Abstract
With the exponential increase in the world’s human population, improving agricultural productivity is among the top of the researchers’ agendas till the 2050 deadline. One of the potential solutions to this global issue is genome editing because of the precision, fastness, and probably low cost involved compared to other traditional methods. It is in the spotlight especially from the last decade due to the discovery of sequence-specific-based nuclease technology including CRISPR/Cas9 tool. Initially, this tool was applied only in protoplasts and calli. However, due to the modifications in vectors, Cas9 variants, cassettes, cloning systems, multiplexing, and delivery methods, this platform has revolutionized the plant science field. It has been exploited in such a manner that about 16 crop plants have been already edited in the last few years. Out of all crops, most of the editing has been done in the case of rice (Oryza sativa L., Family: Poaceae), a cereal staple food. Therefore, in the current chapter, we have highlighted about the CRISPR/Cas9-edited rice for agronomic traits, stress tolerance/resistance, and biofortification. Additionally, we have presented an overview of various tools, databases, and commercial service providers devoted solely to CRISPR/Cas9 genome-editing technology.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
In today’s world, the human population is increasing exponentially and is expected to cross the whopping mark of 9.7 billion by the year 2050 (Valin et al. 2014; Baltes et al. 2017; Figueroa 2019). Furthermore, the whole scenario is expected to be affected greatly by the need to generate more space, reduce the overexploitation of natural resources, and tackle the uncertainties of climatic conditions and global warming (Cazzolla Gatti 2016; Oldeman et al. 2017; Morton et al. 2017; Subramanian 2018; Philander 2018; Pradinaud et al. 2019). In addition to this challenges, international food security, fighting chronic malnourishment, increasing awareness, and interest for healthier functional foods are at the top of the agendas (Siro et al. 2008; Abuajah et al. 2015; Martirosyan and Singh 2015; Atkins and Bowler 2016; Baltes et al. 2017; Pratim Roy 2019).
As our contemporary agricultural lands are degrading, it necessitates to re-think about the current agricultural practices, generation of elite varieties as well as efficient distribution of food (Wingeyer et al. 2015; Morton et al. 2017; Glenn et al. 2017; Banasik et al. 2017; Zhang et al. 2018; Dillard 2019). Solutions to all these challenges are unlikely to come from cross-breeding and mutation breeding (Kantar et al. 2019; Belkhodja 2018; Chen et al. 2019; Kleter et al. 2019; Mehta et al. 2019a; Singh et al. 2019; Rahman et al. 2019). Cross-breeding takes a large span of years to introduce desirable alleles (Darwin 2010; Scheben et al. 2017). Furthermore, this is limited by greatly reduced genetic variability. On the other hand, mutation breeding usually employs agents like ethyl methanesulfonate (EMS) and gamma rays to expand genetic variation by introducing random mutations (Bado et al. 2015, 2017; Pacher and Puchta 2017; Xuan et al. 2019). However, it is restricted by the large-scale mutant screening, high randomness, low efficiency, and stochastic nature. Furthermore, these approaches cannot keep pace with the whopping demand for increased crop production.
As a result, one of the potent approaches that can withstand the increasing crop productivity is genetic engineering (Marco et al. 2015; Baret and Vanloqueren 2017; Knott and Doudna 2018). It has been the spotlight around the globe to create new crop varieties (Sticklen 2008; Marco et al. 2015; Azadi et al. 2016; Arzani and Ashraf 2016; Kumari et al. 2018; Waltz 2018; Banerjee and Roychoudhury 2019; Zhang et al. 2019). Generally, it is defined as the targeted modification of DNA of any living organism belonging to any kingdom of classification using various tools (Baltes et al. 2017). In accordance with the current and future scenario challenges, it easily addresses questions like (1) which traits need to be introduced, (2) which crops need to be focused on, (3) which DNA modifications must be done to generate the desired traits in the selected crops, (4) how to introduce these DNA modifications in the crop’s genome, (5) how to overcome the bottlenecks of existing tools for crop improvement particularly, and (6) how to shift the agendas in accordance with the changing challenges. Due to the wide-ranging use, the enormous number of application falls under the big umbrella of genome engineering. As a result there is a wide range of potential products that could address food security/quality issues (Hsu et al. 2014; Wu et al. 2016; Nielsen and Keasling 2016; Khalid et al. 2017; Knott and Doudna 2018; Shigaki 2018; Waltz 2018; Pray et al. 2018; Merga et al. 2019; Zhang 2019).
Nonetheless, one of the significant tools that has been used enormously in agriculture is genome editing (Upadhyay et al. 2013; Laible et al. 2015; Alagoz et al. 2016; Ricroch et al. 2017; Gao 2018; Eş et al. 2019; Lassoued et al. 2019; Yin and Qiu 2019). This is truly reflected in numerous improved cultivars which have emerged within the last decade (Laible et al. 2015; Alagoz et al. 2016; Yin et al. 2017; Gao 2018; Yin and Qiu 2019). Here, we have highlighted different types of genome-editing tools for plants. Additionally, we have focused on the CRISPR/Cas9-edited rice for various traits and the current limitations and challenges within this field.
2 Genome-Editing Techniques for Plants
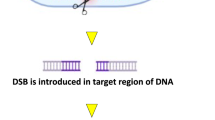
Perhaps the availability of numerous tools for DNA/RNA modifications, the sequence-specific nucleases have been the spotlight for the entire last decade (Porteus and Carroll 2005; Wright et al. 2005; Christian et al. 2010; Voytas 2013; Sprink et al. 2015; Zischewski et al. 2017; Waltz 2018; Novak 2019). These nucleases introduce targeted DNA double-strand breaks (DSBs) which are repaired by the cells itself by two evolved pathways, i.e., homologous recombination (HR) and nonhomologous end joining (NHEJ) (Puchta and Fauser 2014). However, in comparison, NHEJ is naturally an error-prone pathway which frequently results in small indels at the repair sites. Therefore, the researchers utilize this for targeted mutagenesis at a locus of interest. This NHEJ pathway exists in somatic, meiotic, and mitotic cells throughout the cell cycle, whereas HR-mediated repair pathway occurs only within the G2 and S phases of cells having mitotic activity (Huang and Puchta 2019; Jun et al. 2019).
In the present scenario, mostly genome editing is done by multiple technologies like meganucleases (Certo et al. 2012; Daboussi et al. 2015; Youssef et al. 2018), zinc-finger nucleases (ZFNs) (Porteus and Carroll 2005; Wright et al. 2005; Bilichak and Eudes 2016; Novak 2019), TALENs (Christian et al. 2010; Bilichak and Eudes 2016; Hensel and Kumlehn 2019), and CRISPR/Cas9 systems in plants (Bilichak and Eudes 2016; Knott and Doudna 2018; Waltz 2018; Huang and Puchta 2019). A detailed comparison of all these editing technologies is tabulated in Table 23.1. For more detailed information, the readers can look for publications by Gaj et al. (2013), Puchta and Fauser (2014), Sprink et al. (2015), Bilichak and Eudes (2016), Noman et al. (2016), Baltes and Voytas (2015), Baltes et al. (2017), Malzahn et al. (2017), Kamburova et al. (2017), Lino et al. (2018), Shah et al. (2018), and Novak (2019).
3 CRISPR/Cas9 System for Fathomless Genetic Engineering
Currently, the most popular genetic cargo technology is CRISPR/Cas9 (Shan et al. 2013; Belhaj et al. 2013; Miao et al. 2013). This system has truly revolutionized the plant science research (Bilichak and Eudes 2016; Knott and Doudna 2018; Waltz 2018; Huang and Puchta 2019). As a result, various articles have been published throughout the last few years (Belhaj et al. 2013; Shan et al. 2014; Gao et al. 2015; Bilichak and Eudes 2016; Liu et al. 2017a, b; Liang et al. 2017; Knott and Doudna 2018; Butt et al. 2018; Abbott and Qi 2018; Huang and Puchta 2019). This is even supported by the fact that the keyword “CRISPR/Cas” in the paper title fetched about 5610 publications in Google Scholar (https://scholar.google.co.in/).
