Abstract
Cullin-5 (Cul-5) was originally identified as an arginine vasopressin (AVP) receptor due to its homology to a vasopressin-activated calcium-mobilizing protein 1 (VACM-1). Cul-5 has subsequently gained much attention after being identified as the key component of CRL-5 (Cullin-RING ligase-5) that mediates ubiquitylation and degradation of several key cellular proteins associated with human cancers and viral infections. Structurally, Cul-5 interacts with the Elongin B/C complex, a RING finger protein (RBX2/SAG), and a SOCS protein to form a CRL-5 E3 ubiquitin ligase protein complex. CRL-5, by controlling turnover of a variety of substrates, is implicated in several biological processes and human diseases. Activation of CRL-5 requires Cul-5 neddylation, catalyzed by a neddylation enzyme cascade, consisting of the E1 NEDD8-activating enzyme (NAE), the E2 neddylation conjugating enzyme (UBE2F), and E3 neddylation ligase (RBX2/SAG). RBX2/SAG, therefore, serves as both Cul-5 neddylation E3 and CRL-5 ubiquitylation E3. Here, we review the current knowledge on CRL-5, its activation by the UBE2F-SAG, its regulation of various signaling pathways via substrate degradation, and its implications in human cancers.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
16.1 Introduction
Cul-5 is the least conserved member of the cullin family, but highly homologous among various species (Byrd et al. 1997). In kidney cells, Cul-5 localizes to the cell membrane where it binds to AVP to control body fluid and blood pressure, thus retaining the homeostasis (Burnatowska-Hledin et al. 1995). Otherwise, Cul-5, as a scaffold component, complexes with adaptor proteins Elongin B/C, a RING finger protein RBX2/SAG, and a substrate receptor SOCS protein to form a CRL-5 E3 ligase complex. Structurally, Cul-5 possesses a long stalklike amino-terminal domain (NTD), which contains three cullin repeats (CR1, CR2, CR3), and a globular carboxyl-terminal domain (CTD), harboring a highly conserved signature cullin homology (CH) domain (Sarikas et al. 2011), along with the lysine residue (Lys724) for covalent NEDD8 attachment, a process known as neddylation for CRL-5 activation (Fig. 16.1). The amino-terminal helices H2 and H5 of CR1 are used to anchor the cognate adaptor, Elongin B/C. The CTD of Cul-5 binds to SAG, which recruits the ubiquitin-loaded E2 conjugating enzymes for catalysis of ubiquitylation reaction (Petroski and Deshaies 2005). The fourth component of CRL-5 is the suppressor of cytokine signaling (SOCS) protein, responsible for the recognition of cellular substrates, involved in various cellular functions. Interestingly, some viral proteins, including the viral infectivity factor (Vif) in human immunodeficiency virus-1 (HIV-1), adenovirus proteins E4orf6 and E1B55K, and latency-associated nuclear antigen (LANA) in Kaposi’s sarcoma-associated herpesvirus (KSHV), can hijack host CRL-5 complex to trigger the ubiquitylation and degradation of host defensive proteins (Yu et al. 2003; Querido et al. 2001; Cai et al. 2006). Thus, CRL-5 is involved in regulation of both cellular functions and viral infections.
Domain structures of Cul-5 and SOCS box containing proteins. Cullin repeat 1 (CR1) anchors the cognate adaptor proteins, and the cullin homology (CH) domain at the carboxyl-terminus is critical for the binding of the RING finger protein. The N8 site indicates the position of the neddylation site. The SOCS box consists of a BC box and a Cul5 box in the order indicated. SH2 Src homology 2 phosphotyrosine-binding domain, WD40 WD40 repeats, SPRY Sp1A/ryanodine receptor domain, Ank ankyrin repeats
16.2 The Family of Substrate Receptors and Their Substrates
In mammalian cells, four families of substrate receptors were identified in CRL-5 E3 ligase, and every family member contains a C-terminal SOCS box, consisting of a BC box for Elongin B/C binding and a Cul-5 box for Cul-5 binding (Fig. 16.1). CRL-5 ligases target a wide range of proteins for ubiquitylation and degradation with substrate specificity determined by substrate receptors.
16.2.1 Cytokine-Inducible SH2 (Src Homology 2) Domain-Containing SOCS Box Proteins
The SH2 domain-containing SOCS box proteins (SOCS1-7) have a central SH2 domain and a C-terminally located SOCS box, which consists of a BC box and a Cul-5 box with an approximately 40-amino acid motif (Fig. 16.1). The SOCS box interacts with Cul-5 via its amino acid sequence LPΦP (Φ represents a hydrophobic residue) within the Cul5 box (Mahrour et al. 2008; Okumura et al. 2012), and the BC box is responsible for binding to Elongin B/C (Endo et al. 1997) (Fig. 16.1). SOCS1 and SOCS3 have been extensively studied in leukocytes with the activity to inhibit JAK family tyrosine kinase signaling, which are mediated by both Cul5-independent and Cul5-dependent mechanisms (Mahrour et al. 2008; Kazi et al. 2014; Linossi and Nicholson 2015). As the substrate-recognizing subunits, the family of SOCS proteins is involved in ubiquitylation and subsequent proteasomal degradation of a variety of cellular proteins by CRL-5. Specifically, SOCS1 suppresses the signal transduction via targeting for degradation of a variety of cellular proteins, including Vav (De Sepulveda et al. 2000), focal adhesion kinase (FAK) (Liu et al. 2003), the NF-κB family member p65/RelA (Ryo et al. 2003), myeloid differentiation primary-response gene 88 adaptor-like protein (MAL) (Mansell et al. 2006), the Janus kinase 2 (JAK2) (Ungureanu et al. 2002), the TEL-JAK2 onco-fusion protein (Kamizono et al. 2001; Frantsve et al. 2001), Cdh1 (Parrillas et al. 2013), HPV E7 (Kamio et al. 2004), and insulin receptor substrates IRS1 and IRS2 (Rui et al. 2002). SOCS3 specifically binds to the phosphorylated immunoreceptor tyrosine-based inhibitory motif of CD33, resulting in accelerated CD33 degradation (Orr et al. 2007). SOCS6, on the other hand, is involved in the degradation of Cas and other unidentified Src substrates to inhibit Src-dependent cell transformation (Teckchandani et al. 2014). SOCS6 also targets p56lck and c-KIT for degradation (Choi et al. 2010; Lamsoul et al. 2016), whereas SOCS7 degrades Dab1 (Simo and Cooper 2013), an essential protein for neuron migration and positioning (Tissir and Goffinet 2003), thus playing an important role in normal neuron positioning during cerebral development.
16.2.2 Ankyrin Repeat-Containing SOCS Box (ASB) Family
The ASB family includes 18 members from ASB1 to ASB18, all of which contain two functional domains, the C-terminal SOCS box domain and an upstream ankyrin repeat region (Kile et al. 2002). Several members of the ASB family are able to interact with Cul5-SAG to form ubiquitin ligase complexes (Kohroki et al. 2005). ASB1 is expressed widely in a variety of organs, and ASB1 knockout mice show no significantly phenotypes, but with a diminution of spermatogenesis (Kile et al. 2001). On the other hand, ASB2 promotes the polyubiquitylation of the actin-binding protein filamins A and B for degradation, thereby modulating actin remodeling and regulating the cell differentiation (Heuze et al. 2008; Burande et al. 2009). ASB2 has also been shown to promote the degradation of mixed-lineage leukemia (MLL) protein, a factor required for hematopoietic differentiation, through interaction with its PHD/bromodomain region (Wang et al. 2012). ASB3 interacts with the C-terminus of the tumor necrosis factor receptor 2 (TNF-R2) and triggers its ubiquitylation and subsequent degradation, thereby suppressing TNF-R2-mediated JNK activation and apoptosis induction (Chung et al. 2005). ASB4 colocalizes and interacts with the insulin receptor substrate 4 (IRS4) in neurons of the hypothalamus to promote IRS4 ubiquitylation and degradation, thus modulating neuron sensitivity to circulating insulin levels (Li et al. 2011). ASB4 also binds to and promotes ubiquitylation and degradation of inhibitor of DNA binding 2 (ID2), thus mediating vascular differentiation in the placenta (Townley-Tilson et al. 2014). ASB6 is restrictedly expressed in adipose tissue. In 3T3-L1 adipocytes, ASB6 regulated the insulin signaling pathway by targeted ubiquitylation and degradation of the adapter protein with a pleckstrin homology and Src homology 2 domain (APS) upon activation of the insulin receptor (Wilcox et al. 2004). ASB7 is involved in the regulation of cell division by promoting the degradation of DDA3, a critical factor that controls chromosome compression and segregation via modulating the dynamics of the mitotic spindle (Uematsu et al. 2016). ASB9 is predominantly expressed in the kidney and testes, where it promotes the ubiquitination and degradation of brain-type cytosolic creatine kinase (CKB) (Debrincat et al. 2007) and ubiquitous mitochondrial creatine kinase (uMtCK) (Kwon et al. 2010). ASB10 is induced by inflammation cytokines and involved in protein degradation pathways in glaucoma (Keller and Wirtz 2017). The observation that ASB10 forms a complex with Cul-5, SAG, and Elongin B/C (Andresen et al. 2014) suggests a possible role of CRL-5 in the process. Finally, ASB11 is an endoplasmic reticulum (ER)-related ubiquitin ligase, which promotes ubiquitylation and degradation of ribophorin 1, an integral protein of the oligosaccharyltransferase (OST) glycosylation complex (Andresen et al. 2014; Kelleher et al. 1992). Furthermore, ASB11 affects the neural progenitor compartment of the embryos by specifically ubiquitylating Delta A for degradation, thereby regulating the canonical Delta-Notch signaling pathway (Diks et al. 2008).
16.2.3 SPSB (SplA/Ryanodine Receptor) Domain-Containing SOCS Box Proteins
The SPSB family is characterized by a central SPRY (SplA/ryanodine receptor)/domain and a C-terminal SOCS box with four members (Perfetto et al. 2013; Hilton et al. 1998). SPSB1, SPSB2, and SPSB4 were reported to promote ubiquitylation and degradation of inducible nitric oxide synthase (iNOS/NOS2) (Nishiya et al. 2011; Kuang et al. 2010; Lewis et al. 2011). Given that iNOS is responsible for sustained production of NO upon stimulation by microbes or cytokines (Lowenstein and Padalko 2004), SPSB1 and SPSB4, therefore, play important roles in preventing the overproduction of NO by triggering iNOS degradation (Lewis et al. 2011; Matsumoto et al. 2011). At the physiological aspect, SPSB2 knockout in macrophages results in excessive production of iNOS and NO to kill more Leishmania major parasites (Kuang et al. 2010). Furthermore, SPSB1 was reported to negatively regulate the TGF-β signaling pathway through an interaction with type II TGFβ receptor (TβRII) via its SPRY domain, leading to enhanced ubiquitylation and degradation of TβRII (Liu et al. 2015), whereas SPSB3 overexpression significantly inhibits tumor metastasis by promoting polyubiquitylation and degradation of SNAIL upon phosphorylation mediated by GSK-3β (Liu et al. 2018).
