Abstract
Regarded as highly successful in their clinical applications, therapeutic proteins are now widely applied for the precise treatment of several diseases. Common forms of therapeutic proteins include enzymes, antibodies, and recombinant proteins. Here, we discuss different aspects of the clinical applications of protein-based therapeutics, including therapeutic proteins, their mechanisms and metabolism, challenges, precision medicine, and computer-aided drug designing. In addition, an overview of recently approved therapeutic proteins is provided. Conclusively, this chapter delivers comprehensive information on clinical applications of protein-based therapeutics, emerging trends, and challenges.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Therapeutic proteins
- Precision medicine
- Computer-aided drug designing
- Enzymes
- Antibodies
- Recombinant proteins
2.1 Introduction
Proteins, undoubtedly one of the most versatile macromolecules, have been extensively studied for their widespread biomedical and therapeutic applications [1]. Due to their properties like natural origin, biodegradability, biocompatibility, recognition by cells, reduced immunogenic potential, and natural bioactivity, therapeutic proteins offer several advantages compared to synthetic therapeutic molecules. In addition, they are easy to functionalize and engineered for specific locations or applications through alteration of their primary amino acid sequences. Presently, proteins form a dominating segment of the pharmaceutical industry as they have tremendous therapeutic potential against various diseases and syndromes. The demand for protein-based medicines has significantly augmented, mainly due to increased medical awareness and the prevalence of chronic diseases. From $140 billion in 2016, the global protein therapeutics market is projected to reach $566 billion by 2030 [2]. However, the high costs of therapeutic proteins and stringent government regulations remain significant challenges that negatively impact market growth.
Based on their molecular types, therapeutic proteins are classified into several types, such as antibodies, enzymes, fusion proteins, recombinant proteins, blood factors, anticoagulants, growth factors, interferon, hormones, etc. They act differently on biological or drug targets. Some common pharmacological activities of therapeutic proteins are replacing deficient or abnormal proteins, interfering with molecules or pathways, and delivering other molecules [3]. Protein-based therapeutics have evolved a lot with technological advances in the fields of drug discovery and protein engineering. For customized drug designing, the most critical aspects of protein-based molecules are understanding their mechanism of action and the structure-function relationship. Continuous improvements in traditionally existing therapies and methods to identify drug targets have resulted in developing drugs with better efficacy and targeted clinical applications [4]. This chapter provides a comprehensive overview of protein-based therapeutics, their mechanism, clinical applications, and challenges. A detailed illustration is provided about enzymes and antibodies as therapeutic proteins, followed by the introduction, applications, and prospects of precision medicine, an emerging, highly innovative, and targeted medicine approach that looks into an individual’s genetics, environment, and lifestyle. The discovery and development of drugs consist of very complex and time-consuming processes. However, recent decades have seen much growth in this field due to the application of computer-aided drug design (CADD). Thus, we also briefly discuss CADD approaches in drug designing and their application in developing protein therapeutics. The chapter concludes with a discussion of recently approved therapeutic proteins, emerging trends, challenges, and opportunities in the field, followed by the safety and efficacy of protein-based drugs.
2.2 Enzyme as Biologics
In contrast to regular therapeutics, biologics are comparatively larger molecules having higher molecular weights. Most of these molecules are unstable at room temperature and require refrigeration for storage. Also, biologics are produced by complex processes, and at times, even slight changes in their formulation might lead to degradation of their efficacy in disease management. It is nearly impossible to produce an exact copy of biologics, and thus, nearly similar biologics, compared to the original one, are manufactured that are referred to as biosimilars.
In 1878, a German physiologist Wilhelm Kühne coined the word “enzyme” [5, 6]. The enzymes are responsible for biological catalysis and are also called biocatalysts. Biocatalysis is a remarkable property of enzymes to speed up the specific biological reaction in living organisms. The study of enzyme kinetics provides information about a diverse range of reactions, metabolism, cell regulation, and how poisons and drugs affect the enzymes [7, 8]. The first enzyme discovered was diastase (a mixture of amylases), which catalyzes the hydrolysis of starch into maltose. It has a wide range of clinical, food, forensic, biochemical, medicinal, pharmaceutical, and environmental applications [7]. Almost all enzymes are proteins, and a functional enzyme has different components, such as holoenzyme (functional unit of enzyme and conjugated protein), apoenzyme (polypeptide segment of the enzyme and inactive precursor), coenzyme (small organic moiety), or zymogens (simple protein enzymes, which are secreted in an inactive form). Enzymes are classified into six functional classes that catalyze a specific reaction, that is, oxidoreductases, transferases, hydrolases, lyases, isomerases, and ligases/lyases [9]. This classification enables the identification and separation of diverse chemical reactions in living organisms. The functionality of an enzyme is intrinsically linked to its three-dimensional (3D) structure and is determined by the shape when it binds to the substrate that creates an ideal fit for catalysis.
2.2.1 Biological Process: How Enzymes Work?
A large number of biochemical reactions occur in the human body to carry out essential metabolic processes. Thousands of enzymes produced in the human body help to accelerate metabolism, growth, digestion, building muscle, healing, destroying toxins, reproduction, liver function, nerve function, and so on [8, 10]. The functions of enzymes are strongly affected by pH and temperature [7, 11]. The lock and key model related to substrate-enzyme interaction was postulated by Emil Fischer. The key signifies a substrate, and the lock and keyhole represent the enzyme and its active site. The shape and size of the substrate are complementary to the active site of the enzyme [5, 7]. The substrate perfectly binds at the active site and forms the enzyme-substrate complex, allowing rapid biochemical reaction.
2.2.2 Therapeutic Enzymes
The concept of therapeutic enzymes has been exploited for several decades [10]. Some enzymes are the preferred markers of various diseases, including cancer, infectious disease, myocardial infarction, clotting, pancreatitis, inherited diseases, neurodegenerative disorders, etc. These markers help in disease management via the diagnosis, prognosis, and assessment of responses to a therapeutic intervention [6]. Effective therapeutic solutions via various enzymes are helpful for the treatment of several diseases. The wide variety of uses of therapeutic enzymes is depicted in Table 2.1 and Fig. 2.1.
2.3 Antibodies as Biologics
Antibodies are the proteins produced to bind a specific antigen. The main task of antibodies is to get associated with the specific substance chemically considered alien by our body, such as bacteria, viruses, and other foreign substances in the blood, to neutralize them. Biochemically, antibodies are immunoglobulins, protective proteins produced by our immune system when it recognizes the presence of any foreign substance, commonly referred to as an antigen.
Biologics have the potential to mount an immune response against them, reducing their efficacy, and sometimes, it becomes life-threatening due to the generation of antibiologics antibodies. To compensate, other cotherapy options are used to treat a particular disease. It is always advisable to keep track of antibiologics antibodies in a patient’s blood using therapeutic drug monitoring. It is one of the most prominent techniques that detects the presence of biologics and antibiologics antibodies in the blood [12]. If antibiologics antibodies are present above a permissible limit and the amount of biologics is insufficient to reduce the inflammation, immune-modulators are introduced into the treatment regimen [13]. Antibody-based biologics can be subdivided into three major categories: monoclonal antibody (mAb) products, non-mAb products, and vaccines [14]. The working mechanism of vaccines relies solely on mounting immune response and is mainly used for only prophylactic purposes rather than therapeutic uses and, thus, not elaborated here.