This CRISPR/Cas9 tool is popular due to the advantages such as simplicity, easy design, and easiness in delivery (Upadhyay et al. 2013; Baltes et al. 2017; Langner et al. 2018; Soda et al. 2018; Chen et al. 2019). CRISPR/Cas9 stand for clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated endonuclease 9 (Cas9) (Shan et al. 2013; Baltes et al. 2017). Both are integral components of the adaptive immunity system present within bacteria and archaea for protection against bacteriophages (Horvath and Barrangou 2010; Bondy-Denomy et al. 2013; Sampson et al. 2013; Shan et al. 2014). Based on this immunity mechanism, the CRISPR/Cas9 plant transformation vectors have been designed which carries guide RNA (gRNA) and Cas9 (Cong et al. 2013). In the initial days, it was applied in protoplast, calli, germ cells, and somatic cells (Shan et al. 2013; Feng et al. 2013, 2014a, b; Shen et al. 2014; Xing et al. 2014; Yin et al. 2015; Bhowmik et al. 2018). Until now, various modifications have been done in CRISPR/Cas9 plant vectors (Shen et al. 2014; Ma et al. 2015; Mikami et al. 2015a, b; Osakabe et al. 2016; Tsutsui and Higashiyama 2017; Wang et al. 2018; Wu et al. 2018; Mahas et al. 2019). This is even supported by the fact that the optimized protocols are available for many plant species (Miao et al. 2013; Xing et al. 2014; Lowder et al. 2015; Char et al. 2017; Bhowmik et al. 2018; Osakabe et al. 2018; Li and Zhang 2019). Additionally, there is a plethora of available tools and databases devoted to the various omics technologies (Anamika et al. 2019) as well as CRISPR/Cas9 (Tables 23.2 and 23.3). Furthermore, there are many commercial service providers in the marketplace which provide many services and products related to the CRISPR/Cas9 technology (Table 23.4).
Furthermore, this CRISPR/Cas9-mediated genome-editing tool has been successfully implied in various plants (Cong et al. 2013; Upadhyay et al. 2013; Feng et al. 2014a, b; Shan et al. 2014; Svitashev et al. 2015; Malnoy et al. 2016; Alagoz et al. 2016; Liu et al. 2017a, b; Soda et al. 2018). For more detailed information, the researchers are advised to look for publication from the Korotkova and group (Korotkova et al. 2017, 2019).
Recently, Korotkova and colleagues published a cataloging article entitled “Current achievements in modifying crop genes using CRISPR/Cas system” (Korotkova et al. 2019). They studied all the published research articles on crop genome modifications from the Scopus database. In their article, they reported CRISPR/Cas-based genome-editing technology has been applied largely to the rice. The probable reason is being an established model plant which simultaneously counted as the highly valued cash crop worldwide (Khush 2005). This is even boosted by the availability of the rice genome sequence, sequence maps and multiple databases (Goff et al. 2002; Yu et al. 2002; Project, I.R.G.S. and Sasaki 2005; Smita et al. 2011; Zhao et al. 2014; Copetti et al. 2015; Zhang et al. 2016; Crossa et al. 2017). This is even supported by the surge in the number of publications related to the CRISPR/Cas9 (Fig. 23.1). Figure 23.2 highlights the key developments in the field of CRISPR/Cas9 technology for rice.
Typically, the CRISPR/Cas9 system success in rice relies mostly on two factors: (1) type of plant transformation vector and (2) the used delivery system. In general, the vectors carry essentially Cas9 (a endonuclease/nickase), T-DNA border region, selectable marker genes (plant and bacterial), ori site, and gRNA(s) (Alok et al. 2018) depending on the type of strategy-employed binary system, co-transformation, and/or multiplexing (Fig. 23.3). For more detailed information about the CRISPR/Cas9 vector components, the readers can look for publication by Alok et al. (2018).
Similarly, the CRISPR/Cas9-editing reagents (DNA/RNa,RNPs) are delivered into plant cells by particle bombardment (Shan et al. 2014; Sun et al. 2016; Li et al. 2016a, b, c, 2019), Agrobacterium-mediated transformation (Shan et al. 2013, 2014; Xu et al. 2014; Hu et al. 2016; Lu and Zhu 2017; Wang et al. 2019), or protoplast transfection (Xie and Yang 2013; Tang et al. 2019; Lin et al. 2018). The overall workflow for rice genome editing using CRISPR/Cas9 is depicted in Fig. 23.3.
4 CRISPR/Cas9 in Rice for Increasing Food Production
One way to address the global food demand is to increase the crop yield. However, it affects various factors including selection of high-yielding/stress-tolerant cultivars, modification of existing cultivars, nutrient supply, water supply, and weed–pest management. In the past 5 years, the use of genome editing was in its infancy; however, there are numerous successful reports currently in the literature.
4.1 Agronomic Traits Improvement
The most common way to improve the overall yield is to increase the grain number, weight, and size (Sakamoto and Matsuoka 2008; Xing and Zhang 2010; Baltes et al. 2017). Genetically the underlying grain number, weight, and size are directly linked with hundreds of genes and quantitative trait loci. Various major genes/QTLs have been molecularly characterized and edited using the CRISPR/Cas9 system in rice. Gene editing through CRISPR/Cas9 in rice cultivar Zhonghua for loss of function mutation in genes for grain number (Gn1a), grain size (GS3), panicle architecture (DEP1), and plant architecture (IPA1). Mutated rice plants exhibit higher grain number, larger grain size, and dense panicle, and ipa1 mutant shows lesser as well as higher panicle number depending on mutation in the target site of miR156 (Li et al. 2016a, b, c). Knockout mutation in Japonica rice, Kitaake cultivar for LAZY1 gene, exhibits higher tiller number (Miao et al. 2013).OsCAld5H1 gene knockout by CRISPR/Cas9 in rice leads to enrichment of G units in lignins and reveals its role in the synthesis of non-c-p-coumaroylated S lignin units (Takeda et al. 2019). Mutation in abscisic acid receptor family of genes, PYLs through CRISPR/Cas9 in rice, leads to improved growth and enhanced productivity (Miao et al. 2018). Grain weight in rice is regulated by GW2, GW5, and TGW6. The multiple gene editing of all three genes by CRISPR/Cas9 in rice shows larger grain size as compared to non-edited rice (Xu et al. 2016). Multiplex editing of genes Hd2, Hd4, and Hd5 mediated by CRISPR/Cas9 leads to early maturity in rice (Li et al. 2017a, b). All these results together provide information regarding already edited genes in various rice cultivars for enhancing agronomic traits (Table 23.5).
4.2 Enhanced Stress Tolerance/Resistance
A major bottleneck to the current rice productivity is due to the losses incurred by pests, pathogens, and weeds. These biotic stresses are estimated to decline global agricultural productivity by 40% (Mew et al. 1993; Oerke 2006; Savary et al. 2012). In a favorable environment, blast disease causes 60–100% yield loss in rice-growing area (Kihoro et al. 2013). Blast is one of the most devastating diseases in rice caused by Magnaporthe oryzae (Zhang et al. 2014). Great efforts were made in the last few decades for developing blast-resistant rice cultivar through the application of genomics tools. Through conventional breeding approaches, blast-resistant rice has been developed (Fukuoka et al. 2014; Ashkani et al. 2015). Conventional breeding approaches are tedious in nature and need a longer duration. Other limitations like the existence of pathogen variability and the emergence of new pathotype cause breakdown of resistance barrier leading to severe disease infestation. Recent advanced technologies like CRISPR/Cas9, TALEN, and ZFNs could be alternative approaches for engineering rice genome for acquiring disease-resistant phenotype. Blast disease-resistant phenotype is reported in rice by the disruption of ethylene-responsive factor 922 (OsERF922) gene-mediated through CRISPR/Cas9 (Wang et al. 2016). Targeted mutation through CRISPR/Cas9 in the ethylene responsive factor 922 provides blast disease resistance in rice (Liu et al. 2012). Expression of OsSWEET13 gene in rice responsible for bacterial blight disease and indica rice, IR24, with improved resistance for bacterial blight disease has been developed through CRISPR/Cas9 knockout targeting promoter of OsSWEET13 (Zhou et al. 2015). By disrupting, promoter of OsSWEET14 gene by TALEN technology results in resistance towards bacterial blight in rice (Li et al. 2012). TALEN technology is employed for modifying the promoter of Os09g29100 gene to nullifying EBEtal7 interaction, which could provide tolerance to BLB disease in rice (Cai et al. 2017). TALENs targeting effector binding elements (EBEs) of AvrXa7 and Tal5 disrupt their interaction with the susceptible gene Sweet14. The edited rice plants were resistant to Xanthomonas infection. These technologies are quite helpful in developing rice cultivar with tolerant phenotype for Xoo (Li et al. 2012).