16.2.4 WD Repeat-Containing SOCS Box Protein 1 (WSB1)
WSB1 is another member of SOCS protein responsible for ubiquitylation and degradation of several key regulatory proteins. First, WSB1 promotes ubiquitylation and degradation of homeodomain-interacting protein kinase 2 (HIPK2) (Choi et al. 2008), a nuclear protein kinase that triggers apoptosis in part by activation of p53 (Puca et al. 2009). Thus, approaches that block WSB1-mediated HIPK2 degradation, such as treatment with adriamycin or cisplatin, enhance the DNA damage-induced apoptosis (Choi et al. 2008). Second, WSB1 was reported to promote ubiquitylation and degradation of von Hippel-Lindau tumor suppressor (pVHL), thus stabilizing hypoxia-inducible factor-α (HIF-α) under both normoxic and hypoxic conditions. The highly level of HIF-α maintained by WSB1 is responsible for enhanced cancer metastasis (Kim et al. 2015). Third, WSB1 promotes degradation of Rho-binding protein RhoGDI2 under tumor hypoxic environment, thereby inducing Rac1 activation to stimulate osteosarcoma cell migration and invasion (Cao et al. 2015). Fourth, a recent study reported that WSB1 triggers the ubiquitylation and degradation of ATM to bypass the oncogene-induced senescence, contributing to abnormal cell proliferation and cellular transformation (Kim et al. 2017). Finally, WSB1 was reported to promote ubiquitylation of yet two additional proteins, not for degradation, but for functional modulation. The first protein is the mutant of leucine-rich repeat kinase 2 protein (LRRK2), whose expression in neurons causes abnormal neurite process and nuclear condensation, indicating neuronal toxicity, frequently seen in Parkinson’s disease (Lim et al. 2019). WSB1 promotes ubiquitylation of LRRK2 through the K27 and K29 linkages, which contributes to the formation of LRRK2 aggregation for neuronal protection (Nucifora et al. 2016). Therefore, WSB1 knockdown exhibits enhanced neuronal toxicity with decreased protein aggregation in LRRK2 mutant Drosophila model (Nucifora et al. 2016). The second protein is thyroid hormone-activating enzyme type II iodothyronine deiodinase (D2). The WD-40 propeller of WSB-1 is capable of interacting with an 18-amino-acid loop in D2 to cause D2 polyubiquitylation. Such ubiquitylated D2 subsequently induces parathyroid hormone-related peptide (PTHrP) to regulate chondrocyte differentiation (Dentice et al. 2005).
16.2.5 Other SOCS Box Proteins
Rab40 was reported to interact with Elongin B/C and Cul-5 at the Golgi apparatus of Xenopus, forming a ubiquitin ligase complex to regulate the ubiquitylation and localization of the Rap2 GTPase, thereby playing an essential role in the non-canonical Wnt pathway (Kamura et al. 2001). MUF1 was shown to have a ubiquitin ligase activity after complexing with the Cul-5/Elongin BC complex. However, its specific substrate has not been identified (Kamura et al. 2001). Finally, in response to UV irradiation, Elongin A was reported to bind with Elongin B/C to form an Elongin ABC complex and then assembles with the Cul-5 and SAG module to promote ubiquitylation and degradation of the large subunit of RNA polymerase II B1 (Rpb1) (Yasukawa et al. 2008). VHL, a SOCS box-like protein, but lacking the C-terminal sequence of the SOCS box, is well-known to interact with endogenous Elongin B/C, Cul-2, and RBX1 to form an active E3 ubiquitin ligase for HIF-1α degradation (Kamura et al. 2004). Our earlier study showed that VHL is also associated with SAG/Cul-5, particularly under hypoxic conditions, to facilitate HIF-1α degradation to keep HIF-1α levels under control (Tan et al. 2008). A summary of the CRL-5 receptor components and their corresponding substrates is listed in Table 16.1.
16.3 Cul-5 Neddylation and CRL-5 Activation
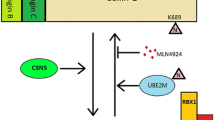
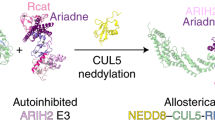
It is well-known that activation of CRL-5 requires the attachment of NEDD8, a ubiquitin-like protein onto the Lys724 residue located at the C-terminus of Cul-5, in a process known as neddylation (Duda et al. 2008). More specifically, NEDD8 is first activated by NAE E1 and then transferred from the active site Cys of NAE onto the active site Cys of UBE2F. NEDD8-loaded UBE2F is then recognized by SAG E3 on the same surface recognized by NAE, to catalyze the transfer of the NEDD8 molecule from UBE2F onto the Lys724 residue at the winged-helix B motif of Cul-5. Such modulation of Cul-5 leads to a conformation change of the cullin-RING interface and results in the catalytically active CRL state (Rabut and Peter 2008) (Fig. 16.2). Conversely, inactivation of cullins occurs through removal of NEDD8 from cullins by a process known as deneddylation via the COP9 signalosome complex (CSN) (Lyapina et al. 2001). Cullin neddylation activates Cullin-E3 ligase activity via several mechanisms. First, it prevents the inhibitory binding of cullin-associated neddylation-dissociated 1 (CAND1) to cullins (Duda et al. 2008); second, it induces the conformational change at the cullin-RBX interface, allowing the ubiquitin-loaded E2s to move closer to the acceptor lysine residue of substrate proteins; third, NEDD8 is capable of promoting the formation of higher-order cullin-RBX complexes to increase the catalytic efficiency of some Cullin-E3 ligases (Soucy et al. 2009). Mammalian cells contain a single neddylation E1, a heterodimer of catalytic subunit UBA3/NAEβ and regulatory subunit APPBP1/NAE1, two neddylation E2s, UBE2M (also known as UBC12) and UBE2F, and several neddylation E3s, mainly consisting of a few RING domain-containing proteins, such as RBX1 and SAG (Zhou et al. 2018a).
Neddylation activation of Cul-5 and SAG-Cul5 E3 ligase activity. CRL-5 activation requires Cul-5 neddylation, which is catalyzed by neddylation E1 NAE, neddylation E2 UBE2F, and neddylation E3 SAG with Cul-5 as the substrate (left panel). Upon Cul-5 neddylation, SAG binds to Cul-5 via its N-terminus and ubiquitylation E2, UBE2C and UBE2S, via its RING domain, and acts as a ubiquitylation E3 to catalyze the ubiquitin transfer from E2 to a substrate and form polyubiquitylation chain via the K11 linkage. On the other hand, SAG complexes with Cul-1 to bind with E2 UBCH5C to catalyze polyubiquitylation of substrate via the K48 linkage (right panel). Thus, SAG is a dual E3 for both neddylation and ubiquitylation; and SAG binds to two types of E2 responsible for polyubiquitylation chains via the K11 and K48 linkage, respectively
16.3.1 Neddylation E2: UBE2F and UBE2M
Two neddylation E2s have distinct features by structural comparison between UBE2F and UBE2M. Both E2s bind to ubiquitin-fold domain and UBA3 hydrophobic groove of E1 through its core domain and N-terminal motif, respectively (Huang et al. 2004). Biochemically, these two E2s have a certain degree of binding selectivity; UBE2M couples with RBX1 to neddylate Cul1-Cul4, whereas UBE2F is relatively specific for SAG to promote Cul-5 neddylation (Huang et al. 2009). In both E2s, the N-terminal methionine is acetylated, which facilitates their respective binding to the PONY domain pocket of DCNL, another neddylation E3, thus enhancing the efficiency of cullin neddylation (Huang et al. 2009; Monda et al. 2013). Biologically, in NIH3T3 cells with knockdown of UBE2M, but not of UBE2F, suppress cell growth (Huang et al. 2009). While both E2s are recruited to DNA damage sites in response to IR or other DNA-damaging agents, only depletion of UBE2M, but not UBE2F, sensitizes cells to DNA damaging agents (Brown et al. 2015; Cukras et al. 2014).
We recently found that UBE2F is subjected to negative regulation by its family member, UBE2M (Zhou et al. 2018b). Specifically, UBE2M is a stress-inducible protein. Upon transcriptional induction by hypoxia or mitogen, UBE2M acts as a dual E2 for both neddylation and ubiquitylation. It promotes Cul-3 neddylation to activate CRL3Keap1 E3 ligase; it also serves as E2 for ubiquitylation E3 Parkin/DJ-1, leading to ubiquitylation and degradation of UBE2F (Zhou et al. 2018b).
16.3.2 Neddylation and Ubiquitylation E3: SAG
SAG, also known as RBX2, ROC2, or RNF7, was first cloned by us as a redox-inducible protein (Duan et al. 1999), and it was later found to be the second family member of RBX RING protein with ligase activity when complexed with other CRL components (Swaroop et al. 2000). Human SAG gene consists of four exons and three introns, which is mapped onto chromosome 3q22-24 with three splicing variants and two family pseudogenes (Swaroop et al. 2001). Structurally, SAG encodes a protein of 113 amino acids, with 12 cysteine residues and a zinc-binding C3H2C3 RING finger domain at the C-terminus (Duan et al. 1999). This characteristic of SAG structure confers its dual functionalities. First, when acting alone, SAG has a nonenzymatic antioxidant activity to scavenge ROS at the expense of self-oligomerization via formation of intra- and intermolecule disulfide bonds (Swaroop et al. 2001). Second, when complex with other components of CRL, SAG possesses an intrinsic ubiquitin E3 ligase activity (Tan et al. 2010). Furthermore, when cooperating with UBE2F, SAG acts as a neddylation E3 to neddylate Cul-5 and activates CRL-5 (Huang et al. 2009).
16.3.2.1 Antioxidant Role of SAG
SAG has well-defined antioxidant activity by generating thiol/disulfide redox buffer and chelating metals with enriched cysteine residues. SAG forms oligomers by hydrogen peroxide, reversible by antioxidant dithiothreitol (DTT), or by the alkylating agent NEM, indicating that SAG oligomerization is induced by the formation of inter- or intramolecular disulfide bonds (Swaroop et al. 2001). SAG also binds to metal ions, such as zinc, iron, or copper, to inhibit the copper-induced lipid peroxidation in vitro (Duan et al. 1999; Swaroop et al. 2001). Importantly, the antioxidant protective role of SAG can also be expanded to the in vivo mouse models. Injection of SAG-expression adenovirus or purified cell-penetrable SAG protein (Tat-SAG) attenuates ischemia/oxidative stress-induced damages in the mouse brain (Kim et al. 2010; Yang et al. 2001).
