Abstract
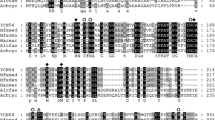
The nitrite oxidoreductase (NOR) from the facultative nitrite-oxidizing bacterium Nitrobacter hamburgensis X14 was investigated genetically. In order to develop a probe for the gene norB, the N-terminal amino acid sequence of the NOR β-subunit (NorB) was determined. Based on that amino acid sequence, an oligo-nucleotide was derived that was used for the identification and cloning of gene norB. Sequence analysis of DNA fragments revealed three adjacent open reading frames in the order norA, norX, norB. The DNA sequences of norX and norB represented complete genes while the open reading frame of norA was truncated by the cloning site. The deduced amino acid sequence of protein NorB contained four cysteine clusters with striking homology to those of iron-sulfur centers of bacterial ferredoxins. NorB shares significant sequence similarity to the β-subunits (NarH, NarY) of the two dissimilatory nitrate reductases (NRA, NRZ) of Escherichia coli. Additionally, the derived amino acid sequence of the truncated open reading frame of norA showed striking resemblance to the α-subunits (NarG, NarZ) of the E. coli nitrate reductases.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Smith JA, Seidman JG, Struhl K (eds) (1989) Current protocols in molecular biology. Wiley, New York.
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7: 1513–1519.
Blasco F, Iobbi C, Giordano G, Chippaux M, Bonnefoy V (1989) Nitrate reductase of Escherichia coli: completion of the nucleotide sequence of the nar operon and reassessment of the role of the α and β subunits in iron binding and electron transfer. Mol Gen Genet 218: 249–256.
Blasco F, Iobbi C, Ratouchniak J, Bonnefoy V, Chippaux M (1990) Nitrate reductases of Escherichia coli: sequence of the second nitrate reductase and comparison with that encoded by the narGHJI operon. Mol Gen Genet 222: 104–111.
Bock E, Sundermeyer-Klinger H, Stackebrandt E (1983) New facultative lithoautotrophic nitrite-oxidizing bacteria. Arch Microbiol 136: 281–284.
Bruschi M, Guerlesquin F (1988) Structure, function and evolution of bacterial ferredoxins. FEMS Microbiol Rev 54: 155–175.
Chaudhry GR, McGregor CH (1983) Escherichia coli nitrate reductase subunit A: its role as the catalytic site and evidence for its modification. J Bacteriol 154: 387–394.
Darlison MG, Guest JR (1984) Nucleotide sequence encoding the iron-sulfur protein subunit of the succinate dehydrogenase of Escherichia coli. Biochem J 223: 507–517.
Edman P (1956) Mechanism of phenylisothiocyanate-hydantoin degradation of peptides. Nature 177: 667–668.
Elvin CM, Thompson PR, Argall ME, Hendry P, Stamford NPJ, Lilley PE (1990) Modified bacteriophage lambda promoter vectors for overproduction of protein in Escherichia coli. Gene 87: 123–126.
Freitag A, Bock E (1990) Energy conservation in Nitrobacter. FEMS Microbiol Lett 66: 157–162.
Guigliarelli B, Asso M, More A, Augier V, Blasco F, Pommier F, Giordano G, Bertrand P (1992) EPR and redox centers in nitrate reductases-A and reductases-Z from Escherichia coli —Evidence for a high-potential and low-potential class and their relevance in the electron-transfer mechanism. Eur J Biochem 207: 61–68.
Henikoff S, Wallace JC, Brown J (1990) Finding protein similarities with nucleotide sequence databases. Methods Enzymol 183: 111–132.
Hochstein LI, Tomlinson GA (1988) The enzymes associated with denitrification. Annu Rev Microbiol 42: 231–261.
Johnson MK, Bennet DE, Morningstar JE (1985) The iron-sulfur cluster composition of Escherichia coli nitrate reductase. J Biol Chem 260: 5456–5463.
Koops HP, Harms H (1985) Deoxyribonucleic acid homologies among 96 strains of ammonia-oxidizing bacteria. Arch Microbiol 141: 214–218.
Krüger B, Meyer O, Nagel M, Andreesen JR, Meincke M, Bock E, Blümle S, Zumft WG (1987) Evidence for the presence of bactopterin in the eubacterial molybdoenzymes nicotinic acid dehydrogenase, nitrite oxidoreductase, and respiratory nitrate reductase. FEMS Microbiol Lett 48: 225–227.
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of the bacteriophage T4. Nature 227: 680–685.
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from microorganism. J Microbiol 3: 208–218.
Matsudaira I (1987) Sequence from picomole quantities of protein electroblottet onto polyvinyliden difluoride membranes. J Biol Chem 262: 10035–10038.
Meincke M, Bock E, Kastrau D, Kroneck PMH (1992) Nitrite oxidoreductase from Nitrobacter hamburgensis: redox centers and their catalytic role. Arch Microbiol 158: 127–131.
Milde K, Bock E (1984) Isolation and partial characterization of inner and outer membrane fractions of Nitrobacter hamburgensis. FEMS Microbiol Lett 21: 137–141.
Sambrook J, Fritsch EF, Maniatis T (eds) (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74: 5463–5467.
Schuler GD, Altschul SF, Lipman DJ (1991) A workbench for multiple alignment construction and analysis. Protein 9: 180–190.
Sundermeyer-Klinger H, Meyer W, Warninghoff B, Bock E (1984) Membrane-bound nitrite oxidoreductase of Nitrobacter: evidence for a nitrate reductase system. Arch Microbiol 140: 153–158.
Tanaka Y, Fukumori Y, Yamanaka T (1983) Purification of cytochrome a 1c1 from Nitrobacter agilis and characterization of nitrite oxidation system of the bacterium. Arch Microbiol 135: 265–271.
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of M13mp18 and pUC19 vectors. Gene 33: 103–119.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kirstein, K., Bock, E. Close genetic relationship between Nitrobacter hamburgensis nitrite oxidoreductase and Escherichia coli nitrate reductases. Arch. Microbiol. 160, 447–453 (1993). https://doi.org/10.1007/BF00245305
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00245305