Rice production constrained by viral disease including rice tungro disease (RTD) plays a major role in reducing rice production in rice-growing areas (Azzam and Chancellor 2002; Muralidharan et al. 2003; Chancellor et al. 2006). Through the development of near-isogenic lines (NILs), it is confirmed by the researchers that resistance to RTSV and RTBV depend on the translation and in-frame mutation of initiation factor 4 gamma (eIF4G) gene respectively (Lee et al. 2010; Macovei et al. 2018). In-frame mutation in eIF4G gene in rice confers resistant phenotype for RTSV (Macovei et al. 2018).
In addition to pathogen resistance, weed management is also considered as a critical factor in optimizing the crop yield. One of the effective ways is the application of herbicides on the field. Herbicide-resistant gene, bentazon-sensitive lethal (BEL) knockout by CRISPR/Cas9, and biallelic mutated rice confer sensitivity to bentazon. This trait could be successfully utilized for hybrid seed production (Xu et al. 2014). CRISPR/Cas9-mediated gene replacement of 5-enolpyruvylshikimate-3-phosphate synthase (EPSP) having the desired substitution gives glyphosate-resistant phenotype in rice (Li et al. 2016a, b, c). Herbicide-tolerant rice cultivar is generated by mutation in the ALS gene by genome editing (Li et al. 2016a, b, c; Sun et al. 2016). TALEN technology was used for creating double-point mutation mediated through homology-directed repair (HR) in OsALS rice gene (Li et al. 2016a, b, c). Rice ALS gene also mutated at multiple points using CRISPR/Cas9 HR and edited rice plant shows tolerance to bispyribac sodium (BS) spraying, and wild-type rice died after 36 days of herbicide spray (Sun et al. 2016). A point mutation generated in acetolactate synthase (ALS) gene through CRISPR/Cas9 coupled with cytidine deaminase confers tolerance to imazamox herbicide (Shimatani et al. 2017a, b) (Table 23.6).
Next to biotic stress, abiotic stresses are considered a factor that controls the rice productivity (Mehta et al. 2019a, b). It includes flooding, drought, heavy metal stress, metalloid stress, and heat stress (Dhakate et al. 2019). Rice plant is extremely sensitive under low temperature especially during the early stage of development. Therefore, the improvement of rice varieties for cold tolerance could significantly enhance productivity in rice. For enhancing cold tolerance in rice, TIFY1b and its homology gene TIFY1a were edited through CRISPR/Cas9 (Huang et al. 2017). Osmotic stress/ABA-activated protein kinase 2 (OsSAPK2) knockout mutant-mediated by CRISPR/Cas9 exhibits higher sensitivity for drought and reactive oxygen species than control rice plant (Lou et al. 2017).
4.3 Biofortification
Next to increasing food production, improving food nutritional value is the biggest hurdle to the researchers. This demand has increased globally with the hike in household incomes and food-related awareness in developing countries. As a result, nowadays consumers require food with properties such as reduced cholesterol, biofortified whole grains, and low wax. As a result, various researchers have successfully used CRISPR/Cas9 technology for biofortification especially in rice.
Loss of function mutation through CRISPR/Cas9 of waxy gene in rice has reduced amylose content (Zhang et al. 2018). CRISPR/Cas9-mediated loss-of-function mutation of the starch debranching enzymes SBEI and SBEIIb has higher amylose content and resistant starch (Sun et al. 2017). Knockout of ISA1 gene through CRISPR/Cas9 in rice exhibit reduced amylose and amylopectin contents. The mutant seeds were altered with shrunken endosperm and lesser grain weight (Chao et al. 2019). Loss-of-function mutation of Nramp5 through CRISPR/Cas9 in rice have low cadmium content when grown in cadmium-contaminated field (Tang et al. 2017a, b). Targeted mutations through modified CRISPR/Cas9 (nCas9 containing cytidine deaminase) for OsPDS and OsSBEIIb in rice were generated (Li et al. 2017a, b). Knockout of Badh2 gene mediated by CRISPR/Cas9 in rice exhibits enhanced aroma (Shao et al. 2017). Table 23.7 summarizes the successful reports of rice biofortification.
5 Insights into the CRISPR/Cpf1: An Alternative to CRISPR/Cas9
In addition to Cas9, scientists have reported other Cas family members for genome editing in the last 5 years. One of the promising members is Cpf1 (CRISPR from Prevotella and Francisella1) (Zetsche et al. 2015). The mechanisms of CRISPR/Cpf1 and CRISPR/Cas9 are compared in Table 23.8. In order to draw out more information, the researchers are suggested to refer to the publications by Endo et al. (2016), Wang et al. (2017), Xu et al. (2017), and Jun et al. (2019).
All the successful reports regarding the application of CRISPR/Cpf1 in rice are enlisted in Table 23.9.
6 Conclusion
In the past five decades, crop improvement via traditional breeding has significantly contributed to acquiring food security for the every second whopping human population. However, various developments require more manpower, time duration, efforts along with high chance of failures in getting the “desirable traits”. Additionally, other conventional technologies like chemical mutagenesis, somaclonal variation, in vitro tissue culture and physical irradiation have also multiple loopholes. For increasing crop production under the changing climate as well as fulfilling the calorific and nutritional demands of mankind, the most recent, advanced nuclease-based technologies have emerged as the most suitable candidate in many crops including rice. Among all these technologies, the CRISPR/Cas9 tool is more precise, easy to handle, and also employed for avoiding backcrossing of a huge number of inbred lines. The varietal development using CRISPR/Cas9 technology consumes less time and is easy to introduce/restore desired changes in the existing elite rice germ plasm. Recently, multiple genes have been stacked together to get the desired phenotype in rice. Additionally, due to the advances like base editing, gene targeting, and DNA-free genome editing, the rice researchers have affirmatively taken a big leap towards the biggest milestone, i.e., super rice generation. Furthermore, due to the technical advances in the post-genomic era, the researchers have characterized a plethora of negative regulatory genes, SNPs, and QTLs for various traits. Taking these points, we hope that our children will be eating the socially accepted, highly nutritious super rice in the long run in the future.