16.3.2.2 SAG as a RING Component of SCF E3 Ubiquitin Ligase
Our earlier study found that purified SAG binds to Cul-1, and SAG-Cul1 complex promotes the formation of high molecular weight smears in a ligase reaction mixture containing ubiquitin, E1, E2, and ATP, indicative of polyubiquitylation in the E1- and E2-dependent manner (Swaroop et al. 2000). The polyubiquitylation activity of SAG-Cul1 is dependent on the RING structure of SAG, since SAG RING mutants completely abrogate this reaction. Furthermore, like the RBX1-Cul1 complex, the SAG-Cul1 complex promotes polyubiquitylation of phosphorylated IκB in an in vitro ubiquitylation assay (Tan et al. 2010). Under overexpressed conditions, both RBX1 and SAG are capable of binding to six members of the cullin proteins (Cul1-3, Cul-4A-B, and Cul-5) (Ohta et al. 1999), whereas under physiological conditions, RBX1 is preferentially associated with Cullin 1–4, whereas SAG is selectively to interact with Cul-5 (Kamura et al. 2004; Huang et al. 2009). These two RBX family members, along with seven cullins, many adaptors, and substrate receptors, assemble into numerous ubiquitin ligase complexes (Cardozo and Pagano 2004; Jia and Sun 2009), which are responsible for ubiquitylation of 20% cellular proteins doomed for proteasome degradation (Soucy et al. 2009).
Most recently, we found that SAG is a dual ubiquitin E3, capable of promoting substrate polyubiquitylation via both K48 and K11 linkages. Specifically, on one hand, SAG binds to UBCH5C E2 and couples with Cul-1 to promote polyubiquitylation of the substrates via the K48 linkage. On the other hand, SAG binds to UBE2C/2S E2 and couples with Cul-5 to promote polyubiquitylation of the substrates via the K11 linkage (Kuang et al. 2016) (Fig. 16.2), even though both are for targeted degradation via proteasome system. In contrast, RBX1, another SAG family member, is only capable of binding with various K48-linked E2s, but not K11-linked UBE2C/2S, for substrate polyubiquitylation via the K48 linkage (Kuang et al. 2016). This distinct difference in biochemical feature may explain why SAG and RBX1 are functionally non-redundant during mouse development (Tan et al. 2009, 2011).
16.3.2.3 SAG as a Neddylation E3 Ligase
NEDD8 E3 ligases catalyze the process of transferring the NEDD8 from the E2 conjugating enzyme onto the target substrate. Until now, all reported NEDD8 E3 ligases are capable of functioning as ubiquitin E3s (Enchev et al. 2015), with a majority of belonging to the RING domain-containing subclass (Deshaies and Joazeiro 2009). Two well-known NEDD8 E3 ligases are RBX1 and SAG. As mentioned before, RBX1 specifically couples with UBE2M E2 to promote neddylation of cullins 1–4, while SAG couples with UBE2F E2 to promote Cul-5 neddylation (Huang et al. 2009). Both the E3 RING domains and the UFD of NAE bind to the same surface on the NEDD8 E2 enzymes, resulting in toggling of relative affinities to ensure the unidirectionality of the neddylation process (Eletr et al. 2005). The interaction between the E3 RING domain and E2-bound NEDD8 is required for catalyzing the transfer of the NEDD8 molecule into a Lys residue or the N terminus of the target substrate. Furthermore, cullin neddylation by SAG or RBX1 is aided by E3 DCNLs (Enchev et al. 2015). A summary of SAG acting as a dual E3 for ubiquitylation and neddylation is shown in Fig. 16.2.
16.4 Substrates of CRL-5 E3 Ligase with Unknown Substrate Receptors
An array of CRL-5 substrates with corresponding receptor proteins are listed in Table 16.1. Several CRL-5 substrates with significant biological functions, but unknown corresponding receptors, are reviewed in the following.
16.4.1 DEPTOR
DEPTOR is a naturally occurring inhibitor of mTORC1 and mTORC2 through a direct binding to mTOR and acts as a tumor suppressor in a context-dependent manner (Peterson et al. 2009). Our earlier study, along with two other groups, showed that DEPTOR accumulates in starvation conditions and contributes to autophagy induction, whereas upon stimulation by serum or growth factors, DEPTOR is phosphorylated and recognized by βTrCP, followed by ubiquitylation and degradation by SCF E3 ligase (also known as CRL1) to ensure mTOR activation (Zhao et al. 2011; Duan et al. 2011; Gao et al. 2011). Our later study showed that SAG can complex with either Cul-1 or Cul-5 to promote ubiquitylation and degradation of DEPTOR (Tan et al. 2016). Importantly, negative regulation of DEPTOR by SAG has biological consequences. In cell culture model, SAG knockdown suppresses growth, survival, and migration of human prostate cancer cells via inactivation of the PI3K/AKT/mTOR signaling axis through DEPTOR accumulation; whereas in a mouse prostate cancer model, Sag deletion significantly inhibits prostate tumorigenesis triggered by Pten loss as a result of suppressed proliferation due to DEPTOR accumulation (Tan et al. 2016). Furthermore, another group has shown that the Cul5/Elongin B complex promotes ubiquitylation and degradation of DEPTOR to negatively regulate autophagy under nutrient-rich conditions (Antonioli et al. 2014).
16.4.2 Heat Shock Protein 90 (Hsp90) Client Proteins
Hsp90, a molecular chaperone with approximately 350 client proteins, is responsible for the correct folding of proteins, which facilitates proteins to attain their proper stabilization and activity (Taipale et al. 2012). Two studies showed that upon treatment of human cancer cells with the clinical HSP90 inhibitor 17-AAG, Cul5/SAG E3 are actively involved or required for degradation of several HSP90 clients, including ErbB2, HIF-1α, BRAFV600E, AKT, and CDK4 (Ehrlich et al. 2009; Samant et al. 2014). Thus, it appears that Cul-5 E3 plays a role in regulation of the cellular response to HSP90 inhibition.
16.4.3 NOXA
NOXA is a pro-apoptotic member of Bcl-2 protein family, which can form hetero- or homodimers and act as pro-apoptotic regulator (Oda et al. 2000). Our recent studies showed that Cul5/SAG E3 targets NOXA for degradation to protect cancer cells from apoptosis, ensuring an apoptosis-escaping mechanism in lung cancer cells. Specifically, UBE2F NEDD8-E2 incorporates with SAG E3 to induce Cul-5 neddylation, leading to activation of CRL-5 E3 to promote NOXA polyubiquitylation via K11 linkage for proteasomal degradation (Jia et al. 2010; Zhou et al. 2017). Most recently, we further found that upon stress stimuli (e.g., hypoxia), UBE2M was induced and then formed a complex with Parkin/DJ-1 to promote UBE2F ubiquitylation and degradation, leading to Cul5/E3 inactivation and subsequent NOXA accumulation for apoptosis induction (Zhou et al. 2018b).
16.4.4 TRAF6
A recent study showed that Cul-5 directly binds to TRAF6 via the C-terminal domain of Cul-5 and the TRAF-C domain of TRAF6 and promotes TRAF6 polyubiquitylation via the K63 linkage in response to lipopolysaccharide (LPS) stimulation (Zhu et al. 2016). While homozygous deletion of Cul-5 is embryonic lethal, heterozygous Cul-5 deletion improves mouse survival and reduces proinflammatory cytokine production in response to LPS challenge due to reduced activation of NF-κB and MAPK signals (Zhu et al. 2016). Given that TRAF6 is an intrinsic E3 ligase, capable of self-poly-ubiquitylation, it remains to be determined whether TRAF6 is indeed a true Cul-5 substrate without involving a substrate receptor protein (Zhu et al. 2016).
Finally, the neddylated Cul5/SAG complex was shown to interact with and surprisingly enhances rather than inhibits the E3 ligase activity of TRIAD1 (two RING finger and double RING finger linked) (Kelsall et al. 2013), a distinct class of E3 ubiquitin ligases implicated in the process of hematopoiesis, mainly inhibiting myeloid colony formation (Marteijn et al. 2005).
16.5 Virus-Mediated Hijacking of CRL-5
Several strains of human virus were reported to hijack CRL-5 to promote ubiquitylation and degradation of a variety of host antiviral proteins. This unique feature makes CRL-5 E3 as a promising antivirus target for drug discovery efforts.
16.5.1 Human Immunodeficiency Virus-1 (HIV-1)
Apolipoprotein B editing complex 3G (APOBEC3G/A3G) is a potent anti-retroviral cytidine deaminase with a broad antiviral activity by inducing C to U mutations in the viral minus DNA strand during reverse transcription, which causes the deleterious G to A mutations in the coding strand (Suspene et al. 2004). To overcome host antiviral protective system, the HIV Vif protein hijacks CRL-5 to promote ubiquitylation and degradation of A3G (Sheehy et al. 2003). The BC box and SOCS box of Vif are required for the interaction with Elongin B/C and Cul-5, respectively. The knockdown of SAG, but not its family member RBX1, impairs Vif-induced A3G degradation (Wang et al. 2015). Moreover, neddylation of Cul-5 by UBE2F/SAG is required for Vif-mediated degradation of A3G, since pharmacological inhibition of the NEDD8 E1 with MLN4924 or knockdown of UBE2F bypasses the effect of Vif, restoring the restriction potential of A3G (Stanley et al. 2012). A non-NEDD8ylatable mutant Cul-5(m) was also shown to inhibit Vif-induced ubiquitination and degradation of A3G (Yu et al. 2003). Notably, A3G without ubiquitylation is still degraded in a Vif-dependent manner, suggesting that the polyubiquitylation of Vif, rather than polyubiquitylation of AG3, serves as a vehicle to transport A3G into proteasomes for degradation (Dang et al. 2008). Another antiviral factor, APOBEC3F/A3F, was also degraded by HIV-1 Vif via hijacking CRL-5 (Liu et al. 2005).
16.5.2 Human Adenoviruses (HAdV)
E4orf6 is 34 kDa product from open reading frame 6 of human adenovirus early region 4 (E4) with three BC boxes. E4orf6 cooperates with the viral E1B55K protein product to form an E3 ubiquitin ligase with Cul-5 to reduce the level of the p53 via the proteasome pathway (Querido et al. 2001; Steegenga et al. 1998), by triggering Cul-5 localization from the cytoplasm to the nucleus and CRL-5 activation via facilitating neddylation (Guo et al. 2019). Furthermore, as a consequence of adenovirus infection, E4orf6/E1B55K-Cul5 complex was shown to promote the degradation of DNA ligase IV, an enzyme that plays a pivotal role in repairing of double-stranded DNA breaks (DSBs) by performing the joining step of the nonhomologous end-joining DNA repair system (NHEJ) (Baker et al. 2007). The E4orf6/E1B55K-Cul5 E3 ligase also promotes the degradation of Mre11, a member of the MRN DNA repair complex (Stracker et al. 2002). Furthermore, the de novo-expressed, preassembled capsid proteins and Rep52 are also degraded by E4orf6/E1B55K-Cul5 E3 (Nayak et al. 2008), and this degradative activity of E4Orf6 can be overcome by virus-associated RNA, thereby increasing the capsid proteins and Rep52 to the levels necessary for efficient virus production (Nayak and Pintel 2007). Finally, the E4orf6/E1B55K-Cul5 E3 ligase complex is involved in the degradation of α3, a component of integrin α3β1, which plays an important role in the regulation of cellular adhesion through the binding with a variety of extracellular matrix substrates, including bronectin, collagen, vitronectin, and laminins, thereby playing an important role in virus spread (Dallaire et al. 2009).