2.3.1 Monoclonal Antibody Products as Biologics
Monoclonal antibodies (mAbs) are immunoglobulin G (IgG) that imitate the natural IgG function within the body. Their role is to bind to the foreign particles to neutralize them. Fc Fusion proteins (FcFPs), consisting of the Fc receptor of the IgG, can also bind to a modified protein. Like natural IgGs, mAbs, and FcFPs bind to extracellular targets, cells, or pathogens to neutralize them by disrupting their functions and removing them from circulation or modulating or imitating their activity. For example, inflammatory cytokines such as tumor necrosis factor-α (TNF-α) or interleukin-1b (IL-1b) neutralize the infected cells and result in immunosuppression [15]. The mAbs are usually derived from mice and rats and humanized to various degrees by engineering amino acid substitutions that make them similar to the human gene sequence through recombinant DNA (rDNA) technologies.
2.3.1.1 Biological Characteristics of mAbs
The mAbs are monospecific antibodies made from identical clones of a unique parent cell [16]. The essential biological characteristics of mAbs are listed below.
-
1.
They show edacity effects against target cells.
-
2.
They have the ability to obstruct protein-protein interactions with different targets like the serum, extracellular, and membrane-bound proteins.
-
3.
They can mediate multiple processes like antibody-dependent cellular cytotoxicity (ADCC), complement-dependent cytotoxicity (CDC), antibody-dependent cellular phagocytosis (ADCP), antibody-mediated immune complex formation with clearance, or a completely silent Fc with none of the above activities [17].
-
4.
They can penetrate certain tissues.
-
5.
The mAbs have minimal drug-drug interactions.
-
6.
They show little or no non-mechanism-mediated toxicity.
-
7.
They also have little or no off-target activity or drug-metabolism-related issues.
2.3.1.2 Applications of mAbs
Modifications in mAbs, such as Fab fragments, and bifunctional antibodies, are used to produce various biologics to treat a variety of diseases.
2.3.1.2.1 Fab Fragments
Fab fragments are the single binding site for the antigen. The important clinical applications of Fab fragments are as follows:
-
1.
Caplacizumab is a humanized, bivalent, variable-domain-only fragment with a high affinity for the von Willebrand factor (VWF). The interaction between VWF and platelet plays a central role in microvascular thromboses in patients with thrombotic thrombocytopenic purpura (TTP). Caplacizumab disrupts the interactions between VWF multimers and platelets and is used to treat acquired TTP conditions [18].
-
2.
Ranibizumab is a recombinant humanized Fab fragment that binds to and inhibits the human vascular endothelial growth factor A (VEGF-A) [19]. It inhibits the binding of VEGF to its receptors and slows down the related vision loss, and is used in treating age-related macular degeneration.
-
3.
Abciximab is a Fab antibody fragment derived from a chimeric human-murine mAb (7E3) that binds to platelet IIb/IIIa receptors, resulting in steric hindrance and thus inhibiting the platelet aggregation [20]. Abciximab has been used in unstable angina and reduction of thrombosis in various coronary stenting procedures.
2.3.1.2.2 Bifunctional Antibodies
Bifunctional antibodies are antibodies with dual specificity. Both the immunoglobin chains are fused together to form a single antibody molecule. A few examples of bifunctional antibodies are as follows:
-
1.
Emicizumab binds to two coagulation factors (factor IXa and factor X), taking the place of activated factor VIII (factor VIIIa) in the coagulation cascade [21]. The mAb is used for the prophylaxis of hemophilia patients.
-
2.
Blinatumomab is a bispecific T cell and B cell engager molecule that binds to the cell surface proteins, CD3 present on T cells and CD19, present on precursor B-cell acute lymphoblastic leukemia (ALL) cells, and takes the site of cytotoxic T cells to recognize malignant B cells [22].
-
3.
Catumaxomab is a bispecific trifunctional antibody that binds to the T-cell surface molecule CD3 and epithelial cell adhesion molecule (EpCAM), a tumor cell surface marker, and receptor Fc region on dendritic cells [23]. This combination of antigen-binding helps in maintaining antitumor immune responses.
2.3.2 Non-Monoclonal Antibody Products as Biologics
The function of some natural proteins, enzymes, hormones, or peptides is disrupted in a healthy individual, resulting in the rise of physiological-function-related diseases. To counter this, non-monoclonal antibody (non-mAb) products with similar physiological effects are given to the patients [24]. Non-mAb therapy helps patients to recover by filling the physiological gaps. In general, the molecular weight of non-MAb products is more than 700 Da. These products are usually homogenous in nature and can be heterogenous only if they are glycosylated. Non-mAb products may include hormones, enzymes, interferons, interleukins, growth factors, or even natural or mimetic peptides [25]. The recent advancements in rDNA technology lead to the development of many hormones and non-MAbs under the biologics category to treat various diseases (Table 2.2).
2.4 Precision Medicine
The application of medicines is patient specific for treating a particular disease. It works better in certain patients compared to others, but the reason for this differential effectiveness was unknown a few decades back. Some patients face severe side effects, and others have fewer adverse effects when treated with anticancer drugs [36]. Over the past six decades, evidence has emerged indicating that a substantial portion of the variability in drug response is genetically determined as age, nutrition, health status, environmental exposure, and epigenetic factors play critical contributory roles [37]. The unique genetic constitution and differential gene expression in an individual is responsible for variation in drug responses. Precision medicine is an emerging practice that uses an individual’s genetic profile to guide decisions regarding disease prevention, diagnosis, and treatment [38, 39]. The genetic profiling of every patient is necessary before treating them with a particular medication to increase the treatment efficacy with fewer side effects. One can take guidance from the knowledge of an individual’s genome profile to preselect the treatment protocols that minimize adverse side effects or ensure more successful outcomes. After completing the human genome project, many advancements have occurred in the field of precision medicine. The individual genomic sequence data can indicate their susceptibility to certain diseases before they manifest, allowing physicians and patients to design a plan for monitoring and prevention [40]. The science of studying how the genetic variations affect drug responses in an individual is pharmacogenomics, an evolving field to better understand an individual’s responses to different treatments.
2.4.1 Benefits of Precision Medicine
Though the current use of precision medicine is limited, it has the potential to offer a wide range of applications in the coming years.
-
1.
Better medication selection: The adverse reactions to medicines, one of the drawbacks during treatment, leads to the death of many individuals. Although the Food and Drug Administration (FDA) approved drugs have to qualify the stringent parameters before coming to the market, there is either less or no information related to their response when given to certain individuals. The medication may appear safe for a large population; however, some patients may experience harmful side effects due to genetic variations. The study of pharmacogenomics may help to predict how a particular group of individuals will be able to respond to one specific medication and can play a deciding role in its selection.
-
2.
Safer dosing options: Following FDA approval, the standard dosage of a medication is decided based on factors such as liver or kidney function, weight, age, etc. However, these parameters might not be sufficient. Standard dosage may work well for one group of individuals but may be toxic for another group because of underlying genetic variations. Currently, clinicians generally decide which medication is appropriate for treating a particular disease based on their diagnosis. Once the field of pharmacogenomics matures, clinicians can directly consider an individual’s genetic profile to decide the optimal medication dosage.
-
3.
Improvements in drug development: Pharmaceutical companies often spend years conducting research and clinical trials of a new drug before it reaches the market. Diagnostic and device firms and pharmaceutical companies typically have to test a product in a large cohort to ensure its safety and efficacy. The study of pharmacogenomics may help these companies to ensure the efficacy of drug testing. For example, if a company has an advanced idea that the drug can optimally work in participants with a particular type of genetic variations, and may cause adverse reactions to others, then those participants having adverse reaction can be excluded from the clinical trials. This will speed up the whole clinical trial process, and a specific population can be treated with the same medication.