References
Abuajah CI, Ogbonna AC, Osuji CM (2015) Functional components and medicinal properties of food: a review. J Food Sci Technol 52(5):2522–2529
Abbott TR, Qi LS (2018) Evolution at the cutting edge: CRISPR-mediated directed evolution. Mol Cell 72(3):402–403
Alagoz Y, Gurkok T, Zhang B, Unver T (2016) Manipulating the biosynthesis of bioactive compound alkaloids for next-generation metabolic engineering in opium poppy using CRISPR-Cas 9 genome editing technology. Sci Rep 6:30910
Alok A, Kumar J, Jogam P, Sandhya D (2018) CRISPR/Cas9-mediated gene editing tool and fathomless genetic and metabolic engineering applications in plants. In: Recent trends and techniques in plant metabolic engineering. Springer, Singapore, pp 167–179
Arzani A, Ashraf M (2016) Smart engineering of genetic resources for enhanced salinity tolerance in crop plants. Crit Rev Plant Sci 35(3):146–189
Ashkani S, Rafii MY, Shabanimofrad M, Miah G, Sahebi M, Azizi P, Tanweer FA, Akhtar MS, Nasehi A (2015) Molecular breeding strategy and challenges towards improvement of blast disease resistance in rice crop. Front Plant Sci 6:886
Atkins P, Bowler I (2016) Food in society: economy, culture, geography. Routledge, Abingdon
Anamika MS, Singh B, Patra A, Islam MA (2019) Databases: a weapon from the arsenal of bioinformatics for plant abiotic stress research. In: Recent approaches in omics for plant resilience to climate change. Springer, Cham, pp 135–169
Azadi P, Bagheri H, Nalousi AM, Nazari F, Chandler SF (2016) Current status and biotechnological advances in genetic engineering of ornamental plants. Biotechnol Adv 34(6):1073–1090
Azzam O, Chancellor TC (2002) The biology, epidemiology, and management of rice tungro disease in Asia. Plant Dis 86(2):88–100
Bado S, Forster BP, Nielen S, Ali AM, Lagoda PJ, Till BJ, Laimer M (2015) Plant mutation breeding: current progress and future assessment. Plant Breed Rev 39:23–88
Bado S, Yamba NG, Sesay JV, Laimer M, Forster BP (2017) Plant mutation breeding for the improvement of vegetatively propagated crops: successes and challenges. CAB Reviews 12(028):1–21
Bae S, Park J, Kim JS (2014) Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30(10):1473–1475
Baltes NJ, Voytas DF (2015) Enabling plant synthetic biology through genome engineering. Trends Biotechnol 33(2):120–131
Baltes NJ, Gil-Humanes J, Voytas DF (2017) Genome engineering and agriculture: opportunities and challenges. In: Progress in molecular biology and translational science, vol 149. Academic Press, Cambridge, pp 1–26
Banasik A, Kanellopoulos A, Claassen GD, Bloemhof-Ruwaard JM, van der Vorst JG (2017) Assessing alternative production options for eco-efficient food supply chains using multi-objective optimization. Ann Oper Res 250(2):341–362
Banerjee A, Roychoudhury A (2019) Genetic engineering in plants for enhancing arsenic tolerance. In: Transgenic plant technology for remediation of toxic metals and metalloids. Academic Press, Cambridge, pp 463–475
Baret PV, Vanloqueren G (2017) How agricultural research systems shape a technological regime that develops genetic engineering but locks out agroecological innovations. In: Food sovereignty, agroecology and biocultural diversity. Routledge, Abingdon, pp 77–112
Belhaj K, Chaparro-Garcia A, Kamoun S, Nekrasov V (2013) Plant genome editing made easy: targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods 9(1):39
Belkhodja R (2018) Breeding small grain cereal crops in a climate change scenario. https://hdl.handle.net/20.500.11766/9623
Bhowmik P, Ellison E, Polley B, Bollina V, Kulkarni M, Ghanbarnia K, Song H, Gao C, Voytas DF, Kagale S (2018) Targeted mutagenesis in wheat microspores using CRISPR/Cas9. Sci Rep 8
Bilichak A, Eudes F (2016) Genetic engineering of plants using Zn fingers, TALENs, and CRISPRs. In: Genome stability. Academic Press, Cambridge, pp 187–201
Bondy-Denomy J, Pawluk A, Maxwell KL, Davidson AR (2013) Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493(7432):429
Butt H, Jamil M, Wang JY, Al-Babili S, Mahfouz M (2018) Engineering plant architecture via CRISPR/Cas9-mediated alteration of strigolactone biosynthesis. BMC Plant Biol 18(1):174
Cai L, Cao Y, Xu Z, Ma W, Zakria M, Zou L, Cheng Z, Chen G (2017) A transcription activator-like effector Tal7 of Xanthomonasoryzaepv. oryzicola activates rice gene Os09g29100 to suppress rice immunity. Sci Rep 7(1):5089
Cazzolla Gatti R (2016) Trends in human development and environmental protection. Int J Environ Stud 73(2):268–276
Certo MT, Gwiazda KS, Kuhar R, Sather B, Curinga G, Mandt T, Brault M, Lambert AR, Baxter SK, Jacoby K, Ryu BY (2012) Coupling endonucleases with DNA end–processing enzymes to drive gene disruption. Nat Methods 9(10):973
Chancellor TCB, Holt J, Villareal S, Tiongco ER, Venn J (2006) Spread of plant virus disease to new plantings: a case study of rice tungro disease. Adv Virus Res 66:1–29
Chao S, Cai Y, Feng B, Jiao G, Sheng Z, Luo J, Tang S, Wang J, Wei X, Hu P (2019) Editing of the rice isoamylase gene ISA1 provides insights into its function in starch formation. Rice Sci 26(2):77–87
Char SN, Neelakandan AK, Nahampun H, Frame B, Main M, Spalding MH, Becraft PW, Meyers BC, Walbot V, Wang K, Yang B (2017) An Agrobacterium-delivered CRISPR/Cas9 system for high-frequency targeted mutagenesis in maize. Plant Biotechnol J 15(2):257–268
Chari R, Mali P, Moosburner M, Church GM (2015) Unraveling CRISPR/Cas9 genome engineering parameters via a library-on-library approach. Nat Methods 12(9):823
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, Voytas DF (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186(2):757–761
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823
Copetti D, Zhang J, El Baidouri M, Gao D, Wang J, Barghini E, Cossu RM, Angelova A, Roffler S, Ohyanagi H, Wicker T (2015) RiTE database: a resource database for genus-wide rice genomics and evolutionary biology. BMC Genomics 16(1):538
Crossa J, Pérez-Rodríguez P, Cuevas J, Montesinos-López O, Jarquín D, de los Campos G, Burgueño J, González-Camacho JM, Pérez-Elizalde S, Beyene Y, Dreisigacker S (2017) Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Sci 22(11):961–975
Daboussi F, Stoddard TJ, Zhang F (2015) Engineering meganuclease for precise plant genome modification. In: Advances in new technology for targeted modification of plant genomes. Springer, New York, pp 21–38
Darwin C (2010) The variation of animals and plants under domestication. Cambridge University Press, Cambridge
Dhakate P, Sharma P, Mehta S, Akter J, Bhatt V, Chandanshive S, Chakravarty D, Rahman M, Islam MA (2019) Beneficial role of metalloids in plants: molecular understanding and applicability. In: Recent approaches in omics for plant resilience to climate change. Springer, Cham, pp 357–387
Dillard HR (2019) Global food and nutrition security: from challenges to solutions. Food Sec 11:249–252
Ding D, Chen K, Chen Y, Li H, Xie K (2018) Engineering introns to express RNA guides for Cas9-and Cpf1-mediated multiplex genome editing. Mol Plant 11(4):542–552
Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I, Sullender M, Ebert BL, Xavier RJ, Root DE (2014) Rational design of highly active sgRNAs for CRISPR/Cas9–mediated gene inactivation. Nat Biotechnol 32(12):1262