16.5.3 Epstein-Barr Virus (EBV)
EBV is a human γ-herpesvirus that is associated with several B cell and epithelial cell malignancies. BZLF1 (known as Zta, EB1, or ZEBRA) is a transcriptional transactivator that promotes an EBV lytic cycle cascade by inducing EBV early gene expression (Chevallier-Greco et al. 1986). Importantly, BZLF1 couples with Cul-5 to form an active ubiquitin ligase to promote ubiquitylation and degradation of p53, a required process for efficient viral propagation in the lytic replication stage (Sato et al. 2009).
16.5.4 Kaposi’s Sarcoma-Associated Herpesvirus (KSHV)
The KSHV-encoded latency-associated nuclear antigen (LANA) complex was initially identified as a DNA binding, nuclear transcription factor that contributes to KSHV latent replication and regulates virus latency. LANA contains a putative SOCS box and can form a complex with Elongin B/C and Cul-5 for ubiquitination and degradation of pVHL and p53 tumor suppressor proteins (Cai et al. 2006).
16.5.5 Murid Herpesvirus-4 (MuHV-4)
MuHV-4 is a gamma herpesvirus that is genetically related to the human pathogens EBV and KSHV (Simas and Efstathiou 1998). The latency-associated protein ORF73 encoded by MuHV-4 is able to interact with Elongin C and Cul-5 to reconstitute an active E3 ubiquitin ligase to target the NF-κB family member p65/RelA for polyubiquitylation and subsequent proteasomal degradation. Such viral inhibition of NF-κB activity is critical for the establishment of a propitious environment for the maintenance of latent infection and progression of KSHV-associated tumors (Rodrigues et al. 2009).
16.6 The Physiological Role of Cul-5 in Human Cancers
The role of CRL-5 components in human cancers has been extensively reviewed. For example, the SOCS family members are extensively involved in inflammation and cancer, largely acting in suppression of proliferation (Inagaki-Ohara et al. 2013; Jiang et al. 2017). The RING component SAG is largely oncogenic (Sun and Li 2013), required for lung tumorigenesis triggered by Kras activation (Li et al. 2014) or for prostate tumorigenesis triggered by Pten loss (Tan et al. 2016), but it is tumor suppressive in skin tumorigenesis, triggered by Kras activation (Xie et al. 2015). This book chapter will only focus on the potential role of Cul-5 in human cancers.
Most studies imply that Cul-5 exerts a tumor suppressive role, as evidenced mainly by frequently downregulation of Cul-5 in various human cancers (Kalla et al. 2007; Fay et al. 2003; Xu et al. 2012; Devor et al. 2016; Tapia-Laliena et al. 2019). Specifically, in a cancer profiling array study, Cul-5 expression was approximately 2.2-fold lower in the breast cancer tissues versus the matched normal tissues (Fay et al. 2003). Ectopic Cul-5 overexpression significantly suppressed the proliferation of breast cancer cells (Burnatowska-Hledin et al. 2004), by inhibiting MAPK phosphorylation to reduce nuclear localization of estrogen receptor ER, eventually leading to suppression of growth of estrogen-dependent cells (Johnson et al. 2007). Conversely, inhibition of Cul-5 by microRNA-19a and microRNA-19b significantly promoted proliferation and invasion of cervical carcinoma cells and gastric cancer cells (Xu et al. 2012; Zhu et al. 2019). The siRNA-based Cul-5 knockdown caused centriole overduplication and mitotic errors and also induced structural chromosomal damage in renal cell carcinoma (Tapia-Laliena et al. 2019). Furthermore, an immunohistochemistry staining study showed that Cul-5 expression is frequently lower in renal cell carcinoma and a reduced Cul-5 expression or Cul-5 deletion is associated with a significantly worse overall patient survival by the analysis of The Cancer Genome Atlas (TCGA) database (https://tcga-data.nci.nih.gov/tcga) (Tapia-Laliena et al. 2019). In small cell lung cancers (SCLC), genetic deletion of Cul-5 or SOCS3 by CRISPR/Ccas9 impaired CRL-5-mediated degradation of integrin β1, leading to stabilization of integrin β1 to activate the downstream focal adhesion kinase/SRC (FAK/SRC) signaling, eventually driving growth and metastasis of SCLC (Zhao et al. 2019). Moreover, low expression of Cul-5 and SOCS3 and relatively high expression of integrin β1 are significantly associated with worse patient survival, suggesting Cul-5 is a tumor suppressor (Zhao et al. 2019). However, in non-small lung cancers (NSCLC), CRL-5 specifically targets NOXA for polyubiquitylation via the K11 linkage and proteasomal degradation to inhibit apoptosis and increase the survival of lung cancer cells (Zhou et al. 2017). Furthermore, elevated expression of Cul-5, coupled with low expression of NOXA, was found to predict poor patient survival in NSCLC patients, suggesting Cul-5 has oncogenic activity (Zhou et al. 2017).
To better understand the expression pattern and prognostic value of Cul-5 in human cancers, we examined Cul-5 expression in 32 types of human cancers using the UALCAN, an easy-to-use, interactive web portal to perform in-depth analyses of TCGA gene expression data (http://ualcan.path.uab.edu/) (Chandrashekar et al. 2017). Notably, compared to normal tissues, Cul-5 mRNA is upregulated in four types, but downregulated in nine types of human cancers (Fig. 16.3). Kaplan-Meier analysis revealed that higher levels of Cul-5 mRNA are associated with a worse patient survival for liver hepatocellular carcinoma and kidney chromophobe, but a better patient survival for kidney renal clear cell carcinoma, prostate adenocarcinoma, and rectum adenocarcinoma (Fig. 16.4). We further analyzed the significance of Cul-5 protein levels in human cancers using the database of the Human Protein Atlas (https://www.proteinatlas.org). Notably, Cul-5 staining is mainly detected at the moderate levels in the moderate cytoplasm in most of cancer tissues, and in more than 50% cases of lymphomas, melanomas, and lung, ovarian, and cervical cancers, the Cul-5 staining is weak or negative (Fig. 16.5a). At the protein levels, only one type of human cancer showed a statistically significant correlation between the expression and the patient survival. That is, the higher Cul-5 staining predicts a better prognosis in renal cancer patients (Fig. 16.5b).
Expression of Cul-5 mRNA in human cancer tissues. The search of the UALCAN database (http://ualcan.path.uab.edu) revealed that Cul-5 mRNA is upregulated in four types of human cancers and downregulated in nine types of human cancers. The abbreviations are as follows: CHOL cholangiocarcinoma, COAD colon adenocarcinoma, LIHC liver hepatocellular carcinoma, LUSC lung squamous cell carcinoma, GBM glioblastoma multiforme, KIRC kidney renal clear cell carcinoma, KTRP kidney renal papillary cell carcinoma, PRAD prostate adenocarcinoma, READ rectum adenocarcinoma, TGCT testicular germ cell tumors, THYM thymoma, THCA thyroid carcinoma, UCEC uterine corpus endometrial carcinoma
Cul-5 mRNA levels in various human cancer tissues associated with patient survival. The UALCAN (http://ualcan.path.uab.edu) search also revealed that elevated levels of Cul-5 mRNA are associated with the worse survival for kidney chromophobe and liver hepatocellular carcinoma, but the better survival for kidney renal clear cell carcinoma, prostate adenocarcinoma, and rectum adenocarcinoma. KICH kidney chromophobe, LIHC liver hepatocellular carcinoma, KIRC kidney renal clear cell carcinoma, PRAD prostate adenocarcinoma, READ rectum adenocarcinoma
The Cul-5 protein immunostaining in various human cancer tissues and their association with patient survival. The Human Protein Atlas data (https://www.proteinatlas.org) search revealed that most cancer tissues have moderate cytoplasmic immunoreactivity of Cul-5, whereas most cases of lymphomas, melanomas, and lung, ovarian, and cervical cancers are stained weakly or negative (a). High levels of Cul-5 protein are associated with favorable prognosis for renal cancer (p = 0.000011) (b). COCA colorectal cancer, ENCA endometrial cancer, HNCA head and neck cancer, TECA testis cancer, THCA thyroid cancer, URCA urothelial cancer, STCA stomach cancer, BRCA breast cancer, PRCA prostate cancer, SKCA skin cancer, PACA pancreatic cancer, CARC carcinoid, GLIO glioma, RECA renal cancer, MELA melanoma, LICA liver cancer, OVCA ovarian cancer, CECA cervical cancer, LUCA lung cancer, LYMP lymphoma
16.7 Future Perspectives
The UBE2F-SAG-Cul-5 axis is exclusively presented in metazoans by phylogenetic analyses, whereas the UBE2M-RBX1-Cul1-4 axis appears in all eukaryotes; the UBE2F-SAG-Cul5 is, therefore, regarded as a distinct pathway in regulation of CRL-5 E3 ligase activity (Huang et al. 2009). CRL-5 is likely involved in regulation of many biological processes, given that a variety of important signal molecules are its substrates (Table 16.1) for proteasomal degradation. However, many of reported studies were conducted in the cell culture setting under overexpressed conditions; the physiological relevance or significance is questionable. Furthermore, total Cul-5 knockout is embryonic lethal (Zhu et al. 2016), whereas no study on tissue specific Cul-5 knockout has been reported. The study of Cul-5 involvement in tumorigenesis under physiological setting is, therefore, lacking. The future studies focusing on the activity and functions of CRL-5 E3 ligase should be directed in the following aspects.
16.7.1 Generation of Conditional Mouse Models to Study the Role of Cul-5 in Tumorigenesis
It is urgent to generate and characterize conditional Cul-5 knockout mice model to study its role in tumorigenesis in various organs triggered by oncogene activation or tumor suppressor inactivation induced by chemical carcinogens or radiation. Given all the data collected from human clinical tumor tissues with regard to Cul-5 expression levels and prognosis correlation only provide an association with the particular type(s) of cancer without elucidation of any cause-consequence relationship, these mouse studies will provide under the physiological settings whether Cul-5 is a cooperative oncogene or tumor suppressor in a tissue-/context-dependent manner.
16.7.2 Generation of Conditional Mouse Models to Study the Roles of Each Receptor Subunit in Tumorigenesis
Among the substrate receptor families, there are many members (Fig. 16.1), and each member is capable of complexing with the rest three components to constitute a variety of CRL-5 E3s. This large array of E3s must play a variety of functions under certain physiological or pathological conditions or in response to environmental stresses, which has not been addressed in many current studies. One feasible approach is to generate conditional knockout mouse models for each receptor family member to fully understand their functions under the physiological settings or during tumorigenesis, triggered by either oncogene activation or tumor suppressor inactivation or by environmental insults.
16.7.3 Identification of Additional Substrates of CRL-5 E3s
Many members of CRL-5 receptors are the orphan receptor without corresponding substrates identified and characterized, which limited our full understanding of CRL-5 functions. Thus, the use of various current technologies and methodologies to identify and characterize additional downstream ubiquitin substrates, especially under biological significant and physiological relevant settings, is much needed.