2.4.2 Applications of Precision Medicine
A promising application of precision medicine lies in the discovery and manipulation of potential drug targets for the treatment of cancer. Precision medicine is used to treat chronic myeloid leukemia (CML). However, the discovery of molecular predispositions, that is, the presence of genetic variants, in various diseases, such as CML, has made it possible to design and develop specific therapeutic agents against novel molecular targets. With this applicability, precision medicine has identified a novel molecular target, Bcr/Abl tyrosine kinase. This kinase is an oncoprotein expressed in more than 95% of CML patients, and administration of a competitive inhibitor helps to achieve almost 80% cytogenetic responses in newly diagnosed CML patients [40].
In 2017, FDA approved more number of precision medicines. One of the drugs approved was pembrolizumab [41], which was marked as the first robust cancer therapy approved for clinical use based on a specific biomarker rather than a tumor’s location. Similarly, trastuzumab-dskt (Ogivri™) was approved as the first biosimilar agent that targets both stomach and breast tumors overexpressing the HER2 gene, facilitating competition and lowering healthcare costs [42]. Since these drugs are developed based on specific biomarkers, a need of companion genetic tests is needed to identify all the biomarkers. MSK-IMPACT™ (screens 468 genes) and Foundation One CdX™ (screens 324 genes) assays are examples of companion genetic tests. Both are solid tumor tests and massively parallel sequencing in vitro diagnostic tests [43]. These tests allow screening of multiple oncogenes to identify variants that might assist in the clinical management of the patients.
2.4.3 Future Prospects of Precision Medicine
In the past, physicians practiced intuition-based diagnosis and used their knowledge to provide medicine for the treatment of diseases. In the present time, clinicians rely on evidence-based diagnosis and treatment. They recommend the medicine based on evidence produced by scientific research, including clinical trials. In the future, precision medicines will be used according to algorithms that will consider the comprehensive information of an individual patient, including their genome, epigenetics, and lifestyle. Therefore, medicine in the twenty-first century must focus on attaining the four P’s: prediction, prevention, personalization, and participation, as stated by Dr. Leroy E. Hood [44]. Currently, patients are treated based on symptoms and diagnosis, which requires a transformation using precision medicine where the treatment is planned using the genetic profile of an individual. This evolution of medical treatment in the past, present, and future is summarized in Fig. 2.2.
2.5 Computer-Aided Drug Design
The discovery and development of new therapeutics is a complex and time-consuming process requiring much experimentation and research. Traditionally, a drug discovery takes an average of 10 to 15 years before it reaches the market for sale, with an estimated cost of 58.8 billion USD in 2015 [45, 46]. The high investment cost and failure rate of traditional methods prompted a need to utilize computational methods to aid drug discovery. Computer-aided drug design (CADD) refers to the use of computational tools and available data resources for designing, storing, analyzing, and modeling lead molecules to establish them as candidate drugs. This will facilitate studying chemical and biological interactions between the lead compounds and their biological targets. The technique systematically evaluates the potential lead candidates before their synthesis and in vitro and in vivo testing [47]. The computational resources comprise a screening process to select the best possible candidates and later estimate their physicochemical properties, such as absorption, distribution, metabolism, excretion, and toxicity (ADMET). CADD is routinely applied to discover and improve the quality of identified lead compounds quickly. Nowadays, the different applications of CADD techniques are used to speed up the drug design and discovery process.
2.5.1 Approaches of CADD in Designing Protein-Based Therapeutics
CADD strategies rely on the accessibility and availability of the 3D structure information of biological target and candidate molecules. This technique can be broadly divided into structure-based drug design (SBDD) and ligand-based drug design (LBDD), based on the information available for a protein receptor and ligands, respectively. The availability of the 3D structure of a receptor leads to the implementation of structure-based drug design methods. If only the ligand information is known, then ligand-based drug design can be adopted.
2.5.1.1 Structure-Based Drug Design Approach
This approach can only be used in the drug design process if the structure information of the protein receptor target is available. One can identify the active site and analyze the key amino acid residues responsible for its biological functions using the 3D structure. This information can then be used to create protein-based therapeutics that can outcompete the natural ligands, thereby interfering with the biological pathways to prevent the disease. The foundation for structure-based drug design was laid by the easy identification of binding cavities due to the availability of 3D structures of a large number of therapeutically important proteins. It is a precise, efficient, and rapid process, because it involves the 3D structure of a protein and knowledge about the disease at the molecular level [48]. SBDD is a multicycle process that leads to the development of potential lead candidates for clinical trials. The most notable success story involves FDA-approved drugs that inhibit the human immunodeficiency virus (HIV)-1, such as amprenavir, an inhibitor of HIV protease discovered through protein structure modeling and molecular dynamics (MD) simulations [49].
Along with success stories, some failures have also been reported. For example, RPX00023 was claimed to be an antidepressant with agonistic activity toward receptor 5-HT1A, but it behaved as an inhibitor of the receptor. Such failures highlight the limitations of SBDD strategies. To overcome these limitations, continuous improvements and developments have been incorporated into SBDD techniques, but still, consistent solutions need to be developed.
2.5.1.2 Ligand-Based Drug Design Approach
Ligand-based drug design (LBDD) is another method where information on ligand molecules is essential to use on previously unknown drug targets. LBDD methods are used when the experimental 3D structure of a receptor is not available. The structural and physicochemical properties of the known ligands that bind to the known drug target are analyzed to study their desired pharmacological activity [50]. The relationship between physicochemical properties and drug activity is known as a structure-activity relationship (SAR), which can be used to optimize known drugs or help design new drugs with improved activity [51]. LBDD methods also include substrate analogues that interact with the target molecule to produce the desired pharmacological effect.
The preparation of small-molecule libraries is the initial step of LBDD, where chemical structures of different compounds are created, processed, and analyzed in the form of molecular graphs. A molecular graph comprises a network of nodes and edges, in which atoms are represented as nodes and bonds between different atoms as edges. The molecular graphs communicate by using connection tables and their linear notations. The different sections and sub-sections of a connection table contain information related to atoms, atom types, connection types, and their coordinate positions in the 3D or 2D space. Specific file formats are used to store the ligand information, such as .mol2, .sdf, .pdb, etc. Simplified molecular-input line-entry specification (SMILE) and Wiswesser line notation are examples of linear notations where alphanumeric characters are used to store the ligand information. Linear notation is preferred for storing or transferring millions of small molecules due to its compactness compared to connection tables [52]. The quantitative structure-activity relationship (QSAR) and pharmacophore modeling concepts are used for designing drugs based on LBDD approaches.
2.5.2 Quantitative Structure-Activity Relationship
The quantitative structure-activity relationship (QSAR) is a computational method for determining the relationship between the structural properties of chemical compounds and their biological activities [47]. It is based on the principle that different structural properties yield different biological activities [53]. Structural properties include physicochemical properties, whereas biological activities correspond to pharmacokinetics, that is, ADMET, of drug molecules. The development of a QSAR model begins with recognizing a group of chemical entities or lead molecules that exhibit the desired biological activity. Then, suitable molecular descriptors are identified that are associated with various structural and physicochemical properties of the molecules of interest. Molecular descriptors are mathematical representations of molecular properties generated by associated algorithms. Finding the set of molecular descriptors is a significant step in constructing QSAR models. It helps reduce computational time, improve prediction performance, and better understand data in machine learning [54]. Further, statistical methods are employed to derive a quantitative relationship between molecular descriptors and their associated biological activity. Finally, the developed QSAR model is validated and tested for structural stability and predictive power. A QSAR model helps predict the movements of untested chemicals and aids in rational drug design through computer-aided simulation, molecular modeling, and virtual screening of suitable compounds.