Durai S, Mani M, Kandavelou K, Wu J, Porteus MH, Chandrasegaran S (2005) Zinc finger nucleases: custom-designed molecular scissors for genome engineering of plant and mammalian cells. Nucleic Acids Res 33(18):5978–5990
Endo A, Masafumi M, Kaya H, Toki S (2016) Efficient targeted mutagenesis of rice and tobacco genomes using Cpf1 from Francisella novicida. Sci Rep 6:38169
Eş I, Gavahian M, Marti-Quijal FJ, Lorenzo JM, Khaneghah AM, Tsatsanis C, Kampranis SC, Barba FJ (2019) The application of the CRISPR/Cas9 genome editing machinery in food and agricultural science: current status, future perspectives, and associated challenges. Biotechnol Adv 37(3):410–421
Feng Z, Zhang B, Ding W, Liu X, Yang DL, Wei P, Cao F, Zhu S, Zhang F, Mao Y, Zhu JK (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23(10):1229
Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L (2014a) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in arabidopsis. Proc Natl Acad Sci 111(12):4632–4637
Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L (2014b) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in arabidopsis. Proc Natl Acad Sci U S A 111(12):4632–4637
Figueroa A (2019) Population and the quality of society. In: The quality of society. Palgrave Macmillan, Cham, pp 115–137
Fukuoka S, Yamamoto SI, Mizobuchi R, Yamanouchi U, Ono K, Kitazawa N, Yasuda N, Fujita Y, Nguyen TT, Koizumi S, Sugimoto K (2014) Multiple functional polymorphisms in a single disease resistance gene in rice enhance durable resistance to blast. Sci Rep 4:4550
Gaj T, Gersbach CA, Barbas CF III (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405
Gao C (2018) The future of CRISPR technologies in agriculture. Nat Rev Mol Cell Biol 19(5):275–276
Gao J, Wang G, Ma S, Xie X, Wu X, Zhang X, Wu Y, Zhao P, Xia Q (2015) CRISPR/Cas9-mediated targeted mutagenesis in nicotianatabacum. Plant Mol Biol 87(1–2):99–110
Goff SA, Ricke D, Lan TH, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296(5565):92–100
Glenn KC, Alsop B, Bell E, Goley M, Jenkinson J, Liu B, Martin C, Parrott W, Souder C, Sparks O, Urquhart W (2017) Bringing new plant varieties to market: plant breeding and selection practices advance beneficial characteristics while minimizing unintended changes. Crop Sci 57(6):2906–2921
Haeussler M, Schönig K, Eckert H, Eschstruth A, Mianné J, Renaud JB, Schneider-Maunoury S, Shkumatava A, Teboul L, Kent J, Joly JS (2016) Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol 17(1):148
Heigwer F, Kerr G, Boutros M (2014) E-CRISP: fast CRISPR target site identification. Nat Methods 11(2):122
Hensel G, Kumlehn J (2019) Genome engineering using TALENs. In: Barley. Humana Press, New York, NY, pp 195–215
Horvath P, Barrangou R (2010) CRISPR/Cas, the immune system of bacteria and archaea. Science 327(5962):167–170
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, Cradick TJ (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31(9):827
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR/Cas9 for genome engineering. Cell 157(6):1262–1278
Hu X, Wang C, Fu Y, Liu Q, Jiao X, Wang K (2016) Expanding the range of CRISPR/Cas9 genome editing in rice. Mol Plant 9(6):943–945
Hu X, Wang C, Liu Q, Fu Y, Wang K (2017) Targeted mutagenesis in rice using CRISPR-Cpf1 system. J Genet Genomics 44(1):71–73
Huang TK, Puchta H (2019) CRISPR/Cas-mediated gene targeting in plants: finally a turn for the better for homologous recombination. Plant Cell Rep 38(4):443–453
Huang X, Zeng X, Li J, Zhao D (2017) Construction and analysis of tify1a and tify1b mutants in rice (Oryza sativa) based on CRISPR/Cas9 technology. J Agric Biotechnol 25(6):1003–1012
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res 41(20):e188
Jun R, Xixun H, Kejian W, Chun W (2019) Development and application of CRISPR/Cas system in rice. Ric Sci 26(2):69–76
Kamburova VS, Nikitina EV, Shermatov SE, Buriev ZT, Kumpatla SP, Emani C, Abdurakhmonov IY (2017) Genome editing in plants: an overview of tools and applications. Int J Agron 2017:1–15
Kantar MB, Runck BC, Raghavan B, Joglekar AB, Senay S, Krohn B, Neyhart J, Bradeen J, Soto Gomez M, Kjelgren R (2019) The many-faced Janus of plant breeding. Plants People Planet 1(4):306–309. https://doi.org/10.1002/ppp3.30
Kaur K, Tandon H, Gupta AK, Kumar M (2015) CrisprGE: a central hub of CRISPR/Cas-based genome editing. Database 2015:bav055
Khalid A, Zhang Q, Yasir M, Li F (2017) Small RNA based genetic engineering for plant viral resistance: application in crop protection. Front Microbiol 8:43
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol 59(1):1–6
Kihoro J, Bosco NJ, Murage H, Ateka E, Makihara D (2013) Investigating the impact of rice blast disease on the livelihood of the local farmers in greater Mwea region of Kenya. Springerplus 2(1):308
Kim HJ, Cho SW, Jeong JS, Kim JK, Lim YP, Kim JS, Kang BC (2012) Construction, selection, and application of zinc finger nucleases as a tool for genome editing in rice. 한국원예학회학술발표요지. Seoul National University Open Repository. pp 45–46. http://s-space.snu.ac.kr/handle/10371/121043
Kleter GA, Kuiper HA, Kok EJ (2019) Gene-edited crops: towards a harmonized safety assessment. Trends Biotechnol 37(5):443–447
Knott GJ, Doudna JA (2018) CRISPR-Cas guides the future of genetic engineering. Science 361(6405):866–869
Korotkova AM, Gerasimova SV, Shumny VK, Khlestkina EK (2017) Crop genes modified using the CRISPR/Cas system. Russian J Genet Appl Res 7(8):822–832
Korotkova AM, Gerasimova SV, Khlestkina EK (2019) Current achievements in modifying crop genes using CRISPR/Cas system. Vavilov J Genet Breed 23(1)
Kumari P, Rastogi A, Shukla A, Srivastava S, Yadav S (2018) Prospects of genetic engineering utilizing potential genes for regulating arsenic accumulation in plants. Chemosphere 211:397–406
Labun K, Montague TG, Gagnon JA, Thyme SB, Valen E (2016) CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res 44(W1):W272–W276
Laible G, Wei J, Wagner S (2015) Improving livestock for agriculture–technological progress from random transgenesis to precision genome editing heralds a new era. Biotechnol J 10(1):109–120
Langner T, Kamoun S, Belhaj K (2018) CRISPR crops: plant genome editing toward disease resistance. Annu Rev Phytopathol 56:479–512
Lassoued R, Macall DM, Hesseln H, Phillips PW, Smyth SJ (2019) Benefits of genome-edited crops: expert opinion. Transgenic Res 28(2):247–256
Lee JH, Muhsin M, Atienza GA, Kwak DY, Kim SM, De Leon TB, Angeles ER, Coloquio E, Kondoh H, Satoh K, Cabunagan RC (2010) Single nucleotide polymorphisms in a gene for translation initiation factor (eIF4G) of rice (Oryza sativa) associated with resistance to Rice tungro spherical virus. Mol Plant Microbe Interact 23(1):29–38
Li C, Zhang B (2019) Genome editing in cotton using CRISPR/Cas9 system. In: Transgenic cotton. Humana Press, New York, pp 95–104
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30(5):390
Li J, Meng X, Zong Y, Chen K, Zhang H, Liu J, Li J, Gao C (2016a) Gene replacements and insertions in rice by intron targeting using CRISPR–Cas9. Nat Plants 2(10):16139