16.7.4 Dynamic Regulation of CRL-5 E3 Ligase Activity
The enzymatic activity of each of CRLs can be regulated at the level of the subunit assembly (see earlier chapters of this book). The regulation of CRL assembly and activity is a very dynamic and precise process. The expression levels of some subunits fluctuate at particular cellular or developmental stages (e.g., at the different phases of cell cycle), which dictate the assembly of a given CRL (Bennett et al. 2010). Also the assembly of CRL is dependent upon CAND1 that can act as a subunit exchange factor (Bennett et al. 2010). Furthermore, the kinetics of CSN association with CRLs and the subsequent deneddylation are also subject to precise regulation. The thorough elucidation of these mechanism regulations, particularly for CRL-5, would lead to a better understanding of its biochemical activity and the consequent biological functions under the physiological or pathological conditions.
16.7.5 CRL-5 as an Attractive Antivirus Drug Target
Several infectious viruses are capable of hijacking CRL-5 to degrade host antiviral proteins, eventually obtaining anti-host property to propagate in the host cells. Inhibition of CRL-5 by MLN4924, a small molecular inhibitor of cullin neddylation, leading to general inactivation of all CRLs (Soucy et al. 2009), indeed showed potent antivirus activity (Stanley et al. 2012; Guo et al. 2019; Becker et al. 2019; Hughes et al. 2015; Kraus et al. 2017; Le-Trilling et al. 2016). Since a general inhibition of all CRLs by MLN4924 likely has some cytotoxic effect, the discovery of selective CRL-5 inhibitors as a novel class of antivirus therapeutic drugs would be an ideal approach.
Abbreviations
- APOBEC3G:
-
Apolipoprotein B editing complex 3G
- APS:
-
Adapter protein with a pleckstrin homology and Src homology 2 domain
- AVP:
-
Arginine vasopressin
- CAND1:
-
Cullin-associated neddylation-dissociated 1
- CH:
-
Cullin homology
- CKB:
-
Cytosolic creatine kinase
- CR:
-
Cullin repeats
- CRL-5:
-
Cullin RING ligase 5
- CRLs:
-
Cullin-RING ligases
- CSN:
-
COP9 signalosome complex
- CTD:
-
Carboxyl-terminal domain
- Cul-5:
-
Cullin-5
- DCNL:
-
The defective in cullin neddylation protein-like proteins
- DSBs:
-
Double-stranded DNA breaks
- EBV:
-
Epstein-Barr virus
- ER:
-
Endoplasmic reticulum
- FAK:
-
Focal adhesion kinase
- HAdV:
-
Human adenoviruses
- HIF-α:
-
Hypoxia-inducible factor-α
- HIPK2:
-
Homeodomain-interacting protein kinase 2
- HIV-1:
-
Human immunodeficiency virus-1
- HSP90:
-
Heat shock protein 90
- ID2:
-
Inhibitor of DNA binding 2
- iNOS:
-
Inducible nitric oxide synthase
- IRS4:
-
Insulin receptor substrate 4
- JAK2:
-
Janus kinase 2
- KSHV:
-
Kaposi’s sarcoma-associated herpesvirus
- LANA:
-
Latency-associated nuclear antigen
- LPS:
-
Lipopolysaccharide
- LRRK2:
-
Leucine-rich repeat kinase 2
- MCK:
-
Mitochondrial creatine kinase
- MLL:
-
Mixed-lineage leukemia
- MuHV-4:
-
Murid herpesvirus-4
- NAE:
-
NEDD8-activating enzyme
- NHEJ:
-
Nonhomologous end-joining DNA repair system
- NSCLC:
-
Non-small lung cancers
- NTD:
-
Amino-terminal domain
- PTHrP:
-
Parathyroid hormone-related peptide
- RBX2:
-
RING-box protein 2
- ROS:
-
Reactive oxygen species
- Rpb1:
-
RNA polymerase II B1
- SAG:
-
Sensitive to apoptosis gene
- SCLC:
-
Small cell lung cancers
- SH2:
-
Src homology 2
- SOCS:
-
Suppressor of cytokine signaling
- TCGA:
-
The Cancer Genome Atlas
- TNF-R2:
-
Tumor necrosis factor receptor 2
- TRIAD1:
-
Two RING finger and double RING finger linked 1
- VACM-1:
-
Vasopressin-activated calcium-mobilizing protein 1
References
Andresen CA, Smedegaard S, Sylvestersen KB, Svensson C, Iglesias-Gato D, Cazzamali G, Nielsen TK, Nielsen ML, Flores-Morales A (2014) Protein interaction screening for the ankyrin repeats and suppressor of cytokine signaling (SOCS) box (ASB) family identify Asb11 as a novel endoplasmic reticulum resident ubiquitin ligase. J Biol Chem 289(4):2043–2054
Antonioli M, Albiero F, Nazio F, Vescovo T, Perdomo AB, Corazzari M, Marsella C, Piselli P, Gretzmeier C, Dengjel J, Cecconi F, Piacentini M, Fimia GM (2014) AMBRA1 interplay with cullin E3 ubiquitin ligases regulates autophagy dynamics. Dev Cell 31(6):734–746
Baker A, Rohleder KJ, Hanakahi LA, Ketner G (2007) Adenovirus E4 34k and E1b 55k oncoproteins target host DNA ligase IV for proteasomal degradation. J Virol 81(13):7034–7040
Becker T, Le-Trilling VTK, Trilling M (2019) Cellular cullin RING ubiquitin ligases: druggable host dependency factors of cytomegaloviruses. Int J Mol Sci 20(7)
Bennett EJ, Rush J, Gygi SP, Harper JW (2010) Dynamics of cullin-RING ubiquitin ligase network revealed by systematic quantitative proteomics. Cell 143(6):951–965
Brown JS, Lukashchuk N, Sczaniecka-Clift M, Britton S, le Sage C, Calsou P, Beli P, Galanty Y, Jackson SP (2015) Neddylation promotes ubiquitylation and release of Ku from DNA-damage sites. Cell Rep 11(5):704–714
Burande CF, Heuze ML, Lamsoul I, Monsarrat B, Uttenweiler-Joseph S, Lutz PG (2009) A label-free quantitative proteomics strategy to identify E3 ubiquitin ligase substrates targeted to proteasome degradation. Mol Cell Proteomics 8(7):1719–1727
Burnatowska-Hledin MA, Spielman WS, Smith WL, Shi P, Meyer JM, Dewitt DL (1995) Expression cloning of an AVP-activated, calcium-mobilizing receptor from rabbit kidney medulla. Am J Phys 268(6 Pt 2):F1198–F1210
Burnatowska-Hledin MA, Kossoris JB, Van Dort CJ, Shearer RL, Zhao P, Murrey DA, Abbott JL, Kan CE, Barney CC (2004) T47D breast cancer cell growth is inhibited by expression of VACM-1, a cul-5 gene. Biochem Biophys Res Commun 319(3):817–825
Byrd PJ, Stankovic T, McConville CM, Smith AD, Cooper PR, Taylor AM (1997) Identification and analysis of expression of human VACM-1, a cullin gene family member located on chromosome 11q22-23. Genome Res 7(1):71–75
Cai QL, Knight JS, Verma SC, Zald P, Robertson ES (2006) EC5S ubiquitin complex is recruited by KSHV latent antigen LANA for degradation of the VHL and p53 tumor suppressors. PLoS Pathog 2(10):e116
Cao J, Wang Y, Dong R, Lin G, Zhang N, Wang J, Lin N, Gu Y, Ding L, Ying M, He Q, Yang B (2015) Hypoxia-induced WSB1 promotes the metastatic potential of osteosarcoma cells. Cancer Res 75(22):4839–4851
Cardozo T, Pagano M (2004) The SCF ubiquitin ligase: insights into a molecular machine. Nat Rev Mol Cell Biol 5(9):739–751
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, Varambally S (2017) UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19(8):649–658
Chevallier-Greco A, Manet E, Chavrier P, Mosnier C, Daillie J, Sergeant A (1986) Both Epstein-Barr virus (EBV)-encoded trans-acting factors, EB1 and EB2, are required to activate transcription from an EBV early promoter. EMBO J 5(12):3243–3249
Choi DW, Seo YM, Kim EA, Sung KS, Ahn JW, Park SJ, Lee SR, Choi CY (2008) Ubiquitination and degradation of homeodomain-interacting protein kinase 2 by WD40 repeat/SOCS box protein WSB-1. J Biol Chem 283(8):4682–4689
Choi YB, Son M, Park M, Shin J, Yun Y (2010) SOCS-6 negatively regulates T cell activation through targeting p56lck to proteasomal degradation. J Biol Chem 285(10):7271–7280
Chung AS, Guan YJ, Yuan ZL, Albina JE, Chin YE (2005) Ankyrin repeat and SOCS box 3 (ASB3) mediates ubiquitination and degradation of tumor necrosis factor receptor II. Mol Cell Biol 25(11):4716–4726
Cukras S, Morffy N, Ohn T, Kee Y (2014) Inactivating UBE2M impacts the DNA damage response and genome integrity involving multiple cullin ligases. PLoS One 9(7):e101844
Dallaire F, Blanchette P, Groitl P, Dobner T, Branton PE (2009) Identification of integrin alpha3 as a new substrate of the adenovirus E4orf6/E1B 55-kilodalton E3 ubiquitin ligase complex. J Virol 83(11):5329–5338
Dang Y, Siew LM, Zheng YH (2008) APOBEC3G is degraded by the proteasomal pathway in a Vif-dependent manner without being polyubiquitylated. J Biol Chem 283(19):13124–13131
De Sepulveda P, Ilangumaran S, Rottapel R (2000) Suppressor of cytokine signaling-1 inhibits VAV function through protein degradation. J Biol Chem 275(19):14005–14008
Debrincat MA, Zhang JG, Willson TA, Silke J, Connolly LM, Simpson RJ, Alexander WS, Nicola NA, Kile BT, Hilton DJ (2007) Ankyrin repeat and suppressors of cytokine signaling box protein asb-9 targets creatine kinase B for degradation. J Biol Chem 282(7):4728–4737
Dentice M, Bandyopadhyay A, Gereben B, Callebaut I, Christoffolete MA, Kim BW, Nissim S, Mornon JP, Zavacki AM, Zeold A, Capelo LP, Curcio-Morelli C, Ribeiro R, Harney JW, Tabin CJ, Bianco AC (2005) The Hedgehog-inducible ubiquitin ligase subunit WSB-1 modulates thyroid hormone activation and PTHrP secretion in the developing growth plate. Nat Cell Biol 7(7):698–705