2.5.3 Applications of CADD in Protein-Based Therapeutics
Therapeutic proteins are genetically engineered proteins that substitute abnormal or malfunctioned human proteins to cure a disease. In CADD, protein-drug interactions are simulated to determine their binding affinities. Virtual database screening helps screen large libraries efficiently to identify potent drugs that are likely to have high binding affinities to the target. Target may be any enzyme or protein linked to a specific disease. Structural information about the target is also required to learn about its functions. One can harness the structural information of proteins already available in the PDB database. The missing structural information can be predicted using bioinformatics approaches, such as homology modeling, threading, or ab initio predictions. Dhanavade et al. predicted the 3D structure of cysteine protease using molecular modeling, which degrades amyloid-β peptide, a major cause of Alzheimer’s disease (AD) [55,56,57]. In recent years, CADD has successfully identified potential drugs for treating several neurodegenerative disorders. ROCK-I and NOX2 are two of the most promising potential therapeutic targets for various neurodegenerative disorders [58, 59]. Inhibition of these two enzymes can help manage neurodegenerative disorders like autism spectral disorder, AD, and fragile X syndrome. Utilizing this information, Alokam et al. identified chemical entities that behave as dual inhibitors of these enzymes using a combination of pharmacophores and the molecular docking approach of CADD [60]. Also, in vitro validation demonstrated their inhibitory potentials to ROCK-I and NOX2.
In the COVID-19 pandemic, CADD served as a powerful tool for identifying therapeutic proteins against rapidly mutating SARS-CoV-2 [61]. The main protease (Mpro) enzyme is crucial for the survival of pathogen as it is involved in replication and maturation. The structure-based virtual screening successfully identified four compounds having the ability to disrupt the normal functioning of Mpro protein. Later, ADMET analysis, molecular docking, and MD simulations were applied to explore their binding conformational stability at the active site of Mpro protein. The study identified crucial ligand amino acid residues, such as GLN189, SER10, GLU166, ASN142, PHE66, and TRP132, that participate in stabilizing the protein-ligand interaction of SARS-CoV-2 Mpro [62]. Nowadays, machine learning approaches are used in conjunction with CADD to identify repurposed therapeutics [63]. Thus, CADD is serving as a rapid and promising technology in the development of protein-based therapeutics [64, 65].
2.6 Overview of Recently Approved Protein Therapeutics for Clinical Applications
Recombinant human proteins can be used as therapeutics for treating many illnesses such as diabetes mellitus and multiple sclerosis. The production of high-quality and functional recombinant proteins is crucial in drug therapy. Although many applications of recombinant proteins exist as potent therapeutics, the production of antidrug antibodies (ADAs) is a matter of concern that limits its use. The aggregates formed during the formulation of recombinant proteins lead to the breakage of immune system tolerance and result in the production of ADAs. Various strategies are applied to minimize the aggregation and reduce immunogenicity to make protein therapeutics safer and more efficient.
2.6.1 Diabetes
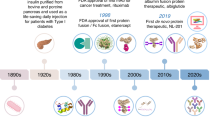
Purified from the porcine and bovine pancreas in the 1920s, insulin was the first therapeutic protein discovered. Since it had a nonhuman origin, its immunogenicity was expected. The patients suffering from diabetes mellitus were treated with insulin for over 80 years [66]. In addition to the source being nonhuman, early purification methods were also not up to the mark, resulting in the development of anti-insulin antibodies in most patients. To overcome the issue, therapeutic insulin is now mainly produced as recombinant human protein, and advanced purification methods take care of purity. Although recombinant insulin is a safer drug, it has been reported that ADAs, including subclasses of immunoglobulins, developed in about 50% of diabetic patients treated with recombinant human insulin [67, 68]. The presence of insulin autoantibodies in diabetes Type I patients hampers the interpretation of clinical data [69]. Several theories have been put forward to explain why ADAs develop against insulin in many patients. One of the most promising theories suggests the involvement of insulin ADAs with themselves and the formation of aggregates as insulin antibodies have a high tendency to self-associate. These aggregates contribute to high immunogenicity [70].
The glucagon-like peptide-1 (GLP-1) receptor agonist lixisenatide (lyxumia 1) was approved for the treatment of type 2 diabetes [71]. The GLP-1 receptor agonist is compared to other antidiabetic drugs, that is, exenatide, insulin glargine, metformin, sitagliptin, liraglutide, or placebo in type 2 diabetes patients [72]. It is linked with other biomolecules like lipids, carbohydrates, polyethylene glycol, or proteins to increase its efficacy. The GLP-1 receptor agonist, along with these conjugates, acts as cell-targeting peptides or cell-penetrating peptides. It induces insulin release and suppresses glucagon release in type 2 diabetes [73]. Another study showed that the C-peptide activates the phosphorylation of insulin receptor tyrosine kinase and glycogen synthase kinase 3 and results in the mobilization of insulin-responsive glucose transporter, increased amino acid uptake, and glycogen synthesis. This suggests that C-peptide signaling may cross-talk with the insulin pathway at the level of the insulin receptor [51]. The clinical studies indicate that the replacement of C-peptides in type 1 diabetic patients shows advantageous effects on somatic and autonomic diabetic peripheral neuropathy (DPN). Apart, the C-peptide also reduces the diabetes-induced glomerular hyperfiltration and, thus, decreases the excretion of urinary albumin [74].
2.6.2 Interferon-β
Relapsing-reemitting multiple sclerosis is generally treated by interferon-β (IFN-β) or recombinant human IFN-β (Rhu IFN-β). Though these are the most promising and efficient anti-inflammatory drugs for treating multiple sclerosis, many patients do not respond to them [75]. As suggested, this can be attributed to the production of neutralizing antibodies (nAbs) against the IFN-β [76]. IFN-β 1a (Avonex®, Rebif®) and IFN-β 1b (Betaseron®) products are available in the market that differ in their source of production, glycosylation pattern, amino acid sequences, and degree of aggregation [77]. These drugs can mount different levels of immunogenicity in patients as their formulation sources, administered routes, dosage, and frequency regimes differ.
Interestingly, patients with a history of developing nAbs, when treated with IFN-β, result in the disappearance of nAbs. This indicates that the production of nAbs does not form the memory and, thus, possibly does not involve in the classical immune response. Besides that, there is an increasing research interest in investigating and characterizing the formation of aggregates in IFN-β formulations and their potency in eliciting an immune response in patients by breaking immune tolerance [75]. One study reported that multiple sclerosis patients who received IFN-β 1b developed more nAbs than those who received IFN-β 1a [78]. This observation is most probably correlated to the levels of aggregates as IFN-β 1b formulation shows a higher degree of aggregation than IFN-β 1a [79]. The self-binding characteristic of IFN-β 1b is high, due to which they cluster together. Also, the lack of glycosylation in these molecules promotes aggregation [80]. The exact cause of the formation of ADAs is not well understood. However, aggregation is considered an essential contributory factor for immunogenicity in almost all cases, which requires comprehensive and exploratory studies to identify and validate the causes of their formation.
2.6.3 Cancer
An array of peptide-based therapeutics has been tested in preclinical models to check their efficacy in curing cancer. Therapeutics are developed based on a synthetic polymeric carrier elastin-like polypeptide (ELP), which can be synthesized in variable sequences and sizes to stabilize the therapeutic peptide and avoid crossing the placental interface to prevent fetal exposure and potential developmental effects [81]. The therapeutic peptides possess a targeting delivery feature to recognize cancer cells effectively. These peptides increase the specificity and efficacy of drug delivery with minimal side effects [82]. The cyclic peptide, cCPGPEGAGC (PEGA), is a homing peptide that can identify cancer cells. In conjugation with the cell-penetrating peptide pVEC, this peptide was selectively taken up by different breast cancer cells [83]. Another peptide, D2A21, and its gel formulations have been used in wound-healing products to treat infected burns, wounds, and several types of cancer. A TAT peptide derived from the N-terminus of p53, fused with a peptide derived from the VHL tumor-suppressor gene, inhibits insulin-like growth factor I receptor (IGF-IR) signaling in renal cell carcinomas [83].