Li T, Liu B, Chen CY, Yang B (2016b) TALEN-mediated homologous recombination produces site-directed DNA base change and herbicide-resistant rice. J Genet Genomics 43(5):297–305
Li M, Li X, Zhou Z, Wu P, Fang M, Pan X, Lin Q, Luo W, Wu G, Li H (2016c) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front Plant Sci 7:377
Li J, Sun Y, Du J, Zhao Y, Xia L (2017a) Generation of targeted point mutations in rice by a modified CRISPR/Cas9 system. Mol Plant 10(3):526–529
Li X, Zhou W, Ren Y, Tian X, Lv T, Wang Z, Fang J, Chu C, Yang J, Bu Q (2017b) High-efficiency breeding of early-maturing rice cultivars via CRISPR/Cas9-mediated genome editing. J Genet Genomics 44(3):175
Li S, Li J, He Y, Xu M, Zhang J, Du W, Zhao Y, Xia L (2019) Precise gene replacement in rice by RNA transcript-templated homologous recombination. Nat Biotechnol 37(4):445–450
Liang Z, Chen K, Li T, Zhang Y, Wang Y, Zhao Q, Liu J, Zhang H, Liu C, Ran Y, Gao C (2017) Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat Commun 8:14261
Lin CS, Hsu CT, Yang LH, Lee LY, Fu JY, Cheng QW, Wu FH, Hsiao HC, Zhang Y, Zhang R, Chang WJ (2018) Application of protoplast technology to CRISPR/Cas9 mutagenesis: from single-cell mutation detection to mutant plant regeneration. Plant Biotechnol J 16(7):1295–1310
Lino CA, Harper JC, Carney JP, Timlin JA (2018) Delivering CRISPR: a review of the challenges and approaches. Drug Deliv 25(1):1234–1257
Liu D, Chen X, Liu J, Ye J, Guo Z (2012) The rice ERF transcription factor OsERF922 negatively regulates resistance to magnaporthe oryzae and salt tolerance. J Exp Bot 63(10):3899–3911
Liu H, Ding Y, Zhou Y, Jin W, Xie K, Chen LL (2017a) CRISPR-P 2.0: an improved CRISPR/Cas9 tool for genome editing in plants. Mol Plant 10(3):530–532
Liu X, Wu S, Xu J, Sui C, Wei J (2017b) Application of CRISPR/Cas9 in plant biology. Acta Pharm Sin B 7(3):292–302
Lloyd A, Plaisier CL, Carroll D, Drews GN (2005) Targeted mutagenesis using zinc-finger nucleases in arabidopsis. Proc Natl Acad Sci 102(6):2232–2237
Lou D, Wang H, Liang G, Yu D (2017) OsSAPK2 confers abscisic acid sensitivity and tolerance to drought stress in rice. Front Plant Sci 8:993
Lowder LG, Zhang D, Baltes NJ, Paul JW, Tang X, Zheng X, Voytas DF, Hsieh TF, Zhang Y, Qi Y (2015) A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol 169(2):971–985
Lu Y, Zhu JK (2017) Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol Plant 10(3):523–525
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8(8):1274–1284
Macovei A, Sevilla NR, Cantos C, Jonson GB, Slamet-Loedin I, Čermák T, Voytas DF, Choi IR, Chadha-Mohanty P (2018) Novel alleles of rice eIF4G generated by CRISPR/Cas9-targeted mutagenesis confer resistance to Rice tungro spherical virus. Plant Biotechnol J 16(11):1918–1927
MacPherson CR, Scherf A (2015) Flexible guide-RNA design for CRISPR applications using Protospacer workbench. Nat Biotechnol 33(8):805
Mahas A, Ali Z, Tashkandi M, Mahfouz MM (2019) Virus-mediated genome editing in plants using the CRISPR/Cas9 system. In: Plant genome editing with CRISPR systems. Humana Press, New York, pp 311–326
Malnoy M, Viola R, Jung MH, Koo OJ, Kim S, Kim JS, Velasco R, NagamangalaKanchiswamy C (2016) DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7:1904
Malzahn A, Lowder L, Qi Y (2017) Plant genome editing with TALEN and CRISPR. Cell Biosci 7(1):21
Marco F, Bitrián M, Carrasco P, Rajam MV, Alcázar R, Tiburcio AF (2015) Genetic engineering strategies for abiotic stress tolerance in plants. In: Plant biology and biotechnology. Springer, New Delhi, pp 579–609
Martirosyan DM, Singh J (2015) A new definition of functional food by FFC: what makes a new definition unique? Funct Food Health Dis 5(6):209–223
Mehta S, Singh B, Dhakate P, Rahman M, Islam MA (2019a) Rice, marker-assisted breeding, and disease resistance. In: Disease resistance in crop plants. Springer, Cham, pp 83–111
Mehta S, James D, Reddy MK (2019b) Omics technologies for abiotic stress tolerance in plants: current status and prospects. In: Recent approaches in omics for plant resilience to climate change. Springer, Cham, pp 1–34
Merga IF, Tripathi L, Hvoslef-Eide AK, Gebre E (2019) Application of genetic engineering for control of bacterial wilt disease of Enset, Ethiopia’s sustainability crop. Front Plant Sci 10:133
Mew TW, Alvarez AM, Leach JE, Swings J (1993) Focus on bacterial blight of rice. Plant Dis 77(1):5–12
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23(10):1233
Miao C, Xiao L, Hua K, Zou C, Zhao Y, Bressan RA, Zhu JK (2018) Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc Natl Acad Sci 115(23):6058–6063
Mikami M, Toki S, Endo M (2015a) Comparison of CRISPR/Cas9 expression constructs for efficient targeted mutagenesis in rice. Plant Mol Biol 88(6):561–572
Mikami M, Toki S, Endo M (2015b) Parameters affecting frequency of CRISPR/Cas9 mediated targeted mutagenesis in rice. Plant Cell Rep 34(10):1807–1815
Minkenberg B, Zhang J, Xie K, Yang Y (2019) CRISPR-PLANT v2: an online resource for highly specific guide RNA spacers based on improved off-target analysis. Plant Biotechnol J 17(1):5–8
Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E (2014) CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res 42(W1):W401–W407
Moreno-Mateos MA, Vejnar CE, Beaudoin JD, Fernandez JP, Mis EK, Khokha MK, Giraldez AJ (2015) CRISPRscan: designing highly efficient sgRNAs for CRISPR/Cas9 targeting in vivo. Nat Methods 12(10):982
Morton LW, Roesch-McNally G, Wilke AK (2017) Upper Midwest farmer perceptions: too much uncertainty about impacts of climate change to justify changing current agricultural practices. J Soil Water Conserv 72(3):215–225
Muralidharan K, Krishnaveni D, Rajarajeswari NV, Prasad AS (2003) Tungro epidemics and yield losses in paddy fields in India. Curr Sci 85(8):1143–1147
Naito Y, Hino K, Bono H, Ui-Tei K (2014) CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31(7):1120–1123
Nielsen J, Keasling JD (2016) Engineering cellular metabolism. Cell 164(6):1185–1197
Noman A, Aqeel M, He S (2016) CRISPR/Cas9: tool for qualitative and quantitative plant genome editing. Front Plant Sci 7:1740
Novak S (2019) Plant biotechnology applications of zinc finger technology. In: Transgenic plants. Humana Press, New York, pp 295–310
Oerke EC (2006) Crop losses to pests. J Agric Sci 144(1):31–43
Oldeman LR, Hakkeling RT, Sombroek WG (2017) World map of the status of human-induced soil degradation: an explanatory note. International Soil Reference and Information Centre, Wageningen
Oliveros JC, Franch M, Tabas-Madrid D, San-León D, Montoliu L, Cubas P, Pazos F (2016) Breaking-Cas—interactive design of guide RNAs for CRISPR-Cas experiments for ENSEMBL genomes. Nucleic Acids Res 44(W1):W267–W271
Osakabe Y, Watanabe T, Sugano SS, Ueta R, Ishihara R, Shinozaki K, Osakabe K (2016) Optimization of CRISPR/Cas9 genome editing to modify abiotic stress responses in plants. Sci Rep 6:26685
Osakabe Y, Liang Z, Ren C, Nishitani C, Osakabe K, Wada M, Komori S, Malnoy M, Velasco R, Poli M, Jung MH (2018) CRISPR–Cas9-mediated genome editing in apple and grapevine. Nat Protoc 13(12):2844
