their Ti plasmids. A review. Mol Plant-Microbe Interact 5:279–287
Otten L, Gérard J-C, de Ruffray P (1992b) The Ti plasmid from the wide host range Agrobacterium vitis strain Tm4; map and homology with other Ti plasmids. Plasmid 29:154–159
Panagopoulos CG, Psallidas PG (1973) Characteristics of Greek isolates of Agrobacterium tumefaciens (EF Smith and Townsend) Conn. J Appl Bacteriol 36:233–240
Paulus F, Huss B, Bonnard G, Ride M, Szegedi E, Tempé J, Petit A, Otten, L (1989a) Molecular systematics of biotype III Ti plasmids of Agrobacterium tumefaciens. Mol Plant-Microbe Interact 2:64–74
Paulus F, Ridé M, Otten L (1989b) Distribution of two Agrobacterium tumefaciens insertion elements in natural isolates: evidence for stable association between Ti plasmids and their bacterial hosts. Mol Gen Genet 219:145–152
Paulus F, Huss B, Tinland B, Herrmann A, Canaday J, Otten L (1991a) Role of T-region borders in Agrobacterium host range. Mol Plant-Microbe Interact 4:163–172
Paulus F, Canaday J, Otten, I (1991b) Limited host range Ti plasmids: recent origin from wide host range Ti plasmids and involvement of a novel IS element, IS868. Mol Plant-Microbe Interact 4:190–197
Paulus F, Canaday J, Vincent F, Bonnard G, Kares C, Otten L (1991c) Sequence of the iaa and ipt region of different Agrobacterium tumefaciens biotype III octopine strains: reconstruction of octopine Ti plasmid evolution. Plant Mol Biol 16:601–614
Regnery RL, Spruill CL, Plikaytis BD (1991) Genotypic diversification of Rickettsiae and estimation of intraspecies sequence divergence for portions of two rickettsial genes. J Bacteriol 173:1576–1589
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edition. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Szegedi E (1985) Host range and specific l(+)-tartrate utilization of biotype 3 Agrobacterium tumefaciens. Acta Phytopathol Acad Sci Hung 20:17–22
Thomashow MF, Knauf VC, Nester EW (1981) Relationship between the limited and wide host range octopine-type Ti plasmids of Agrobacterium tumefaciens. J Bacteriol 146:484–493
Upholt WB (1977) Estimation of DNA sequence divergence from comparison of restriction endonuclease digests. Nucleic Acids Res 4:1257–1265
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Yanofsky M, Montoya A, Knauf VC, Lowe B, Gordon M, Nester, EW (1985a) Limited-host-range plasmids of Agrobacterium tumefaciens: molecular and genetic analysis of transferred DNA. J Bacteriol 163:341–348
Yanofsky M, Lowe B, Montoya A, Rubin R, Krul W, Gordon MP, Nester EW (1985b) Molecular and genetic analysis of factors controlling host range in Agrobacterium tumefaciens. Mol Gen Genet 201:237–246
Author information
Authors and Affiliations
Additional information
Communicated by A. Kondorosi
Rights and permissions
About this article
Cite this article
van Nuenen, M., de Ruffray, P. & Otten, L. Rapid divergence of Agrobacterium vitis octopine-cucumopine Ti plasmids from a recent common ancestor. Molec. Gen. Genet. 240, 49–57 (1993). https://doi.org/10.1007/BF00276883
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00276883