Deshaies RJ, Joazeiro CA (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434
Devor EJ, Schickling BM, Reyes HD, Warrier A, Lindsay B, Goodheart MJ, Santillan DA, Leslie KK (2016) Cullin-5, a ubiquitin ligase scaffold protein, is significantly underexpressed in endometrial adenocarcinomas and is a target of miR-182. Oncol Rep 35(4):2461–2465
Diks SH, Sartori MA, da Silva JL, Hillebrands RJ, Bink HHV, van Rooijen C, Brouwers A, Chitnis AB, Peppelenbosch MP, Zivkovic D (2008) d-Asb11 is an essential mediator of canonical Delta-Notch signalling. Nat Cell Biol 10(10):1190–1198
Duan H, Wang Y, Aviram M, Swaroop M, Loo JA, Bian J, Tian Y, Mueller T, Bisgaier CL, Sun Y (1999) SAG, a novel zinc RING finger protein that protects cells from apoptosis induced by redox agents. Mol Cell Biol 19(4):3145–3155
Duan S, Skaar JR, Kuchay S, Toschi A, Kanarek N, Ben-Neriah Y, Pagano M (2011) mTOR generates an auto-amplification loop by triggering the betaTrCP- and CK1alpha-dependent degradation of DEPTOR. Mol Cell 44(2):317–324
Duda DM, Borg LA, Scott DC, Hunt HW, Hammel M, Schulman BA (2008) Structural insights into NEDD8 activation of cullin-RING ligases: conformational control of conjugation. Cell 134(6):995–1006
Ehrlich ES, Wang T, Luo K, Xiao Z, Niewiadomska AM, Martinez T, Xu W, Neckers L, Yu XF (2009) Regulation of Hsp90 client proteins by a Cullin5-RING E3 ubiquitin ligase. Proc Natl Acad Sci U S A 106(48):20330–20335
Eletr ZM, Huang DT, Duda DM, Schulman BA, Kuhlman B (2005) E2 conjugating enzymes must disengage from their E1 enzymes before E3-dependent ubiquitin and ubiquitin-like transfer. Nat Struct Mol Biol 12(10):933–934
Enchev RI, Schulman BA, Peter M (2015) Protein neddylation: beyond cullin-RING ligases. Nat Rev Mol Cell Biol 16(1):30–44
Endo TA, Masuhara M, Yokouchi M, Suzuki R, Sakamoto H, Mitsui K, Matsumoto A, Tanimura S, Ohtsubo M, Misawa H, Miyazaki T, Leonor N, Taniguchi T, Fujita T, Kanakura Y, Komiya S, Yoshimura A (1997) A new protein containing an SH2 domain that inhibits JAK kinases. Nature 387(6636):921–924
Fay MJ, Longo KA, Karathanasis GA, Shope DM, Mandernach CJ, Leong JR, Hicks A, Pherson K, Husain A (2003) Analysis of CUL-5 expression in breast epithelial cells, breast cancer cell lines, normal tissues and tumor tissues. Mol Cancer 2:40
Frantsve J, Schwaller J, Sternberg DW, Kutok J, Gilliland DG (2001) Socs-1 inhibits TEL-JAK2-mediated transformation of hematopoietic cells through inhibition of JAK2 kinase activity and induction of proteasome-mediated degradation. Mol Cell Biol 21(10):3547–3557
Gao D, Inuzuka H, Tan MK, Fukushima H, Locasale JW, Liu P, Wan L, Zhai B, Chin YR, Shaik S, Lyssiotis CA, Gygi SP, Toker A, Cantley LC, Asara JM, Harper JW, Wei W (2011) mTOR drives its own activation via SCF(betaTrCP)-dependent degradation of the mTOR inhibitor DEPTOR. Mol Cell 44(2):290–303
Guo H, Shen S, Li Y, Bi R, Zhang N, Zheng W, Deng Y, Yang Y, Yu XF, Wang C, Wei W (2019) Adenovirus oncoprotein E4orf6 triggers Cullin5 neddylation to activate the CLR5 E3 ligase for p53 degradation. Biochem Biophys Res Commun 516(4):1242–1247
Heuze ML, Lamsoul I, Baldassarre M, Lad Y, Leveque S, Razinia Z, Moog-Lutz C, Calderwood DA, Lutz PG (2008) ASB2 targets filamins A and B to proteasomal degradation. Blood 112(13):5130–5140
Hilton DJ, Richardson RT, Alexander WS, Viney EM, Willson TA, Sprigg NS, Starr R, Nicholson SE, Metcalf D, Nicola NA (1998) Twenty proteins containing a C-terminal SOCS box form five structural classes. Proc Natl Acad Sci U S A 95(1):114–119
Huang DT, Miller DW, Mathew R, Cassell R, Holton JM, Roussel MF, Schulman BA (2004) A unique E1-E2 interaction required for optimal conjugation of the ubiquitin-like protein NEDD8. Nat Struct Mol Biol 11(10):927–935
Huang DT, Ayrault O, Hunt HW, Taherbhoy AM, Duda DM, Scott DC, Borg LA, Neale G, Murray PJ, Roussel MF, Schulman BA (2009) E2-RING expansion of the NEDD8 cascade confers specificity to cullin modification. Mol Cell 33(4):483–495
Hughes DJ, Wood JJ, Jackson BR, Baquero-Perez B, Whitehouse A (2015) NEDDylation is essential for Kaposi’s sarcoma-associated herpesvirus latency and lytic reactivation and represents a novel anti-KSHV target. PLoS Pathog 11(3):e1004771
Inagaki-Ohara K, Kondo T, Ito M, Yoshimura A (2013) SOCS, inflammation, and cancer. JAKSTAT 2(3):e24053
Jia L, Sun Y (2009) RBX1/ROC1-SCF E3 ubiquitin ligase is required for mouse embryogenesis and cancer cell survival. Cell Div 4:16
Jia L, Yang J, Hao X, Zheng M, He H, Xiong X, Xu L, Sun Y (2010) Validation of SAG/RBX2/ROC2 E3 ubiquitin ligase as an anticancer and radiosensitizing target. Clin Cancer Res 16(3):814–824
Jiang M, Zhang WW, Liu P, Yu W, Liu T, Yu J (2017) Dysregulation of SOCS-mediated negative feedback of cytokine signaling in carcinogenesis and its significance in cancer treatment. Front Immunol 8:70
Johnson AE, Le IP, Buchwalter A, Burnatowska-Hledin MA (2007) Estrogen-dependent growth and estrogen receptor (ER)-alpha concentration in T47D breast cancer cells are inhibited by VACM-1, a cul 5 gene. Mol Cell Biochem 301(1–2):13–20
Kalla C, Scheuermann MO, Kube I, Schlotter M, Mertens D, Dohner H, Stilgenbauer S, Lichter P (2007) Analysis of 11q22-q23 deletion target genes in B-cell chronic lymphocytic leukaemia: evidence for a pathogenic role of NPAT, CUL5, and PPP2R1B. Eur J Cancer 43(8):1328–1335
Kamio M, Yoshida T, Ogata H, Douchi T, Nagata Y, Inoue M, Hasegawa M, Yonemitsu Y, Yoshimura A (2004) SOCS1 [corrected] inhibits HPV-E7-mediated transformation by inducing degradation of E7 protein. Oncogene 23(17):3107–3115
Kamizono S, Hanada T, Yasukawa H, Minoguchi S, Kato R, Minoguchi M, Hattori K, Hatakeyama S, Yada M, Morita S, Kitamura T, Kato H, Nakayama K, Yoshimura A (2001) The SOCS box of SOCS-1 accelerates ubiquitin-dependent proteolysis of TEL-JAK2. J Biol Chem 276(16):12530–12538
Kamura T, Burian D, Yan Q, Schmidt SL, Lane WS, Querido E, Branton PE, Shilatifard A, Conaway RC, Conaway JW (2001) Muf1, a novel Elongin BC-interacting leucine-rich repeat protein that can assemble with Cul5 and Rbx1 to reconstitute a ubiquitin ligase. J Biol Chem 276(32):29748–29753
Kamura T, Maenaka K, Kotoshiba S, Matsumoto M, Kohda D, Conaway RC, Conaway JW, Nakayama KI (2004) VHL-box and SOCS-box domains determine binding specificity for Cul2-Rbx1 and Cul5-Rbx2 modules of ubiquitin ligases. Genes Dev 18(24):3055–3065
Kazi JU, Kabir NN, Flores-Morales A, Ronnstrand L (2014) SOCS proteins in regulation of receptor tyrosine kinase signaling. Cell Mol Life Sci 71(17):3297–3310
Kelleher DJ, Kreibich G, Gilmore R (1992) Oligosaccharyltransferase activity is associated with a protein complex composed of ribophorins I and II and a 48 kd protein. Cell 69(1):55–65
Keller KE, Wirtz MK (2017) Working your SOCS off: the role of ASB10 and protein degradation pathways in glaucoma. Exp Eye Res 158:154–160
Kelsall IR, Duda DM, Olszewski JL, Hofmann K, Knebel A, Langevin F, Wood N, Wightman M, Schulman BA, Alpi AF (2013) TRIAD1 and HHARI bind to and are activated by distinct neddylated Cullin-RING ligase complexes. EMBO J 32(21):2848–2860
Kile BT, Metcalf D, Mifsud S, DiRago L, Nicola NA, Hilton DJ, Alexander WS (2001) Functional analysis of Asb-1 using genetic modification in mice. Mol Cell Biol 21(18):6189–6197
Kile BT, Schulman BA, Alexander WS, Nicola NA, Martin HM, Hilton DJ (2002) The SOCS box: a tale of destruction and degradation. Trends Biochem Sci 27(5):235–241
Kim DW, Lee SH, Jeong MS, Sohn EJ, Kim MJ, Jeong HJ, An JJ, Jang SH, Won MH, Hwang IK, Cho SW, Kang TC, Lee KS, Park J, Yoo KY, Eum WS, Choi SY (2010) Transduced Tat-SAG fusion protein protects against oxidative stress and brain ischemic insult. Free Radic Biol Med 48(7):969–977
Kim JJ, Lee SB, Jang J, Yi SY, Kim SH, Han SA, Lee JM, Tong SY, Vincelette ND, Gao B, Yin P, Evans D, Choi DW, Qin B, Liu T, Zhang H, Deng M, Jen J, Zhang J, Wang L, Lou Z (2015) WSB1 promotes tumor metastasis by inducing pVHL degradation. Genes Dev 29(21):2244–2257
Kim JJ, Lee SB, Yi SY, Han SA, Kim SH, Lee JM, Tong SY, Yin P, Gao B, Zhang J, Lou Z (2017) WSB1 overcomes oncogene-induced senescence by targeting ATM for degradation. Cell Res 27(2):274–293
Kohroki J, Nishiyama T, Nakamura T, Masuho Y (2005) ASB proteins interact with Cullin5 and Rbx2 to form E3 ubiquitin ligase complexes. FEBS Lett 579(30):6796–6802
Kraus RJ, Yu X, Cordes BA, Sathiamoorthi S, Iempridee T, Nawandar DM, Ma S, Romero-Masters JC, McChesney KG, Lin Z, Makielski KR, Lee DL, Lambert PF, Johannsen EC, Kenney SC, Mertz JE (2017) Hypoxia-inducible factor-1alpha plays roles in Epstein-Barr virus’s natural life cycle and tumorigenesis by inducing lytic infection through direct binding to the immediate-early BZLF1 gene promoter. PLoS Pathog 13(6):e1006404