Pacher M, Puchta H (2017) From classical mutagenesis to nuclease-based breeding–directing natural DNA repair for a natural end-product. Plant J 90(4):819–833
Park J, Bae S (2017) Cpf1-database: web-based genome-wide guide RNA library design for gene knockout screens using CRISPR-Cpf1. Bioinformatics 34(6):1077–1079
Park J, Kim JS, Bae S (2016) Cas-database: web-based genome-wide guide RNA library design for gene knockout screens using CRISPR/Cas9. Bioinformatics 32(13):2017–2023
Perez AR, Pritykin Y, Vidigal JA, Chhangawala S, Zamparo L, Leslie CS, Ventura A (2017) GuideScan software for improved single and paired CRISPR guide RNA design. Nat Biotechnol 35(4):347
Philander SG (2018) Is the temperature rising?: the uncertain science of global warming. Princeton University Press, Princeton
Pliatsika V, Rigoutsos I (2015) “Off-Spotter”: very fast and exhaustive enumeration of genomic lookalikes for designing CRISPR/Cas guide RNAs. Biol Direct 10(1):4
Porteus MH, Carroll D (2005) Gene targeting using zinc finger nucleases. Nat Biotechnol 23(8):967
Pradinaud C, Northey S, Amor B, Bare J, Benini L, Berger M, Boulay AM, Junqua G, Lathuillière MJ, Margni M, Motoshita M (2019) Defining freshwater as a natural resource: a framework linking water use to the area of protection natural resources. Int J Life Cycle Assess 24(5):960–974
Pratim Roy M (2019) Malnutrition in children and its determinants: a study from east India. Tropical Doctor. 0049475518824825
Pray C, Huang J, Hu R, Deng H, Yang J, Morin XK (2018) Prospects for cultivation of genetically engineered food crops in China. Glob Food Sec 16:133–137
Prykhozhij SV, Rajan V, Gaston D, Berman JN (2015) CRISPR multitargeter: a web tool to find common and unique CRISPR single guide RNA targets in a set of similar sequences. PLoS One 10(3):e0119372
Project, I.R.G.S, Sasaki T (2005) The map-based sequence of the rice genome. Nature 436(7052):793
Puchta H, Fauser F (2014) Synthetic nucleases for genome engineering in plants: prospects for a bright future. Plant J 78(5):727–741
Rahman M, Sultana S, Nath D, Kalita S, Chakravarty D, Mehta S, Wani SH, Islam MA (2019) Molecular breeding approaches for disease resistance in sugarcane. In: Disease resistance in crop plants. Springer, Cham, pp 131–155
Ricroch A, Clairand P, Harwood W (2017) Use of CRISPR systems in plant genome editing: toward new opportunities in agriculture. Emerg Top Life Sci 1(2):169–182
Sakamoto T, Matsuoka M (2008) Identifying and exploiting grain yield genes in rice. Curr Opin Plant Biol 11(2):209–214
Sampson TR, Saroj SD, Llewellyn AC, Tzeng YL, Weiss DS (2013) A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature 497(7448):254
Savary S, Ficke A, Aubertot JN, Hollier C (2012) Crop losses due to diseases and their implications for global food production losses and food security. Food Sec 4:519–537
Scheben A, Wolter F, Batley J, Puchta H, Edwards D (2017) Towards CRISPR/Cas crops–bringing together genomics and genome editing. New Phytol 216(3):682–698
Shah T, Andleeb T, Lateef S, Noor MA (2018) Genome editing in plants: advancing crop transformation and overview of tools. Plant Physiol Biochem 131:12–21
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL, Gao C (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31(8):686
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9(10):2395
Shao GN, Xie LH, Jiao GA, Wei XJ, Sheng ZH, Tang SQ, Hu PS (2017) CRISPR/CAS9-mediated editing of the fragrant gene Badh2 in rice. Chin J Rice Sci 31(2):216–222
Shen B, Zhang W, Zhang J, Zhou J, Wang J, Chen L, Wang L, Hodgkins A, Iyer V, Huang X, Skarnes WC (2014) Efficient genome modification by CRISPR/Cas9 nickase with minimal off-target effects. Nat Methods 11(4):399–402
Shimatani Z, Kashojiya S, Takayama M, Terada R, Arazoe T, Ishii H, Teramura H, Yamamoto T, Komatsu H, Miura K, Ezura H (2017a) Targeted base editing in rice and tomato using a CRISPR/Cas9 cytidine deaminase fusion. Nat Biotechnol 35(5):441–443
Shimatani Z, Kashojiya S, Takayama M, Terada R, Arazoe T, Ishii H, Teramura H, Yamamoto T, Komatsu H, Miura K, Ezura H (2017b) Targeted base editing in rice and tomato using a CRISPR/Cas9 cytidine deaminase fusion. Nat Biotechnol 35(5):441–443
Shigaki T (2018) Application of biotechnologies in the conservation and utilization of plant genetic resources for food security. In: Food Security and Climate Change, pp 413–431
Singh B, Mehta S, Aggarwal SK, Tiwari M, Bhuyan SI, Bhatia S, Islam MA (2019) Barley, disease resistance, and molecular breeding approaches. In: Disease resistance in crop plants. Springer, Cham, pp 261–299
Siro I, Kápolna E, Kápolna B, Lugasi A (2008) Functional food. Product development, marketing and consumer acceptance—a review. Appetite 51(3):456–467
Smita S, Lenka SK, Katiyar A, Jaiswal P, Preece J, Bansal KC (2011) QlicRice: a web interface for abiotic stress responsive QTL and loci interaction channels in rice. Database 2011:bar037
Soda N, Verma L, Giri J (2018) CRISPR/Cas9 based plant genome editing: significance, opportunities and recent advances. Plant Physiol Biochem 131:2–11
Sprink T, Metje J, Hartung F (2015) Plant genome editing by novel tools: TALEN and other sequence specific nucleases. Curr Opin Biotechnol 32:47–53
Stemmer M, Thumberger T, del Sol Keyer M, Wittbrodt J, Mateo JL (2015) CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS One 10(4):e0124633
Sticklen MB (2008) Plant genetic engineering for biofuel production: towards affordable cellulosic ethanol. Nat Rev Genet 9(6):433–443
Subramanian KR (2018) The crisis of consumption of natural resources. Int J Rec Innov Acad Res 2(8–19):8–19
Sun Y, Zhang X, Wu C, He Y, Ma Y, Hou H, Guo X, Du W, Zhao Y, Xia L (2016) Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of acetolactate synthase. Mol Plant 9(4):628–631
Sun Y, Jiao G, Liu Z, Zhang X, Li J, Guo X, Du W, Du J, Francis F, Zhao Y, Xia L (2017) Generation of high-amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes. Front Plant Sci 8:298
Sun J, Liu H, Liu J, Cheng S, Peng Y, Zhang Q, Yan J, Liu HJ, Chen LL (2019) CRISPR-local: a local single-guide RNA (sgRNA) design tool for non-reference plant genomes. Bioinformatics 35(14):2501–2503
Svitashev S, Young JK, Schwartz C, Gao H, Falco SC, Cigan AM (2015) Targeted mutagenesis, precise gene editing, and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol 169(2):931–945
Takeda Y, Suzuki S, Tobimatsu Y, Osakabe K, Osakabe Y, Ragamustari SK, Sakamoto M, Umezawa T (2019) Lignin characterization of rice coniferaldehyde 5-hydroxylase loss-of-function mutants generated with the CRISPR/Cas9 system. Plant J 97(3):543–554
Tang X, Lowder LG, Zhang T, Malzahn AA, Zheng X, Voytas DF, Zhong Z, Chen Y, Ren Q, Li Q, Kirkland ER (2017a) A CRISPR–Cpf1 system for efficient genome editing and transcriptional repression in plants. Nat Plants 3(3):17018
Tang L, Mao B, Li Y, Lv Q, Zhang L, Chen C, He H, Wang W, Zeng X, Shao Y, Pan Y (2017b) Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep 7(1):14438
Tang X, Ren Q, Yang L, Bao Y, Zhong Z, He Y, Liu S, Qi C, Liu B, Wang Y, Sretenovic S (2019) Single transcript unit CRISPR 2.0 systems for robust Cas9 and Cas 12a mediated plant genome editing. Plant Biotechnol J 17(7):1431–1445