2.7 Emerging Issues and Developments in Proteins-Based Therapeutics
Both native and recombinant therapeutic proteins are an essential class of medicines developed to treat a wide variety of diseases. Therapeutic proteins, including vaccines, antigens, or hormones, are produced using rDNA technology and protein purification methods. The drug developers apply protein engineering to achieve desirable molecular characteristics to make these therapeutic proteins safe and effective. Drug targeting is an important aspect of therapeutics to treat several diseases. So, it is essential to devise better drug targeting and delivery methods to have improved potency and functionality. The knowledge of the mechanism of action and structure-function relationship of a protein is essential for engineering its activity or introducing new desired activities. The customization of existing proteins or the generation of novel therapeutics having specific clinical applications is a developing field in drug design. Besides protein engineering, technological advancements in genetic engineering are also used to develop therapeutic proteins to tackle a wide range of life-threatening conditions. However, there are challenges and limitations associated with the use of therapeutic proteins to combat life-threatening conditions [84], which include (i) optimal utilization of therapeutic proteins and peptides via the oral route, (ii) extensive hepatic first-pass metabolism, (iii) degradation in the gastrointestinal tract, and (iv) large molecular size and poor permeation.
2.7.1 Issue of Demand and Supply
Therapeutic protein development projects are time consuming and budget extensive. Also, the associated development processes have various intricacies of cellular metabolism, pharmacokinetics, and pharmacodynamics, making their development task more difficult. To reduce the complexity and overcome related limitations, pharmaceutical scientists do a lot of preresearch and testing to select only those molecules with a maximum chance of success in clinical trials. It is a well-known and documented fact that from discovery to the pharmaceutical market, a new drug molecule takes more than ten years, and yet, its success rate is not guaranteed [17, 85]. Therefore, much research needs to be done to consider selective molecules to maximize the chances of success.
2.7.2 Issues Related to Immunogenicity
Immunogenicity is a challenging aspect of disease management. A particulate matter, however small in amount, can lead drastically enhance immunogenicity [86, 87]. Protein-based therapeutics develop either from nonhuman or human sources and have chances of producing neutralizing and/or nonneutralizing antibodies [88]. In some cases, using protein therapeutics may also lead to an array of adverse immune reactions from mild inflammation to severe anaphylaxis. The therapeutic proteins given to patients may also neutralize endogenous proteins in some cases and, thus, lead to adverse effects [89]. T cells are a critical arm of the immune system, and their activity is regulated via T cell receptor interactions. Therefore, the prior knowledge of all the T cell epitopes present on the surface of therapeutics can enhance the immune tolerance level and, thus, minimize the unwanted immunogenic responses [90].
The immunogenicity of protein therapeutics can be reduced by designing depleting T cell epitopes (deimmunization process) [91]. This idea led to many deimmunized therapeutic proteins in clinical trials. The quality and quantity of T cell epitopes are measured by using T-cell-based assays. These in vitro methods, along with computational techniques, facilitate the identification and removal of T cell epitopes. The desired mutations can be incorporated into the peptide sequences using in silico tools, which are later implemented into deimmunized T cell epitope protein sequences. The resultant peptides have limited capacity for MHC binding and produce decreased immune responses. Antibodies are mainly deimmunized protein therapeutics. The rise in the unwanted level of immunogenicity is diminished by using deimmunized antibodies. The deimmunized antibodies specific for prostrate membrane antigen have passed different stages of clinical trials and are approved for clinical use. In clinical trials, these antibodies are conjugated with a radioactive probe and do not show antitherapeutic immune responses [92,93,94,95]. So, T cell epitopes are one of the crucial factors taken into consideration to control antitherapeutic antibody responses. Thus, deimmunized protein-based therapeutics may provide a safe class of new biologics.
2.7.3 Issue of Protein Stability
The prolonged stability of therapeutic proteins in a clinical setting is a desirable but challenging trait. It is one of the limitations in making them ideal clinical therapeutics [96]. The possible aggregation of therapeutic proteins increases if they are stored in the high concentrations required for using them on a large-scale [97, 98]. The aggregation decreases their overall activity and results in immunological reactions [97]. This problem can be overcome by spatial aggregation propensity, which identifies aggregation-prone regions in a protein sequence, and then those regions are mutated to engineer stable antibodies [99, 100]. High temperature also influences the stability of therapeutic proteins. The proteins lose their activity and structural integrity when stored at room temperature. The best practice is to keep the purified therapeutic proteins at or below 4 °C for an extended period. During the release of therapeutic proteins, it may also form particles that compromise their stability and induce an immune response in patients when administered [97]. The common strategies to enhance protein stability include (i) the inclusion of desired mutations in the protein [101, 102], (ii) optimization of the formulation of therapeutic proteins [103], (iii) use of thermosensitive polymers [96], (iv) encapsulation of therapeutic proteins, (v) use of biodegradable polymers for the delivery of therapeutic substances, and (vi) use of nontoxic nanostructured materials.
2.7.4 Issues of Metabolism and Elimination
The metabolism and elimination of protein-based therapeutics, such as those used for hepatic diseases (like liver cirrhosis), poses a significant hurdle in their successful clinical uses. The noninvasive administration of protein-based therapeutic by using alternative routes can possibly solve the issue of hepatic metabolism. Moreover, the hepatic first-pass metabolism may also be overcome by using invasive delivery of therapeutic proteins. Another problem with therapeutic proteins is that most have a short half-life. To overcome this issue, therapeutic proteins are encapsulated and/or conjugated with biocompatible polymers [104]. Nowadays, the half-life of these therapeutics is enhanced by using existing fusion protein technology.
The hurdles in developing and delivering therapeutic proteins may be overcome by studying their pharmacokinetic properties and pharmacodynamic effects. Complete identification and analysis of their pharmacokinetic parameters are required for predicting the biodisposition of these agents. Though recent advancements in applied technologies have solved such problems to some extent, some unknown factors are responsible for creating hindrances to efficiently using much-needed therapeutics. For instance, poor intestinal absorption and intestinal first-pass metabolism significantly impact the clearance of protein-based therapeutics if they are given via the oral route. Therefore, in-depth knowledge of the routes of their administration and the underlying mechanism of metabolism is needed to tackle the issues of early clearance. Several such protein-based therapeutics have recently been developed, such as Oral Recosulin, Octreolin®, Sandimmune®, etc., and many are in the clinical stages of development.
2.8 Conclusion and Future Prospects
Protein-based therapeutics are engineered drugs with a wide range of clinical applications. Rapid progress has been made in the last decade toward developing engineered proteins to treat several life-threatening conditions. Clinical safety and efficacy are some of the essential features to overcome by working on various factors such as disease biology, individual genetic profile, and selection of target population on the patient side, while safety margin, route of delivery, half-life, stability, and solubility on the protein-based therapeutics side. The mode of drug delivery is a concern for increasing the efficacy. The oral delivery of therapeutic proteins is one of the efficient ways to replace the invasive routes, only if the problems of poor absorption and intestinal first-pass metabolism are handled. The recent advancements in several cross-cutting technologies have made the oral delivery of therapeutic proteins possible. Apart from that, the stability of protein during its formulation and decreasing its development cost remain a significant challenge in front of research communities. These problems can be handled by parallel use of advanced in vitro, in vivo, and in silico techniques. Many protein-based therapeutics are either FDA approved or in the final stages of approval, with many reaching the global market, and hundreds are in preclinical studies and clinical development. These therapeutics have been successful in treating a variety of conditions, from diabetes mellitus to cancers. The design and development of therapeutic proteins considering novel scaffolds with superior biochemical and physiological activities will be primary areas of research in the coming decades.
