Abstract
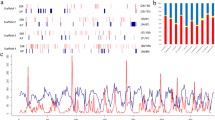
Three Plantago species were surveyed for within- and between-population variation at DNA sequences detected with the M13 minisatellite probe. The levels and patterns of variation detected by this probe correspond to those expected from the mating systems of the species. The highly-selfing species P. major has a relatively low variability of minisatellite sequences within populations and considerable differentiation between populations. The outcrossing species P. lanceolata exhibits higher minisatellite variability within populations and moderate differentiation between populations. P. coronopus, with a mixed mating system, has levels of variation intermediate between P. major and P. lanceolata. The levels of variation within and between populations corresponds, in general, to the levels of allozyme variation determined in an earlier study. Mating system and population structure appear to have a major influence on M13-detected fragment variation.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Brown AHD (1979) Enzyme polymorphism in plant populations. Theor Pop Biol 15:1–42
Burke T, Bruford MW (1987) DNA fingerprinting in birds. Nature 327:149–152
Chakraborty R, Kidd KK (1991) The utility of DNA typing in forensic work. Science 254:1735–1739
Dallas JF (1988) Detection of DNA “fingerprints” of cultivated rice by hybridization with a human minisatellite DNA probe. Proc Natl Acad Sci USA 85:6831–6835
Davis SK, Strassmann JE, Hughes C, Pletscher LS, Templeton AR (1990) Population structure and kinship in Polistes (Hymenoptera, Vespidae): an analysis using ribosomal DNA and protein electrophoresis. Evolution 44:1242–1254
Jeffreys AJ, Wilson V, Thein SL (1985a) Hypervariable ‘minisatellite’ regions in human DNA. Nature 314:67–73
Jeffreys AJ, Wilson V, Thein SL (1985b) Individual-specific ‘fingerprints’ of human DNA. Nature 316:76–79
Jeffreys AJ, Royle NJ, Wilson V, Wong Z (1988) Spontaneous mutation rates to new length alleles at tandem-repetitive hypervariable loci in human DNA. Nature 332:278–281
Layton CR, Ganders FR (1984) The genetic consequences of contrasting breeding systems in Plectritis (Valerianaceae). Evolution 38:1308–1325
Lewontin RC, Hartl DL (1991) Population genetics in forensic DNA typing. Science 254:1745–1750
Lotz LAP (1989) Variation in life-history characteristics between and within populations of Plantago major L. PhD thesis, Nijmegen
Lotz LAP, Blom CWPM (1986) Plasticity in life-history traits of Plantago major L. ssp. pleiosperma Pilger. Oecologia 69:25–30
Lynch M (1990) The similarity index and DNA fingerprinting. Mol Evol Biol 7:478–484
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Nei M (1975). Molecular population genetics and evolution. North Holland, Amsterdam
Nybom H (1990) Genetic variation in ornamental apple trees and their seedlings (Malus, Rosaceae) revealed by DNA ‘fingerprinting’ with the M13 repeat probe. Hereditas 113:17–28
Nybom H, Hall HK (1991) Minisatellite DNA ‘fingerprints’ can distinguish Rubus cultivars and estimate their degree of relatedness. Euphytica 53:107–114
Nybom H, Schaal BA (1990) DNA “fingerprints” reveal genotypic distributions in natural populations of blackberries and raspberries (Rubus L., Rosaceae). Am J Bot 77:883–888
Nybom H, Rogstad SH, Schaal BA (1990) Genetic variation detected by use of the M13 “DNA fingerprint” probe in Malus, Prunus and Rubus (Rosaceae). Theor Appl Genet 79:153–156
Rogstad SH, Patton JC, Schaal BA (1988) M13 repeat probe detects DNA minisatellite-like sequences in gymnosperms and angiosperms. Proc Natl Acad Sci USA 85:9176–9178
Rogstad SH, Herwaldt BL, Schlesinger PH, Krogstad DJ (1989) The M13 repeat probe detects RFLPs between two strains of the protozoan malaria parasite Plasmodium falciparum. Nucleic Acids Res 17:3610
Rogstad SH, Wolff K, Schaal BA (1991) Clonal structure and genetic variation in Asimina triloba revealed by “DNA fingerprinting”. Am J Bot 78:1391–1396
Rogstad SH, Nybom H, Schaal BA (1991) The tetrapod “DNA fingerprinting” M13 repeat probe reveals genetic diversity and clonal growth in quaking aspen (Populus tremuloides, Salicaceae). Plant Syst Evol 175:115–123
Ryskov AP, Jincharadze AG, Prosnyak MI, Ivanov PL, Limborska SA (1988) M13 phage DNA as a universal marker for DNA fingerprinting of animals, plants and microorganisms. Fed Eur Biochem Soc 233:388–392
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphism in barley: Mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbour Laboratory, Cold Spring Harbor, New York
Sokal RR, Rohlf F (1981) Biometry. W. H. Freeman, San Francisco
Van Dijk H, Van Delden W (1981) Genetic variability in Plantago species in relation to their ecology. 1. Genetic analysis of the allozyme variation in Plantago major subspecies. Theor Appl Genet 60:285–290
Van Dijk H, Wolff K, De Vries A (1988) Genetic variability in Plantago species in relation to their ecology. 3. Structure of populations of P. major, P. lanceolata and P. coronopus. Theor Appl Genet 75:518–528
Vassart G, Georges M, Monsieur R, Brocas H, Lequarre AS, Christophe D (1987) A sequence in M13 phage detects hyper-variable minisatellites in humans and animal DNA. Science 235:683–684
Westneat DF, Noon WA, Reeve HK, Aquadro CF (1988) Impro ved hybridization conditions for. DNA ‘fingerprints’ probed with M13. Nucleic Acids Res 16:4161
Wetton JH, Carter RE, Parkin DT, Walters D (1987) Demographic study of a wild house sparrow population by DNA fingerprinting. Nature 327:147–149
Wolff K (1988) Natural selection in Plantago species: a genetical analysis of ecologically-relevant morphological variability. PhD thesis, University of Groningen
Wolff K (1991 a) Genetic analysis of morphological variability in three Plantago species with different mating systems. Theor Appl Genet 81:111–118
Wolff K (1991 b) Genetic analysis of allozyme variability in three Plantago species and a comparison to morphological variability. Theor Appl Genet 81:119–126
Wolff K, Schaal B (1992). Chloroplast DNA variation within and among five Plantago species. J Evol Biol 5:325–344
Wolff K, Friso B, Van Damme JMM (1988) Outcrossing rates and male sterility in natural populations of Plantago coronopus. Theor Appl Genet 76:190–196
Wright S (1965) The interpretation of population structure by F-statistics with special regard to system of mating. Evolution 19:395–420
Wyman AR, White R (1980) A highly-polymorphic locus in human DNA. Proc Natl Acad Sci USA 77:6754–6758
Author information
Authors and Affiliations
Additional information
Communicated by P. M. A. Tigerstedt
Rights and permissions
About this article
Cite this article
Wolff, K., Rogstad, S.H. & Schaal, B.A. Population and species variation of minisatellite DNA in Plantago . Theoret. Appl. Genetics 87, 733–740 (1994). https://doi.org/10.1007/BF00222899
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00222899