Tsutsui H, Higashiyama T (2017) pKAMA-ITACHI vectors for highly efficient CRISPR/Cas9-mediated gene knockout in Arabidopsis thaliana. Plant Cell Physiol 58(1):46–56
Upadhyay SK, Kumar J, Alok A, Tuli R (2013) RNA-guided genome editing for target gene mutations in wheat. G3 3(12):2233–2238
Valin H, Sands RD, Van der Mensbrugghe D, Nelson GC, Ahammad H, Blanc E, Bodirsky B, Fujimori S, Hasegawa T, Havlik P, Heyhoe E (2014) The future of food demand: understanding differences in global economic models. Agric Econ 45(1):51–67
Voytas DF (2013) Plant genome engineering with sequence-specific nucleases. Annu Rev Plant Biol 64:327–350
Waltz E (2018) With a free pass, CRISPR-edited plants reach market in record time. Nat Biotechnol 36(1):6–7
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y, Liu YG, Zhao K (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS One 11(4):e0154027
Wang M, Mao Y, Lu Y, Tao X, Zhu JK (2017) Multiplex gene editing in rice using the CRISPR-Cpf1 system. Mol Plant 10(7):1011–1013
Wang P, Zhang J, Sun L, Ma Y, Xu J, Liang S, Deng J, Tan J, Zhang Q, Tu L, Daniell H (2018) High efficient multisites genome editing in allotetraploid cotton (Gossypium hirsutum) using CRISPR/Cas9 system. Plant Biotechnol J 16(1):137–150
Wang X, Han Y, Feng X, Li YZ, Qin BX, Luo JJ, Wei Z, Qiu YF, Liu F, Li RB (2019) Breeding of Indica glutinous cytoplasmic male sterile line WX209A via CRISPR/Cas9 mediated genomic editing. Czech J Genet Plant Breed 55:93–100
Wingeyer A, Amado T, Pérez-Bidegain M, Studdert G, Varela C, Garcia F, Karlen D (2015) Soil quality impacts of current South American agricultural practices. Sustainability 7(2):2213–2242
Wong N, Liu W, Wang X (2015) WU-CRISPR: characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genome Biol 16(1):218
Wright DA, Townsend JA, Winfrey RJ Jr, Irwin PA, Rajagopal J, Lonosky PM, Hall BD, Jondle MD, Voytas DF (2005) High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J 44(4):693–705
Wu G, Yan Q, Jones JA, Tang YJ, Fong SS, Koffas MA (2016) Metabolic burden: cornerstones in synthetic biology and metabolic engineering applications. Trends Biotechnol 34(8):652–664
Wu R, Lucke M, Jang YT, Zhu W, Symeonidi E, Wang C, Fitz J, Xi W, Schwab R, Weigel D (2018) An efficient CRISPR vector toolbox for engineering large deletions in Arabidopsis thaliana. Plant Methods 14(1):65
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR–Cas system. Mol Plant 6(6):1975–1983
Xie X, Ma X, Zhu Q, Zeng D, Li G, Liu YG (2017) CRISPR-GE: a convenient software toolkit for CRISPR-based genome editing. Mol Plant 10(9):1246–1249
Xing Y, Zhang Q (2010) Genetic and molecular bases of rice yield. Annu Rev Plant Biol 61:421–442
Xing HL, Dong L, Wang ZP, Zhang HY, Han CY, Liu B, Wang XC, Chen QJ (2014) A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol 14(1):327
Xu R, Li H, Qin R, Wang L, Li L, Wei P, Yang J (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice 7(1):5
Xu R, Yang Y, Qin R, Li H, Qiu C, Li L, Wei P, Yang J (2016) Rapid improvement of grain weight via highly efficient CRISPR/Cas9-mediated multiplex genome editing in rice. J Genet Genomics 43(8):529
Xu R, Qin R, Li H, Li D, Li L, Wei P, Yang J (2017) Generation of targeted mutant rice using a CRISPR-Cpf1 system. Plant Biotechnol J 15(6):713–717
Xuan TD, Anh TT, Tran HD, Khanh TD, Dat TD (2019) Mutation breeding of a N-methyl-N-nitrosourea (MNU)-induced Rice (Oryza sativa L. ssp. Indica) population for the yield attributing traits. Sustainability 11(4):1062
Yin K, Qiu JL (2019) Genome editing for plant disease resistance: applications and perspectives. Philos Trans R Soc B 374(1767):20180322
Yin K, Han T, Liu G, Chen T, Wang Y, Yu AY, Liu Y (2015) A geminivirus-based guide RNA delivery system for CRISPR/Cas9 mediated plant genome editing. Sci Rep 5:14926
Yin K, Gao C, Qiu JL (2017) Progress and prospects in plant genome editing. Nat Plants 3(8):17107
Yin X, Anand A, Quick P, Bandyopadhyay A (2019) Editing a stomatal developmental gene in rice with CRISPR/Cpf1. In: Plant genome editing with CRISPR systems. Humana Press, New York, pp 257–268
Youssef D, Nihou A, Partier A, Tassy C, Paul W, Rogowsky PM, Beckert M, Barret P (2018) Induction of targeted deletions in transgenic bread wheat (Triticum aestivum L.) using customized meganuclease. Plant Mol Biol Report 36(1):71–81
Yu J, Hu S, Wang J, Wong GK, Li S, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, Cao M (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296(5565):79–92
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P, Volz SE, Joung J, Van Der Oost J, Regev A, Koonin EV (2015) Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163(3):759–771
Zhang H (2019) Genetically modified organisms (GMOs) as the new hope. In: Securing the ‘rice bowl’. Palgrave Macmillan, Singapore, pp 95–123
Zhang H, Wu Z, Wang C, Li Y, Xu JR (2014) Germination and infectivity of microconidia in the rice blast fungus Magnaporthe oryzae. Nat Commun 5:4518
Zhang Y, Zang Q, Xu B, Zheng W, Ban R, Zhang H, Yang Y, Hao Q, Iqbal F, Li A, Shi Q (2016) IsomiR Bank: a research resource for tracking IsomiRs. Bioinformatics 32(13):2069–2071
Zhang J, Zhang H, Botella JR, Zhu JK (2018) Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Integr Plant Biol 60(5):369–375
Zhang T, Liang J, Wang M, Li D, Liu Y, Chen TH, Yang X (2019) Genetic engineering of the biosynthesis of glycine betaine enhances the fruit development and size of tomato. Plant Sci 280:355–366
Zhao H, Yao W, Ouyang Y, Yang W, Wang G, Lian X, Xing Y, Chen L, Xie W (2014) RiceVarMap: a comprehensive database of rice genomic variations. Nucleic Acids Res 43(D1):D1018–D1022
Zheng Y, Zhang N, Martin GB, Fei Z (2019) Plant genome editing database (PGED): a call for submission of information about genome-edited plant mutants. Mol Plant 12(2):127–129
Zhong Z, Zhang Y, You Q, Tang X, Ren Q, Liu S, Yang L, Wang Y, Liu X, Liu B, Zhang T (2018) Plant genome editing using FnCpf1 and LbCpf1 nucleases at redefined and altered PAM sites. Mol Plant 11(7):999–1002
Zhou J, Peng Z, Long J, Sosso D, Liu B, Eom JS, Huang S, Liu S, Vera Cruz C, Frommer WB, White FF (2015) Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J 82(4):632–643
Zhu LJ, Holmes BR, Aronin N, Brodsky MH (2014) CRISPRseek: a bioconductor package to identify target-specific guide RNAs for CRISPR/Cas9 genome-editing systems. PLoS One 9(9):e108424
Zischewski J, Fischer R, Bortesi L (2017) Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases. Biotechnol Adv 35(1):95–104
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mehta, S. et al. (2020). CRISPR/Cas9-Edited Rice: A New Frontier for Sustainable Agriculture. In: Rakshit, A., Singh, H., Singh, A., Singh, U., Fraceto, L. (eds) New Frontiers in Stress Management for Durable Agriculture. Springer, Singapore. https://doi.org/10.1007/978-981-15-1322-0_23
Download citation
DOI: https://doi.org/10.1007/978-981-15-1322-0_23
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1321-3
Online ISBN: 978-981-15-1322-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)