References
Singh DB, Tripathi T (2020) Frontiers in protein structure, function, and dynamics. Springer Nature, Singapore
Sumant OSS (2017) Protein Therapeutics Market, Market Research Report, Allied Market Research, p 211
Dimitrov DS (2012) Therapeutic proteins. Methods Mol Biol 899:1–26
Tripathi T, Dubey VK (2022) Advances in protein molecular and structural biology methods, 1st edn. Academic Press, Cambridge
Heckmann CM, Paradisi F (2020) Looking back: a short history of the discovery of enzymes and how they became powerful chemical tools. ChemCatChem 12(24):6082–6102
Hemalatha T, UmaMaheswari T, Krithiga G, Sankaranarayanan P, Puvanakrishnan R (2013) Enzymes in clinical medicine: an overview. Indian J Exp Biol 51(10):777–788
Robinson PK (2015) Enzymes: principles and biotechnological applications. Essays Biochem 59:1–41
Tandon S, Sharma A, Singh S, Sharma S, Sarma SJ (2021) Therapeutic enzymes: discoveries, production and applications. J Drug Deliv Sci Technol 63:102455
Martínez Cuesta S, Rahman SA, Furnham N, Thornton JM (2015) The classification and evolution of enzyme function. Biophys J 109(6):1082–1086
Meghwanshi GK, Kaur N, Verma S, Dabi NK, Vashishtha A, Charan PD, Purohit P, Bhandari HS, Bhojak N, Kumar R (2020) Enzymes for pharmaceutical and therapeutic applications. Biotechnol Appl Biochem 67(4):586–601
Vimal A, Kumar A (2019) Chapter 35 - transforming the healthcare system through therapeutic enzymes. In: Kuddus M (ed) Enzymes in food biotechnology. Academic Press, pp 603–625
Van Stappen T, Vande Casteele N, Van Assche G, Ferrante M, Vermeire S, Gils A (2018) Clinical relevance of detecting anti-infliximab antibodies with a drug-tolerant assay: post hoc analysis of the TAXIT trial. Gut 67(5):818
Dalal SR, Cohen RD (2015) What to do when biologic agents are not working in inflammatory bowel disease patients. Gastroenterol Hepatol (N Y) 11(10):657–665
Kalita P, Tripathi T (2022) Methodological advances in the design of peptide-based vaccines. Drug Discov Today 27(5):1367–1380
Kany S, Vollrath JT, Relja B (2019) Cytokines in inflammatory disease. Int J Mol Sci 20(23):6008
Ndoja S, Lima H (2017) Monoclonal antibodies. In: Thomaz-Soccol V, Pandey A, Resende RR (eds) Current developments in biotechnology and bioengineering. Elsevier, pp 71–95
Strohl WR, Knight DM (2009) Discovery and development of biopharmaceuticals: current issues. Curr Opin Biotechnol 20(6):668–672
Scully M, Cataland SR, Peyvandi F, Coppo P, Knöbl P, Kremer Hovinga JA, Metjian A, de la Rubia J, Pavenski K, Callewaert F, Biswas D, De Winter H, Zeldin RK (2019) Caplacizumab treatment for acquired thrombotic thrombocytopenic purpura. N Engl J Med 380(4):335–346
Campochiaro PA, Heier JS, Feiner L, Gray S, Saroj N, Rundle AC, Murahashi WY, Rubio RG (2010) Ranibizumab for macular edema following branch retinal vein occlusion: six-month primary end point results of a phase III study. Ophthalmology 117(6):1102–1112.e1
Tcheng JE, Kandzari DE, Grines CL, Cox DA, Effron MB, Garcia E, Griffin JJ, Guagliumi G, Stuckey T, Turco M, Fahy M, Lansky AJ, Mehran R, Stone GW (2003) Benefits and risks of abciximab use in primary angioplasty for acute myocardial infarction: the controlled abciximab and device investigation to lower late angioplasty complications (CADILLAC) trial. Circulation 108(11):1316–1323
Blair HA (2019) Emicizumab: a review in Haemophilia A. Drugs 79(15):1697–1707
Burt R, Warcel D, Fielding AK (2019) Blinatumomab, a bispecific B-cell and T-cell engaging antibody, in the treatment of B-cell malignancies. Hum Vaccin Immunother 15(3):594–602
Linke R, Klein A, Seimetz D (2010) Catumaxomab: clinical development and future directions. MAbs 2(2):129–136
Strohl WR, Strohl LM (2012) Introduction to biologics and monoclonal antibodies, therapeutic antibody engineering. Woodhead Publishing, pp 1–595
Pradhananga S, Sayers JR (2012) Natural synthesis: biologics, biosimilars and biobetters in protein hormone therapy. Biochemist 34(1):10–15
Kowarski CR, Lin SS, Michalek LM, Kowarski AA (1988) Comparing the biological activity of humulin R and novolin R using an intravenous bioassay. Pharm Res 5(4):245–246
Latrech H, Simon A, Beltrand J, Souberbielle JC, Belmejdoub G, Polak M (2012) Postprandial hyperglycemia corrected by IGF-I (Increlex®) in Laron syndrome. Horm Res Paediatr 78(3):193–200
Lal RA, Hoffman AR (2018) Long-acting growth hormone preparations in the treatment of children. Pediatr Endocrinol Rev 16(Suppl 1):162–167
de Las Cuevas Allende R, Díaz de Entresotos L, Conde Díez S (2021) Anaemia of chronic diseases: pathophysiology, diagnosis and treatment. Med Clin (Barc) 156(5):235–242
Dale DC, Crawford J, Klippel Z, Reiner M, Osslund T, Fan E, Morrow PK, Allcott K, Lyman GH (2018) A systematic literature review of the efficacy, effectiveness, and safety of filgrastim. Support Care Cancer 26(1):7–20
Swartz ML (1991) Intron A (interferon-alpha-2b recombinant) for injection/Schering-Plough Corporation. Gastroenterol Nurs 14(1):40–43
Kleinschnitz C, Niemczyk G, Rehberg-Weber K, Wernsdörfer C (2015) Interferon Beta-1a (AVONEX®) as a treatment option for untreated patients with multiple sclerosis (AXIOM): a prospective, observational study. Int J Mol Sci 16(7):15271–15286
Hadjigeorgiou G, Dardiotis E, Tsivgoulis G, Doskas T, Petrou D, Makris N, Vlaikidis N, Thomaidis T, Kyritsis A, Fakas N, Treska X, Karageorgiou C, Sotirli S, Giannoulis C, Papadimitriou D, Mylonas I, Kouremenos E, Vlachos G, Georgiopoulos D, Mademtzoglou D, Vikelis M, Zintzaras E (2014) Observational study assessing demographic, economic and clinical factors associated with access and utilization of health care services of patients with multiple sclerosis under treatment with interferon beta-1b (EXTAVIA). PLoS One 9(11):e113933
Clark JI, Wong MKK, Kaufman HL, Daniels GA, Morse MA, McDermott DF, Agarwala SS, Lewis LD, Stewart JH, Vaishampayan U, Curti B, Gonzalez R, Lutzky J, Rudraptna V, Cranmer LD, Jeter JM, Hauke RJ, Miletello G, Milhem MM, Amin A, Richart JM, Fishman M, Hallmeyer S, Patel SP, Van Veldhuizen P, Agarwal N, Taback B, Treisman JS, Ernstoff MS, Perritt JC, Hua H, Rao TB, Dutcher JP, Aung S (2017) Impact of sequencing targeted therapies with high-dose Interleukin-2 immunotherapy: an analysis of outcome and survival of patients with metastatic renal cell carcinoma from an on-going observational IL-2 clinical trial: PROCLAIM(SM). Clin Genitourin Cancer 15(1):31–41.e4
