Abstract
Flavivirus NS5 RNA-dependent RNA polymerase (RdRp) is an important drug target. Whilst a number of allosteric inhibitors have been described for Hepatitis C virus RdRp, few have been described for DENV RdRp. In addition, compound screening campaigns have not yielded suitable leads for this enzyme. Using fragment-based screening via X-ray crystallography, we identified a biphenyl acetic acid fragment that binds to a novel pocket of the dengue virus (DENV) RdRp, in the thumb/palm interface, close to its active site (termed “N pocket”). Structure-guided optimization yielded nanomolar inhibitors of the RdRp de novo initiation activity, with low micromolar EC50 in DENV cell-based assays. Compound-resistant DENV replicons exhibited amino acid mutations that mapped to the N pocket. This is the first report of a class of pan-serotype and cell-active DENV RdRp inhibitors and provides a significant opportunity for rational design of novel therapeutics against this proven antiviral target.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Flaviviruses
- Dengue virus
- RNA dependent RNA polymerase
- Anti-viral
- Drug discovery
- Rational design
- X-ray crystallography
- Mechanism of inhibition
- Resistance phenotype
14.1 Introduction
Flavivirus non-structural protein 5 (NS5) is the largest protein encoded by the viral genome. It is also the most highly conserved, with approximately 67-82% amino-acid sequence identity between the four serotypes of dengue virus (DENV-1 to -4). The N-terminal region comprises an RNA methyltransferase (MTase) domain [9] that has both N7 and 2′-O methyl-transferase activities [4, 6, 12, 22]. Together, they are responsible for type 1 cap structure (m7GpppAm) formation at the 5′ end of the viral genome. The C-terminal two-thirds of NS5 harbors an RNA-dependent RNA polymerase domain (RdRp; [8]) which performs de novo RNA synthesis from the 3′ end of the viral template [3]. These two domains are linked via a short amino acid sequence of low conservation and may play a role in regulating RdRp activity [13, 21].
Crystal structures of the DENV and other flavivirus MTase and RdRp domains and the full length NS5 protein, have been solved. The overall architecture of the polymerase resembles a right hand with fingers, palm, and thumb sub-domains, with a fully-encircled active site. In addition, they are well-characterized, structurally and functionally. For more detailed information on these aspects, the readers are encouraged to refer to excellent reviews [1, 3, 5, 14, 21] as well as Chaps. 8, 9, and 19 in this monograph.
Recently, a class of pyridobenzothiazole compounds that inhibit DENV RdRp and DENV infection at low micromolar potencies was reported [23]. This class of compounds binds in a similar site as NITD107, in the DENV RdRp a RNA tunnel [19]. Herein, we describe a class of acyl sulfonamide-thiophene compounds that has nanomolar inhibitory potency against the de novo initiation activities of DENV1-4 RdRp [18] and also exhibit EC50 values of 1–2 micromolar in DENV-infected cells [15, 20, 26].
14.2 Identification of N-Pocket Binding Fragment by DENV RdRp Fragment-Based Screening
14.2.1 Fragment-Based Screening Via DENV RdRp Crystallography
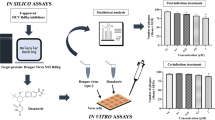
A total of 1408 compounds obtained from the Novartis in-house fragment library were screened with DENV3 RdRp, in pools of 8, totaling 176 separate pools. DENV3 RdRp crystals were first crystallized in conditions as previously described [19] and soaked for 4–6 h in drops containing compound mixtures in crystallization buffer and 10% DMSO, giving a final concentration per compound of 625 μM. The crystals were then frozen in liquid nitrogen after cryo-protection in the same crystallization solution supplemented with 10% glycerol and 10% compound/DMSO. The X-ray diffraction data were collected on beam-line X10SA (PXII) at the Swiss Light Source, integrated using MOSFLM [11] and scaled using SCALA, part of the CCP4 suite [24]. The structures were refined using REFMAC5 [17] starting from the deposited DENV-3 RdRp structure (PDB code 4HHJ) [19]. Useful X-ray diffraction data was collected for 145 out of 176 pools (with resolution better than 2.5 Å). Only one dataset, which was solved to 2.0 Å resolution, showed difference density that could be interpreted as a single contiguous compound. Models for the eight possible fragments from the pool were built into the density and only compound JF-31-MG46 fitted the data satisfactorily. To confirm that the single compound bound in the pocket, the DENV-3 RdRp was co-crystallized with the pure compound and the structure was solved to 2.05 Å (Fig. 14.1a; PDB 5F3T). The co-crystal structure showed that the single compound bound at the same site and orientation with full occupancy. Specifically, it bound in a novel pocket between the thumb and palm sub-domains of the RdRp and the priming loop. This site is designed the “N” pocket and is similar to the Site III (Palm I) in HCV ([3]; See also Summary and Conclusions).
Crystal structure of N pocket inhibitors and the DENV-3 RdRp. Binding of JF-31-MG46 (orange, a), compounds 10 (grey, b) and 15 (cyan, b) and compound 29 (yellow, c) in the N pocket of DENV-3 RdRp domain are shown as sticks. Amino acid residues lining the N pocket are colored in purple (a), cyan (c), or green (palm and thumb subdomains, b) and yellow (priming loop, b). Individual residues are labelled according to their numbering in the DENV3 polymerase and H-bond interactions are indicated with dashed lines
Additional information on this Section can be found in [20].
