Abstract
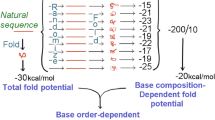
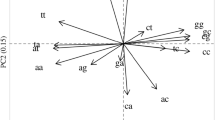
To an approximation Chargaff's rule (%A = %T; %G = %C) applies to single-stranded DNA. In long sequences, not only complementary bases but also complementary oligonucleotides are present in approximately equal frequencies. This applies to all species studied. However, species usually differ in base composition. With the goal of understanding the evolutionary forces involved, I have compared the frequencies of trinucleotides in long sequences and their shuffled counterparts. Among the 32 complementary trinucleotide pairs there is a hierarchy of frequencies which is influenced both by base composition (not affected by shuffling the order of the bases) and by base order (affected by shuffling). The influence of base order is greatest in DNA of 50% G + C and seems to reflects a more fundamental hierarchy of dinucleotide frequencies. Thus if TpA is at low frequency, all eight TpA-containing trinucleotides are at low frequency. Mammals and their viruses share similar hierarchies, with intra- and intergenomic differences being mainly associated with differences in base composition (percentage G + C). E. coli and, to a lesser extent, Drosophila melanogaster hierarchies differ from mammalian hierarchies; this is associated with differences both in base composition and in base order. It is proposed that Chargaff's rule applies to single-stranded DNA because there has been an evolutionary selection pressure favoring mutations that generate complementary oligonucleotides in close proximity, thus creating a potential to form stem-loops. These are dispersed throughout genomes and are rate-limiting in recombination. Differences in (G + C)% between species would impair interspecies recombination by interfering with stem-loop interactions.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Alff-Steinberger C (1984) Evidence for a coding pattern on the noncoding strand of the E. coli. genome. Nucleic Acids Res 12:2235–2241
Bernstein C, Bernstein H (1991) Aging, sex and DNA repair. Academic Press, San Diego
Bird AP (1980) DNA methylation and the frequency of CPO in animal DNA. Nucleic Acids Res 8:1499–1504
Chargaff E (1951) Structure and function of nucleic acids as cell constituents. Fed Proc 10:654–659
Filipski J (1990) Evolution of DNA sequence. Contributions of mutational bias and selection to the origin of chromosomal compartments. Adv Mutagen Res 2:1–54
Forsdyke DR (1995a) A stem-loop “kissing” model for the initiation of recombination and the origin of introns. Mol Biol Evol (in press)
Forsdyke DR (1995b) Conservation of stem-loop potential in introns of snake venom phospholipase A2 genes. An application of FORS-D analysis. Mol Biol Evol (in press)
Forsdyke DR (1995c) Different biological species “broadcast” their DNAs at different (G + C)% “wavelengths.” Proc Can Fed Biol Socs 38:107
Goebel SJ, Johnson GP, Perkus ME, Davis SW, Winslow JP, Paoletti E (1990) The complete DNA sequence of vaccinia virus. Virology 179:247–266
Gribskov M, Devereux J (1991) Sequence analysis primer. Stockton Press, New York
Karkas JD, Rudner R, Chargaff E (1968) Separation of B. subtilis DNA into complementary strands, II. Template functions and composition as determined by transcription with RNA polymerase. Proc Natl Acad Sci USA 60:915–920
Lawn RM, Efstratiadis A, O'Connell C, Maniatis T (1980) The nucleotide sequence of the human β-globin gene. Cell 21:647–651
Le S-Y, Maizel JV (1989) A method for assessing the statistical significance of RNA folding. J Theor Biol 138:495–510
Martin-Gallardo A, McCombie WR, Gocayne JD, Fitzgerald MG, Wallace S, Lee BMB, Lamerdin J, Trapp S, Kelley JM, Liu L-I, Dubnick M, Johnston-Dow LA, Kerlavage AR, Jong P de, Carrano A, Fields C, Venter JC (1992) Automated DNA sequencing and analysis of 106 kilobases from human chromosome 19g13.3. Nature Genet 1:34–39
McGeoch DJ, Dalrymple MA, Davison AJ, Dolan A, Frame MC, MeNab D, Perry LJ, Scott JE, Taylor P (1988) The complete DNA sequence of the long unique region in the genome of herpes simplex virus type 1. J Gen Virol 69:1531–1574
Murchie AIH, Bowater R, Aboul-ela F, Lilley DMJ (1992) Helix opening transitions in supercoiled DNA. Biochem Biophys Acta 1131: 1–15
Nussinov R (1981) Eukaryotic dinucleotide preference rules and their implications for degenerate codon usage. J Mol Biol 149:125–131
Nussinov R (1984) Strong doublet preferences in nucleotide sequences and DNA geometry. J Mol Evol 20:111–119
Pradhu VV (1993) Symmetry observations in long nucleotide sequences. Nucleic Acids Res 21:2797–2800
Proffitt JH, Davie JR, Swinton D, Hattman S (1984) 5-Methylcytosine is not detectable in Saccharomyces cerevisiae DNA. Mol Cell Biol 4:985–988
Urieli-Shoval S, Gruenbaum Y, Sedat J, Razin A (1981) The absence of detectable methylated bases in Drosophila melanogaster DNA. FEBS Lett 146:148–152
Ward GK, McKenzie R. Zannis-Hadjopoulos M, Price GB (1990) The dynamic distribution and quantification of DNA cruciforms in eukaryotic nuclei. Exp Cell Res 188:235–246
Watson JD, Crick FHC (1953) Genetical implications of the structure of deoxyribonucleic acid. Nature 171:964–967
Yomo T, Ohno S (1989) Concordant evolution of coding and noncoding regions of DNA made possible by the universal rule of TA/CG deficiency-TG/CT excess. Proc Natl Acad Sci USA 86: 8452–8456
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Forsdyke, D.R. Relative roles of primary sequence and (G + C)% in determining the hierarchy of frequencies of complementary trinucleotide pairs in DNAs of different species. J Mol Evol 41, 573–581 (1995). https://doi.org/10.1007/BF00175815
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00175815