Kaye JA (1998) FDA licensure of NEUMEGA to prevent severe chemotherapy-induced thrombocytopenia. Stem Cells 16(Suppl 2):207–223
Vogenberg FR, Isaacson Barash C, Pursel M (2010) Personalized medicine: part 1: evolution and development into theranostics. P T 35(10):560–576
Ginsburg GS, Phillips KA (2018) Precision medicine: from science to value. Health Aff (Millwood) 37(5):694–701
Anna M, Nurrani Mustika D, Andi W (2016) Personalized medicine: the future of health care. Indones Biomed J 8(3):46–127
Mathur S, Sutton J (2017) Personalized medicine could transform healthcare. Biomed Rep 7(1):3–5
Gameiro GR, Sinkunas V, Liguori GR, Auler-Júnior JOC (2018) Precision medicine: changing the way we think about healthcare. Clinics (Sao Paulo) 73:e723
Bradford D, Demko S, Jin S, Mishra-Kalyani P, Beckles AR, Goldberg KB, Lemery S, Ward A, Keegan P, Pazdur R (2020) FDA accelerated approval of pembrolizumab for recurrent locally advanced or metastatic Merkel cell carcinoma. Oncologist 25(7):e1077–e1082
Miller EM, Schwartzberg LS (2019) Biosimilars for breast cancer: a review of HER2-targeted antibodies in the United States. Ther Adv Med Oncol 11:1758835919887044–1758835919887044
Bilkey GA, Burns BL, Coles EP, Mahede T, Baynam G, Nowak KJ (2019) Optimizing precision medicine for public health. Front Public Health 7:42
Schmidt C (2014) Leroy Hood looks forward to P4 medicine: predictive, personalized, preventive, and participatory. J Natl Cancer Inst 106(12)
Hay M, Thomas DW, Craighead JL, Economides C, Rosenthal J (2014) Clinical development success rates for investigational drugs. Nat Biotechnol 32(1):40–51
Mullard A (2016) Biotech R&D spend jumps by more than 15. Nat Rev Drug Discov 15(7):447
Song CM, Lim SJ, Tong JC (2009) Recent advances in computer-aided drug design. Brief Bioinform 10(5):579–591
Lionta E, Spyrou G, Vassilatis DK, Cournia Z (2014) Structure-based virtual screening for drug discovery: principles, applications and recent advances. Curr Top Med Chem 14(16):1923–1938
Wlodawer A, Vondrasek J (1998) Inhibitors of HIV-1 protease: a major success of structure-assisted drug design. Annu Rev Biophys Biomol Struct 27:249–284
Güner O, Clement O, Kurogi Y (2004) Pharmacophore modeling and three dimensional database searching for drug design using catalyst: recent advances. Curr Med Chem 11(22):2991–3005
Yu W, MacKerell AD Jr (2017) Computer-aided drug design methods. Methods Mol Biol 1520:85–106
Sharma V, Wakode S, Kumar H (2021) Chapter 2 - Structure- and ligand-based drug design: concepts, approaches, and challenges. In: Sharma N, Ojha H, Raghav PK, Goyal RK (eds) Chemoinformatics and bioinformatics in the pharmaceutical sciences. Academic Press, pp 27–53
Verma J, Khedkar VM, Coutinho EC (2010) 3D-QSAR in drug design--a review. Curr Top Med Chem 10(1):95–115
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26(5):694–701
Dhanavade MJ, Jalkute CB, Barage SH, Sonawane KD (2013) Homology modeling, molecular docking and MD simulation studies to investigate role of cysteine protease from Xanthomonas campestris in degradation of Aβ peptide. Comput Biol Med 43(12):2063–2070
Tripathi T, Kalita P (2019) Synergistic effect of amyloid-β and tau disrupts neural circuits. ACS Chem Neurosci 10(3):1129–1130
Tripathi T, Khan H (2020) Direct interaction between the β-amyloid core and tau facilitates cross-seeding: a novel target for therapeutic intervention. Biochemistry 59(4):341–342
Labandeira-Garcia JL, Rodríguez-Perez AI, Villar-Cheda B, Borrajo A, Dominguez-Meijide A, Guerra MJ (2015) Rho kinase and dopaminergic degeneration: a promising therapeutic target for Parkinson's disease. Neuroscientist 21(6):616–629
Sorce S, Krause KH (2009) NOX enzymes in the central nervous system: from signaling to disease. Antioxid Redox Signal 11(10):2481–2504
Alokam R, Singhal S, Srivathsav GS, Garigipati S, Puppala S, Sriram D, Perumal Y (2015) Design of dual inhibitors of ROCK-I and NOX2 as potential leads for the treatment of neuroinflammation associated with various neurological diseases including autism spectrum disorder. Mol BioSyst 11(2):607–617
Padhi AK, Rath SL, Tripathi T (2021) Accelerating COVID-19 research using molecular dynamics simulation. J Phys Chem B 125(32):9078–9091
Mohan A, Rendine N, Mohammed MKS, Jeeva A, Ji HF, Talluri VR (2021) Structure-based virtual screening, in silico docking, ADME properties prediction and molecular dynamics studies for the identification of potential inhibitors against SARS-CoV-2 M(pro). Mol Divers:1–17
Yadav MK, Ahmad S, Raza K, Kumar S, Eswaran M, Pasha Km M (2022) Predictive modeling and therapeutic repurposing of natural compounds against the receptor-binding domain of SARS-CoV-2. J Biomol Struct Dyn:1–13
Shukla R, Tripathi T (2021) Molecular dynamics simulation in drug discovery: opportunities and challenges. In: Singh SK (ed) Innovations and implementations of drug discovery strategies in rational drug design. Springer Nature, Singapore, pp 295–316
Shukla R, Tripathi T (2020) Molecular dynamics simulation of protein and protein-ligand complexes. In: Singh DB (ed) Computer-aided drug design. Springer Nature, Singapore, pp 133–161
Sauerborn M, Brinks V, Jiskoot W, Schellekens H (2010) Immunological mechanism underlying the immune response to recombinant human protein therapeutics. Trends Pharmacol Sci 31(2):53–59
Greenfield JR, Tuthill A, Soos MA, Semple RK, Halsall DJ, Chaudhry A, O’Rahilly S (2009) Severe insulin resistance due to anti-insulin antibodies: response to plasma exchange and immunosuppressive therapy. Diabet Med 26(1):79–82
Radermecker RP, Renard E, Scheen AJ (2009) Circulating insulin antibodies: influence of continuous subcutaneous or intraperitoneal insulin infusion, and impact on glucose control. Diabetes Metab Res Rev 25(6):491–501
Jaeger C, Winter S, Eckhard M, Hardt P, Brendel MD, Bretzel RG (2008) Binding characteristics and crossreactivity of insulin autoantibodies and insulin antibodies directed to three different insulin molecules. Acta Diabetol 45(3):191–194
Fineberg SE, Kawabata TT, Finco-Kent D, Fountaine RJ, Finch GL, Krasner AS (2007) Immunological responses to exogenous insulin. Endocr Rev 28(6):625–652
Kaspar AA, Reichert JM (2013) Future directions for peptide therapeutics development. Drug Discov Today 18(17–18):807–817
Keservani RK, Sharma AK, Jarouliya U (2015) Protein and peptide in drug targeting and its therapeutic approach. Ars Pharmaceutica (Internet) 56:165–177
Nauck MA, Meier JJ (2005) Glucagon-like peptide 1 and its derivatives in the treatment of diabetes. Regul Pept 128(2):135–148
