Abstract
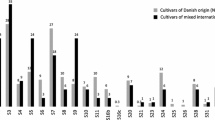
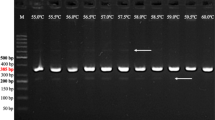
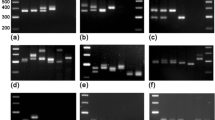
Primers for the polymerase chain reaction (PCR) were tailored to selectively amplify RFLP marker alleles associated with resistance and susceptibility for powdery mildew in cereals. The differentiation between marker alleles for susceptible and resistant genotypes is based on the discrimination of a single nucleotide by using allele-specific oligonucleotides as PCR primers. The PCR assays developed are diagnostic for RFLP alleles at the loci MWG097 in the barley genome and Whs350 in the wheat genome. The first marker locus is closely linked to MlLa resistance in barley, while the latter is linked to Pm2 resistance locus in wheat. PCR analysis of 31 barley and 30 wheat cultivars, with some exceptions, verified the presence or absence of the resistance loci investigated. These rapid PCR-based approaches are proposed as an efficient alternative to conventional procedures for selecting powdery mildew-resistant genotypes in breeding programs.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Anonymous (1995) Beschreibende Sortenliste Getreide, Mais, Ölfrüchte, Leguminosen, Hackfrüchte. Bundessortenamt. Landbuchverlag, Hannover
D'Ovidio R, Anderson OD (1994) PCR analysis to distinguish between alleles of a member of a multigene family correlated with wheat bread-making quality. Theor Appl Genet 88:759–763
Feuillet C, Messmer M, Schachermayr G, Keller B (1995) Genetic and physical characterization of the Lr1 leaf rust resistance locus in wheat (Triticum aestivum L.). Mol Gen Genet 248:553–562
Giese H, Holm-Jensen AG, Jensen HP, Jensen J (1993) Localization of the Laevigatum powdery mildew resistance gene to barley chromosome 2 by the use of RFLP markers. Theor Appl Genet 85:897–900
Görg R, Hollricher K, Schulze-Lefert P (1993) Functional analysis and RFLP-mediated mapping of the Mlg resistance locus in barley. Plant J 3:857–866
Hartl L, Weiss H, Zeller FJ, Jahoor A (1993) Use of RFLP markers for the identification of alleles of the Pm3 locus conferring powdery mildew resistance in wheat (Triticum aestivum L.). Theor Appl Genet 86:959–963
Hartl L, Weiss H, Stephan U, Zeller FJ, Jahoor A (1995) Molecular identification of powdery mildew resistance genes in common wheat (Triticum aestivum L.). Theor Appl Genet 90:601–606
Hilbers S, Fischbeck G, Jahoor A (1992) Localization of the Laevigatum resistance gene MlLa against powdery mildew in the barley genome by the use of RFLP markers. Plant Breeding 109: 335–338
Hinze K, Thompson RD, Ritter E, Salamini F, Schulze-Lefert P (1991) Restriction fragment length polymorphism-mediated targeting of the ml-o resistance locus in barley (Hordeum vulgare). Proc Natl Acad Sci USA 88:3691–3695
Huang M, Arnheim N, Goodman MF (1992) Extension of base mispairs by Taq DNA polymerase: implications for single nucleotide discrimination in PCR. Nucleic Acids Res 20:4567–4573
Islam AKRM, Shepherd KW, Sparrow DHB (1981) Isolation and characterization of euplasmic wheat-barley addition lines. Heredity 46:161–174
Jensen HP, Christensen E, Jørgensen JH (1992) Powdery mildew resistance genes in 127 northwest european spring barley varieties. Plant Breed 108:210–228
Kwok S, Kellogg DE, McKinny N, Spasic D, Goda L, Sninsky JJ (1990) Effects of primer-template mismatches on the polymerase chain reaction: human immunodeficiency virus type-I model studies. Nucleic Acids Res 18:999–1005
Ma ZQ, Sorrells ME, Tanksley SD (1994) RFLP markers linked to powdery mildew resistance genes Pm1, Pm2, Pm3 and Pm4 in wheat. Genome 37:871–875
Niewöhner J, Salamini F, Gebhardt C (1995) Development of a PCR assay diagnostic for RFLP marker alleles closely linked to alleles Gro1 and H1, conferring resistance to the root cyst nematode Globodera rostochiensis potato. Mol Breed 1:65–78
Olson M, Hood L, Cantor CH, Botstein D (1989) A common language for the physical mapping of the human genome. Science 24:1434–1435
Röder MS, Plaschke J, König SU, Börner A, Sorrells ME, Tanksley SD, Ganal MW (1995) Abundance, variability and chromosomal location of microsatellites in wheat. Mol Gen Genet 246:327–333
Schachermayr G, Siedler H, Gale MD, Winzeler H, Winzeler M, Keller B (1994) Identification and localization of molecular markers linked to the Lr9 leaf rust resistance gene of wheat. Theor Appl Genet 88:110–115
Schachermayr GM, Messmer MM, Feuillet C, Winzeler H, Winzeler M, Keller B (1995) Identification of molecular markers linked to the Agropyron elongatum-derived leaf rust resistance gene Lr24 in wheat. Theor Appl Genet 90:982–990
Schönfeld M, Ragni A, Fischbeck G, Jahoor A (1996) RFLP mapping of three new loci for resistance genes to powdery mildew (Erysiphe graminis f. sp. hordei) in barley. Theor Appl Genet 93:48–56
Schüller C, Backes G, Fischbeck G, Jahoor A (1992) RFLP markers to identify the alleles on the Mla locus conferring powdery mildew resistance in barley. Theor Appl Genet 84:330–338
Sears ER (1966) Nullisomic-tetrasomic combinations in hexaploid wheat. In: Riley R, Lewis KR (eds) Chromosome manipulation and plant genetics. Oliver and Boyd, Edinburgh, pp 29–45
Shattuck-Eidens DM, Bell RN, Mitchell JT, McWorther VC (1991) Rapid detection of maize DNA sequence variation. GATA 8:240–245
Talbert LE, Blake NK, Chee PW, Blake TK, Magyar GM (1994) Evaluation of ‘sequence-tagged-site’ PCR products as molecular markers in wheat. Theor Appl Genet 87:789–794
Tragoonrung S, Kanazin V, Hayes PM, Blake TK (1992) Sequencetagged-site-facilitated PCR for barley genome mapping. Theor Appl Genet 84:1002–1008
Williams GK, Kubelik AR, Kenneth JL, Rafalski A, Scott VT (1991) DNA polymorphism amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Xu L, Hall BG (1994) SASA: a simplified, reliable method for allele-specific amplification of polymorphic sites. BioTechniques 6:44–45
Author information
Authors and Affiliations
Additional information
Communicated by J. W. Snape
Rights and permissions
About this article
Cite this article
Mohler, V., Jahoor, A. Allele-specific amplification of polymorphic sites for the detection of powdery mildew resistance loci in cereals. Theoret. Appl. Genetics 93, 1078–1082 (1996). https://doi.org/10.1007/BF00230128
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00230128