Kuang P, Tan M, Zhou W, Zhang Q, Sun Y (2016) SAG/RBX2 E3 ligase complexes with UBCH10 and UBE2S E2s to ubiquitylate beta-TrCP1 via K11-linkage for degradation. Sci Rep 6:37441
Kuang Z, Lewis RS, Curtis JM, Zhan Y, Saunders BM, Babon JJ, Kolesnik TB, Low A, Masters SL, Willson TA, Kedzierski L, Yao S, Handman E, Norton RS, Nicholson SE (2010) The SPRY domain-containing SOCS box protein SPSB2 targets iNOS for proteasomal degradation. J Cell Biol 190(1):129–141
Kwon S, Kim D, Rhee JW, Park JA, Kim DW, Kim DS, Lee Y, Kwon HJ (2010) ASB9 interacts with ubiquitous mitochondrial creatine kinase and inhibits mitochondrial function. BMC Biol 8:23
Lamsoul I, Uttenweiler-Joseph S, Moog-Lutz C, Lutz PG (2016) Cullin 5-RING E3 ubiquitin ligases, new therapeutic targets? Biochimie 122:339–347
Le-Trilling VT, Megger DA, Katschinski B, Landsberg CD, Ruckborn MU, Tao S, Krawczyk A, Bayer W, Drexler I, Tenbusch M, Sitek B, Trilling M (2016) Broad and potent antiviral activity of the NAE inhibitor MLN4924. Sci Rep 6:19977
Lewis RS, Kolesnik TB, Kuang Z, D’Cruz AA, Blewitt ME, Masters SL, Low A, Willson T, Norton RS, Nicholson SE (2011) TLR regulation of SPSB1 controls inducible nitric oxide synthase induction. J Immunol 187(7):3798–3805
Li H, Tan M, Jia L, Wei D, Zhao Y, Chen G, Xu J, Zhao L, Thomas D, Beer DG, Sun Y (2014) Inactivation of SAG/RBX2 E3 ubiquitin ligase suppresses KrasG12D-driven lung tumorigenesis. J Clin Invest 124(2):835–846
Li JY, Chai B, Zhang W, Wu X, Zhang C, Fritze D, Xia Z, Patterson C, Mulholland MW (2011) Ankyrin repeat and SOCS box containing protein 4 (Asb-4) colocalizes with insulin receptor substrate 4 (IRS4) in the hypothalamic neurons and mediates IRS4 degradation. BMC Neurosci 12:95
Lim SY, Tan AH, Ahmad-Annuar A, Klein C, Tan LCS, Rosales RL, Bhidayasiri R, Wu YR, Shang HF, Evans AH, Pal PK, Hattori N, Tan CT, Jeon B, Tan EK, Lang AE (2019) Parkinson’s disease in the Western Pacific region. Lancet Neurol
Linossi EM, Nicholson SE (2015) Kinase inhibition, competitive binding and proteasomal degradation: resolving the molecular function of the suppressor of cytokine signaling (SOCS) proteins. Immunol Rev 266(1):123–133
Liu B, Sarkis PT, Luo K, Yu Y, Yu XF (2005) Regulation of Apobec3F and human immunodeficiency virus type 1 Vif by Vif-Cul5-ElonB/C E3 ubiquitin ligase. J Virol 79(15):9579–9587
Liu E, Cote JF, Vuori K (2003) Negative regulation of FAK signaling by SOCS proteins. EMBO J 22(19):5036–5046
Liu S, Nheu T, Luwor R, Nicholson SE, Zhu HJ (2015) SPSB1, a novel negative regulator of the transforming growth factor-beta signaling pathway targeting the type II receptor. J Biol Chem 290(29):17894–17908
Liu Y, Zhou H, Zhu R, Ding F, Li Y, Cao X, Liu Z (2018) SPSB3 targets SNAIL for degradation in GSK-3beta phosphorylation-dependent manner and regulates metastasis. Oncogene 37(6):768–776
Lowenstein CJ, Padalko E (2004) iNOS (NOS2) at a glance. J Cell Sci 117(Pt 14):2865–2867
Lyapina S, Cope G, Shevchenko A, Serino G, Tsuge T, Zhou C, Wolf DA, Wei N, Shevchenko A, Deshaies RJ (2001) Promotion of NEDD-CUL1 conjugate cleavage by COP9 signalosome. Science 292(5520):1382–1385
Mahrour N, Redwine WB, Florens L, Swanson SK, Martin-Brown S, Bradford WD, Staehling-Hampton K, Washburn MP, Conaway RC, Conaway JW (2008) Characterization of Cullin-box sequences that direct recruitment of Cul2-Rbx1 and Cul5-Rbx2 modules to Elongin BC-based ubiquitin ligases. J Biol Chem 283(12):8005–8013
Mansell A, Smith R, Doyle SL, Gray P, Fenner JE, Crack PJ, Nicholson SE, Hilton DJ, O’Neill LA, Hertzog PJ (2006) Suppressor of cytokine signaling 1 negatively regulates Toll-like receptor signaling by mediating Mal degradation. Nat Immunol 7(2):148–155
Marteijn JA, van Emst L, Erpelinck-Verschueren CA, Nikoloski G, Menke A, de Witte T, Lowenberg B, Jansen JH, van der Reijden BA (2005) The E3 ubiquitin-protein ligase Triad1 inhibits clonogenic growth of primary myeloid progenitor cells. Blood 106(13):4114–4123
Matsumoto K, Nishiya T, Maekawa S, Horinouchi T, Ogasawara K, Uehara T, Miwa S (2011) The ECS(SPSB) E3 ubiquitin ligase is the master regulator of the lifetime of inducible nitric-oxide synthase. Biochem Biophys Res Commun 409(1):46–51
Monda JK, Scott DC, Miller DJ, Lydeard J, King D, Harper JW, Bennett EJ, Schulman BA (2013) Structural conservation of distinctive N-terminal acetylation-dependent interactions across a family of mammalian NEDD8 ligation enzymes. Structure 21(1):42–53
Nayak R, Pintel DJ (2007) Positive and negative effects of adenovirus type 5 helper functions on adeno-associated virus type 5 (AAV5) protein accumulation govern AAV5 virus production. J Virol 81(5):2205–2212
Nayak R, Farris KD, Pintel DJ (2008) E4Orf6-E1B-55k-dependent degradation of de novo-generated adeno-associated virus type 5 Rep52 and capsid proteins employs a cullin 5-containing E3 ligase complex. J Virol 82(7):3803–3808
Nishiya T, Matsumoto K, Maekawa S, Kajita E, Horinouchi T, Fujimuro M, Ogasawara K, Uehara T, Miwa S (2011) Regulation of inducible nitric-oxide synthase by the SPRY domain- and SOCS box-containing proteins. J Biol Chem 286(11):9009–9019
Nucifora FC Jr, Nucifora LG, Ng CH, Arbez N, Guo Y, Roby E, Shani V, Engelender S, Wei D, Wang XF, Li T, Moore DJ, Pletnikova O, Troncoso JC, Sawa A, Dawson TM, Smith W, Lim KL, Ross CA (2016) Ubiquitination via K27 and K29 chains signals aggregation and neuronal protection of LRRK2 by WSB1. Nat Commun 7:11792
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T, Tokino T, Taniguchi T, Tanaka N (2000) Noxa, a BH3-only member of the Bcl-2 family and candidate mediator of p53-induced apoptosis. Science 288(5468):1053–1058
Ohta T, Michel JJ, Schottelius AJ, Xiong Y (1999) ROC1, a homolog of APC11, represents a family of cullin partners with an associated ubiquitin ligase activity. Mol Cell 3(4):535–541
Okumura F, Matsuzaki M, Nakatsukasa K, Kamura T (2012) The role of Elongin BC-containing ubiquitin ligases. Front Oncol 2:10
Orr SJ, Morgan NM, Elliott J, Burrows JF, Scott CJ, McVicar DW, Johnston JA (2007) CD33 responses are blocked by SOCS3 through accelerated proteasomal-mediated turnover. Blood 109(3):1061–1068
Parrillas V, Martinez-Munoz L, Holgado BL, Kumar A, Cascio G, Lucas P, Rodriguez-Frade JM, Malumbres M, Carrera AC, van Wely KH, Mellado M (2013) Suppressor of cytokine signaling 1 blocks mitosis in human melanoma cells. Cell Mol Life Sci 70(3):545–558
Perfetto L, Gherardini PF, Davey NE, Diella F, Helmer-Citterich M, Cesareni G (2013) Exploring the diversity of SPRY/B30.2-mediated interactions. Trends Biochem Sci 38(1):38–46
Peterson TR, Laplante M, Thoreen CC, Sancak Y, Kang SA, Kuehl WM, Gray NS, Sabatini DM (2009) DEPTOR is an mTOR inhibitor frequently overexpressed in multiple myeloma cells and required for their survival. Cell 137(5):873–886
Petroski MD, Deshaies RJ (2005) Function and regulation of cullin-RING ubiquitin ligases. Nat Rev Mol Cell Biol 6(1):9–20
Puca R, Nardinocchi L, Sacchi A, Rechavi G, Givol D, D’Orazi G (2009) HIPK2 modulates p53 activity towards pro-apoptotic transcription. Mol Cancer 8:85
Querido E, Blanchette P, Yan Q, Kamura T, Morrison M, Boivin D, Kaelin WG, Conaway RC, Conaway JW, Branton PE (2001) Degradation of p53 by adenovirus E4orf6 and E1B55K proteins occurs via a novel mechanism involving a Cullin-containing complex. Genes Dev 15(23):3104–3117
Rabut G, Peter M (2008) Function and regulation of protein neddylation. ‘Protein modifications: beyond the usual suspects’ review series. EMBO Rep 9(10):969–976
Rodrigues L, Filipe J, Seldon MP, Fonseca L, Anrather J, Soares MP, Simas JP (2009) Termination of NF-kappaB activity through a gammaherpesvirus protein that assembles an EC5S ubiquitin-ligase. EMBO J 28(9):1283–1295
Rui L, Yuan M, Frantz D, Shoelson S, White MF (2002) SOCS-1 and SOCS-3 block insulin signaling by ubiquitin-mediated degradation of IRS1 and IRS2. J Biol Chem 277(44):42394–42398
Ryo A, Suizu F, Yoshida Y, Perrem K, Liou YC, Wulf G, Rottapel R, Yamaoka S, Lu KP (2003) Regulation of NF-kappaB signaling by Pin1-dependent prolyl isomerization and ubiquitin-mediated proteolysis of p65/RelA. Mol Cell 12(6):1413–1426
Samant RS, Clarke PA, Workman P (2014) E3 ubiquitin ligase Cullin-5 modulates multiple molecular and cellular responses to heat shock protein 90 inhibition in human cancer cells. Proc Natl Acad Sci U S A 111(18):6834–6839
Sarikas A, Hartmann T, Pan ZQ (2011) The cullin protein family. Genome Biol 12(4):220