Luppi P, Cifarelli V, Wahren J (2011) C-peptide and long-term complications of diabetes. Pediatr Diabetes 12(3 Pt 2):276–292
Bermel RA, Rudick RA (2007) Interferon-beta treatment for multiple sclerosis. Neurotherapeutics 4(4):633–646
Farrell R, Kapoor R, Leary S, Rudge P, Thompson A, Miller D, Giovannoni G (2008) Neutralizing anti-interferon beta antibodies are associated with reduced side effects and delayed impact on efficacy of interferon-beta. Mult Scler 14(2):212–218
Kagawa Y, Takasaki S, Utsumi J, Hosoi K, Shimizu H, Kochibe N, Kobata A (1988) Comparative study of the asparagine-linked sugar chains of natural human interferon-beta 1 and recombinant human interferon-beta 1 produced by three different mammalian cells. J Biol Chem 263(33):17508–17515
Bertolotto A, Deisenhammer F, Gallo P, Sölberg Sørensen P (2004) Immunogenicity of interferon beta: differences among products. J Neurol 251:ii15–ii24
Hermeling S, Jiskoot W, Crommelin D, Bornaes C, Schellekens H (2005) Development of a transgenic mouse model immune tolerant for human interferon Beta. Pharm Res 22(6):847–851
Runkel L, Meier W, Pepinsky RB, Karpusas M, Whitty A, Kimball K, Brickelmaier M, Muldowney C, Jones W, Goelz SE (1998) Structural and functional differences between glycosylated and non-glycosylated forms of human interferon-beta (IFN-beta). Pharm Res 15(4):641–649
Bidwell GL 3rd, George EM (2014) Maternally sequestered therapeutic polypeptides - a new approach for the management of preeclampsia. Front Pharmacol 5:201–201
Zhang XX, Eden HS, Chen X (2012) Peptides in cancer nanomedicine: drug carriers, targeting ligands and protease substrates. J Control Release 159(1):2–13
Regberg J, Srimanee A, Langel U (2012) Applications of cell-penetrating peptides for tumor targeting and future cancer therapies. Pharmaceuticals (Basel) 5(9):991–1007
Muheem A, Shakeel F, Jahangir MA, Anwar M, Mallick N, Jain GK, Warsi MH, Ahmad FJ (2016) A review on the strategies for oral delivery of proteins and peptides and their clinical perspectives. Saudi Pharm J 24(4):413–428
Shih H (2013) Discovery process for antibody-based therapeutics, development of antibody-based therapeutics. Translat Consider 9–32
Carpenter JF, Randolph TW, Jiskoot W, Crommelin DJ, Middaugh CR, Winter G, Fan YX, Kirshner S, Verthelyi D, Kozlowski S, Clouse KA, Swann PG, Rosenberg A, Cherney B (2009) Overlooking subvisible particles in therapeutic protein products: gaps that may compromise product quality. J Pharm Sci 98(4):1201–1205
Rosenberg AS (2006) Effects of protein aggregates: an immunologic perspective. AAPS J 8(3):E501–E507
Anjum R, Zahra N, Rehman K, Alam R, Parveen A, Tariq M, Akash MSH (2013) Comparative analysis of serum lipid profile between normotensive and hypertensive. J Mol Genetic Med 7:64
Schellekens H (2005) Factors influencing the immunogenicity of therapeutic proteins. Nephrol Dial Transplant 20(Suppl 6):vi3–vi9
De Groot AS, Moise L, McMurry JA, Wambre E, Van Overtvelt L, Moingeon P, Scott DW, Martin W (2008) Activation of natural regulatory T cells by IgG Fc-derived peptide “Tregitopes”. Blood 112(8):3303–3311
Baker MP, Reynolds HM, Lumicisi B, Bryson CJ (2010) Immunogenicity of protein therapeutics: the key causes, consequences and challenges. Self Nonself 1(4):314–322
Galsky MD, Eisenberger M, Moore-Cooper S, Kelly WK, Slovin SF, DeLaCruz A, Lee Y, Webb IJ, Scher HI (2008) Phase I trial of the prostate-specific membrane antigen-directed immunoconjugate MLN2704 in patients with progressive metastatic castration-resistant prostate cancer. J Clin Oncol 26(13):2147–2154
Morris MJ, Divgi CR, Pandit-Taskar N, Batraki M, Warren N, Nacca A, Smith-Jones P, Schwartz L, Kelly WK, Slovin S, Solit D, Halpern J, Delacruz A, Curley T, Finn R, O’Donoghue JA, Livingston P, Larson S, Scher HI (2005) Pilot trial of unlabeled and indium-111-labeled anti-prostate-specific membrane antigen antibody J591 for castrate metastatic prostate cancer. Clin Cancer Res 11(20):7454–7461
Pandit-Taskar N, O'Donoghue JA, Morris MJ, Wills EA, Schwartz LH, Gonen M, Scher HI, Larson SM, Divgi CR (2008) Antibody mass escalation study in patients with castration-resistant prostate cancer using 111In-J591: lesion detectability and dosimetric projections for 90Y radioimmunotherapy. J Nucl Med 49(7):1066–1074
Vallabhajosula S, Goldsmith SJ, Kostakoglu L, Milowsky MI, Nanus DM, Bander NH (2005) Radioimmunotherapy of prostate cancer using 90Y- and 177Lu-labeled J591 monoclonal antibodies: effect of multiple treatments on myelotoxicity. Clin Cancer Res 11(19 Pt 2):7195s–7200s
Akash MS, Rehman K, Chen S (2013) IL-1Ra and its delivery strategies: inserting the association in perspective. Pharm Res 30(11):2951–2966
Jiskoot W, van Schie RMF, Carstens MG, Schellekens H (2009) Immunological risk of injectable drug delivery systems. Pharm Res 26(6):1303–1314
Shire SJ, Shahrokh Z, Liu J (2004) Challenges in the development of high protein concentration formulations. J Pharm Sci 93(6):1390–1402
Chennamsetty N, Voynov V, Kayser V, Helk B, Trout BL (2009) Design of therapeutic proteins with enhanced stability. Proc Natl Acad Sci U S A 106(29):11937–11942
Voynov V, Chennamsetty N, Kayser V, Helk B, Trout BL (2009) Predictive tools for stabilization of therapeutic proteins. MAbs 1(6):580–582
Lawrence T, Fong C (2010) The resolution of inflammation: anti-inflammatory roles for NF-kappaB. Int J Biochem Cell Biol 42(4):519–523
Strickler SS, Gribenko AV, Gribenko AV, Keiffer TR, Tomlinson J, Reihle T, Loladze VV, Makhatadze GI (2006) Protein stability and surface electrostatics: a charged relationship. Biochemistry 45(9):2761–2766
Schneider CP, Trout BL (2009) Investigation of cosolute-protein preferential interaction coefficients: new insight into the mechanism by which arginine inhibits aggregation. J Phys Chem B 113(7):2050–2058
Tariq M, Akash MSH, Rehman K, Shuqing C (2015) Development of therapeutic proteins: advances and challenges. Turk J Biol 39
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Yadav, M.K. et al. (2023). Clinical Applications of Protein-Based Therapeutics. In: Singh, D.B., Tripathi, T. (eds) Protein-based Therapeutics. Springer, Singapore. https://doi.org/10.1007/978-981-19-8249-1_2
Download citation
DOI: https://doi.org/10.1007/978-981-19-8249-1_2
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-8248-4
Online ISBN: 978-981-19-8249-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)