Sato Y, Shirata N, Kudoh A, Iwahori S, Nakayama S, Murata T, Isomura H, Nishiyama Y, Tsurumi T (2009) Expression of Epstein-Barr virus BZLF1 immediate-early protein induces p53 degradation independent of MDM2, leading to repression of p53-mediated transcription. Virology 388(1):204–211
Sheehy AM, Gaddis NC, Malim MH (2003) The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nat Med 9(11):1404–1407
Simas JP, Efstathiou S (1998) Murine gammaherpesvirus 68: a model for the study of gammaherpesvirus pathogenesis. Trends Microbiol 6(7):276–282
Simo S, Cooper JA (2013) Rbx2 regulates neuronal migration through different cullin 5-RING ligase adaptors. Dev Cell 27(4):399–411
Soucy TA, Smith PG, Milhollen MA, Berger AJ, Gavin JM, Adhikari S, Brownell JE, Burke KE, Cardin DP, Critchley S, Cullis CA, Doucette A, Garnsey JJ, Gaulin JL, Gershman RE, Lublinsky AR, McDonald A, Mizutani H, Narayanan U, Olhava EJ, Peluso S, Rezaei M, Sintchak MD, Talreja T, Thomas MP, Traore T, Vyskocil S, Weatherhead GS, Yu J, Zhang J, Dick LR, Claiborne CF, Rolfe M, Bolen JB, Langston SP (2009) An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458(7239):732–736
Stanley DJ, Bartholomeeusen K, Crosby DC, Kim DY, Kwon E, Yen L, Cartozo NC, Li M, Jager S, Mason-Herr J, Hayashi F, Yokoyama S, Krogan NJ, Harris RS, Peterlin BM, Gross JD (2012) Inhibition of a NEDD8 cascade restores restriction of HIV by APOBEC3G. PLoS Pathog 8(12):e1003085
Steegenga WT, Riteco N, Jochemsen AG, Fallaux FJ, Bos JL (1998) The large E1B protein together with the E4orf6 protein target p53 for active degradation in adenovirus infected cells. Oncogene 16(3):349–357
Stracker TH, Carson CT, Weitzman MD (2002) Adenovirus oncoproteins inactivate the Mre11-Rad50-NBS1 DNA repair complex. Nature 418(6895):348–352
Sun Y, Li H (2013) Functional characterization of SAG/RBX2/ROC2/RNF7, an antioxidant protein and an E3 ubiquitin ligase. Protein Cell 4(2):103–116
Suspene R, Sommer P, Henry M, Ferris S, Guetard D, Pochet S, Chester A, Navaratnam N, Wain-Hobson S, Vartanian JP (2004) APOBEC3G is a single-stranded DNA cytidine deaminase and functions independently of HIV reverse transcriptase. Nucleic Acids Res 32(8):2421–2429
Swaroop M, Wang Y, Miller P, Duan H, Jatkoe T, Madore S, Sun Y (2000) Yeast homolog of human SAG/ROC2/Rbx2/Hrt2 is essential for cell growth, but not for germination: chip profiling implicates its role in cell cycle regulation. Oncogene 19:2855–2866
Swaroop M, Gosink M, Sun Y (2001) SAG/ROC2/Rbx2/Hrt2, a component of SCF E3 ubiquitin ligase: genomic structure, a splicing variant, and two family pseudogenes. DNA Cell Biol 20(7):425–434
Taipale M, Krykbaeva I, Koeva M, Kayatekin C, Westover KD, Karras GI, Lindquist S (2012) Quantitative analysis of HSP90-client interactions reveals principles of substrate recognition. Cell 150(5):987–1001
Tan M, Gu Q, He H, Pamarthy D, Semenza GL, Sun Y (2008) SAG/ROC2/RBX2 is a HIF-1 target gene that promotes HIF-1 alpha ubiquitination and degradation. Oncogene 27(10):1404–1411
Tan M, Davis SW, Saunders TL, Zhu Y, Sun Y (2009) RBX1/ROC1 disruption results in early embryonic lethality due to proliferation failure, partially rescued by simultaneous loss of p27. Proc Natl Acad Sci U S A 106(15):6203–6208
Tan M, Zhu Y, Kovacev J, Zhao Y, Pan ZQ, Spitz DR, Sun Y (2010) Disruption of Sag/Rbx2/Roc2 induces radiosensitization by increasing ROS levels and blocking NF-kappaB activation in mouse embryonic stem cells. Free Radic Biol Med 49(6):976–983
Tan M, Zhao Y, Kim SJ, Liu M, Jia L, Saunders TL, Zhu Y, Sun Y (2011) SAG/RBX2/ROC2 E3 ubiquitin ligase is essential for vascular and neural development by targeting NF1 for degradation. Dev Cell 21(6):1062–1076
Tan M, Xu J, Siddiqui J, Feng F, Sun Y (2016) Depletion of SAG/RBX2 E3 ubiquitin ligase suppresses prostate tumorigenesis via inactivation of the PI3K/AKT/mTOR axis. Mol Cancer 15(1):81
Tapia-Laliena MA, Korzeniewski N, Pena-Llopis S, Scholl C, Frohling S, Hohenfellner M, Duensing A, Duensing S (2019) Cullin 5 is a novel candidate tumor suppressor in renal cell carcinoma involved in the maintenance of genome stability. Oncogene 8(1):4
Teckchandani A, Laszlo GS, Simo S, Shah K, Pilling C, Strait AA, Cooper JA (2014) Cullin 5 destabilizes Cas to inhibit Src-dependent cell transformation. J Cell Sci 127(Pt 3):509–520
Tissir F, Goffinet AM (2003) Reelin and brain development. Nat Rev Neurosci 4(6):496–505
Townley-Tilson WH, Wu Y, Ferguson JE 3rd, Patterson C (2014) The ubiquitin ligase ASB4 promotes trophoblast differentiation through the degradation of ID2. PLoS One 9(2):e89451
Uematsu K, Okumura F, Tonogai S, Joo-Okumura A, Alemayehu DH, Nishikimi A, Fukui Y, Nakatsukasa K, Kamura T (2016) ASB7 regulates spindle dynamics and genome integrity by targeting DDA3 for proteasomal degradation. J Cell Biol 215(1):95–106
Ungureanu D, Saharinen P, Junttila I, Hilton DJ, Silvennoinen O (2002) Regulation of Jak2 through the ubiquitin-proteasome pathway involves phosphorylation of Jak2 on Y1007 and interaction with SOCS-1. Mol Cell Biol 22(10):3316–3326
Wang J, Muntean AG, Hess JL (2012) ECSASB2 mediates MLL degradation during hematopoietic differentiation. Blood 119(5):1151–1161
Wang X, Wang X, Wang W, Zhang J, Wang J, Wang C, Lv M, Zuo T, Liu D, Zhang H, Wu J, Yu B, Kong W, Wu H, Yu X (2015) Both Rbx1 and Rbx2 exhibit a functional role in the HIV-1 Vif-Cullin5 E3 ligase complex in vitro. Biochem Biophys Res Commun 461(4):624–629
Wilcox A, Katsanakis KD, Bheda F, Pillay TS (2004) Asb6, an adipocyte-specific ankyrin and SOCS box protein, interacts with APS to enable recruitment of elongins B and C to the insulin receptor signaling complex. J Biol Chem 279(37):38881–38888
Xie CM, Wei D, Zhao L, Marchetto S, Mei L, Borg JP, Sun Y (2015) Erbin is a novel substrate of the Sag-betaTrCP E3 ligase that regulates KrasG12D-induced skin tumorigenesis. J Cell Biol 209(5):721–737
Xu XM, Wang XB, Chen MM, Liu T, Li YX, Jia WH, Liu M, Li X, Tang H (2012) MicroRNA-19a and -19b regulate cervical carcinoma cell proliferation and invasion by targeting CUL5. Cancer Lett 322(2):148–158
Yang GY, Pang L, Ge HL, Tan M, Ye W, Liu XH, Huang FP, Wu DC, Che XM, Song Y, Wen R, Sun Y (2001) Attenuation of ischemia-induced mouse brain injury by SAG, a redox- inducible antioxidant protein. J Cereb Blood Flow Metab 21(6):722–733
Yasukawa T, Kamura T, Kitajima S, Conaway RC, Conaway JW, Aso T (2008) Mammalian Elongin A complex mediates DNA-damage-induced ubiquitylation and degradation of Rpb1. EMBO J 27(24):3256–3266
Yu X, Yu Y, Liu B, Luo K, Kong W, Mao P, Yu XF (2003) Induction of APOBEC3G ubiquitination and degradation by an HIV-1 Vif-Cul5-SCF complex. Science 302(5647):1056–1060
Zhao G, Gong L, Su D, Jin Y, Guo C, Yue M, Yao S, Qin Z, Ye Y, Tang Y, Wu Q, Zhang J, Cui B, Ding Q, Huang H, Hu L, Chen Y, Zhang P, Hu G, Chen L, Wong KK, Gao D, Ji H (2019) Cullin5 deficiency promotes small-cell lung cancer metastasis by stabilizing integrin beta1. J Clin Invest 129(3):972–987
Zhao Y, Xiong X, Sun Y (2011) DEPTOR, an mTOR inhibitor, is a physiological substrate of SCF(betaTrCP) E3 ubiquitin ligase and regulates survival and autophagy. Mol Cell 44(2):304–316
Zhou L, Zhang W, Sun Y, Jia L (2018a) Protein neddylation and its alterations in human cancers for targeted therapy. Cell Signal 44:92–102
Zhou W, Xu J, Li H, Xu M, Chen ZJ, Wei W, Pan Z, Sun Y (2017) Neddylation E2 UBE2F promotes the survival of lung cancer cells by activating CRL5 to degrade NOXA via the K11 linkage. Clin Cancer Res 23(4):1104–1116
Zhou W, Xu J, Tan M, Li H, Li H, Wei W, Sun Y (2018b) UBE2M is a stress-inducible dual E2 for neddylation and ubiquitylation that promotes targeted degradation of UBE2F. Mol Cell 70(6):1008–1024.e6
Zhu Y, Li L, Hou D, Ouyang Y, Guo X, Wang Y, Li J, Gong K (2019) MicroRNA-19a regulates the proliferation, migration and invasion of human gastric cancer cells by targeting CUL5. Arch Biochem Biophys 662:93–100
Zhu Z, Wang L, Hao R, Zhao B, Sun L, Ye RD (2016) Cutting edge: a Cullin-5-TRAF6 interaction promotes TRAF6 polyubiquitination and lipopolysaccharide signaling. J Immunol 197(1):21–26
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Zhang, S., Sun, Y. (2020). Cullin RING Ligase 5 (CRL-5): Neddylation Activation and Biological Functions. In: Sun, Y., Wei, W., Jin, J. (eds) Cullin-RING Ligases and Protein Neddylation. Advances in Experimental Medicine and Biology, vol 1217. Springer, Singapore. https://doi.org/10.1007/978-981-15-1025-0_16
Download citation
DOI: https://doi.org/10.1007/978-981-15-1025-0_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1024-3
Online ISBN: 978-981-15-1025-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)