14.2.2 Compound JF-31-MG46 and DENV RdRp N Pocket
Compound JF-31-MG46, 3 is a biphenyl acetic acid fragment, and forms interactions with several amino acid residues in the DENV RdRp N pocket which are highly conserved across the four DENV serotypes, and in other flaviviruses such as WNV, YFV, and JEV (Fig.14.1a; PDB 5F3T). Its outer phenyl is sandwiched between Ser796 on one side and Tyr766 and His711 on the other. The carboxyl group on this ring, interacts with the side chain OH of Thr794, and also forms a water-mediated interaction with Arg737. The other substituent on the ring, the methoxy group, is surrounded by residues Cys709, Leu514, His798, and Leu511. The inner phenyl occupies the back of the pocket lined with several hydrophobic residues, including Trp803, Met761, Met765, Ala799, and Leu511, with Ser710 located at the mouth of the pocket. Surface-plasmon resonance (SPR) analyses indicated that JF-31-MG46 binds to DENV-3 RdRp with Kd of 209 μM and with threefold lower binding affinity for DENV-4 RdRp (Kd = 610 μM). No binding was detected by isothermal calorimetry, presumably due to the lack of sufficient sensitivity at the maximum compound solubility. Compound JF-31-MG46 inhibited the de novo initiation activities of DENV-1-4 with IC50 values ranging from 734 to 796 μM and is consistent with the affinities measured by SPR. The calculated ligand efficiency [10] for compound JF-31-MG46 is 0.24.
Additional information on this Section can be found in Lim et al. [15] and [20].
14.3 Structure-Guided Inhibitor Optimization of N-Pocket Inhibitors
14.3.1 Improving Inhibitory Property of N-pocket Inhibitors
Based on the X-ray crystallography finding that a related analog, 4 (IC50 = 769 μM, SPR-Kd > 200 μM, LE 0.26; PDB 5HNO) also bound in the same pocket but in the opposite orientation of the carboxylic acid moiety compared to compound JF-31-MG, the two fragments were merged, to generate the bis-acid, 5. This resulted in stronger binding in the N-pocket and better inhibition (IC50 = 177 μM, ITC-K d = 154 μM; LE = 0.26; PDB 5HMW). The initial optimization efforts focused on exploring the SAR of the inner benzene ring of 5. Substitution with 2′-thiophene ring, 10, provided a tenfold increase in potency (IC50 = 15 μM, ITC-K d = 28 μM). This higher affinity is likely due to non-covalent interaction of the sulfur of the thiophene with the oxygen of the OH side-chain of Ser796 (PDB 5HMX). Computational studies detected a water molecule in the back of the DENV N pocket, which forms H-bond interactions with residues Gln802 and His800. Design strategies to improve inhibitor potency by contacting this water molecule were not fruitful. An alternate approach was conducted to displace the water molecule. A propargyl alcohol extension from the 2-thiophene ring provided 15 (IC50 = 1.7 μM, ITC-K d = 1.4 μM; LE = 0.35), with a 100-fold increase in potency as compared to 5. Co-crystallization of 15 with the NS5 RdRp domain confirmed that the propargyl alcohol filled the narrow cavity and displaced the water molecule to form the H-bond interactions with the side chain of Gln802 and the backbone of His800 as predicted (Fig. 14.1b; PDB 5HMY). Compound 15 displayed inhibitory activity against all four serotypes with IC50 values of 0.3 to 2.2 μM.
Additional information on this Section can be found in Lim et al. [15] and [26].
14.3.2 Improving Cellular Permeability of N-Pocket Inhibitors
Poor membrane permeability rendered compound 15 to be inactive in DENV-infected cells, mostly due to the presence of the negatively charged bis-carboxylic acids. Efforts to identify carboxylic bioisoteres lead to the acylsulfonamide moiety which dramatically improved their potencies and binding affinities in the biochemical and biophysical analyses (compounds 23–29i; Fig. 14.1c; Table 14.1). Crystal structure of DENV-3 RdRp with 23 revealed that the acylsulfonamide formed H-bond interactions with the side chains of Thr794 and Arg729 and the backbone of Trp795 (PDB 5HMZ). The methyl group of the acylsulfonamide moiety was exposed to the solvent space, providing the opportunity to modify the overall physicochemical property of inhibitor without interfering with its affinity to the enzyme. Changing the methyl to benzene sulfonamide 25 (which increased its lipophilicity, whilst retaining similar acidity (pKa) and permeability as 23), provided weak cellular activity (EC50 18-41 μM). Replacing the 6-methyl group of the methoxy benzene ring of 25 with another electron-donating methoxy group, improved cell permeability of 26 (by reducing ionization of the acylsulfonamide) and led to modest improvement in cellular potency (EC50 = 7.2–36 μM). The most potent anti-dengue activity was obtained with the 3-methoxyphenyl sulfonamide derivative, 2 7, which displayed EC50 values of low micromolar against all four serotypes (EC50 1.8–2.3 μM). The exact cause of the significant improvement in the cellular potency of 27 is not known, as it has poorer permeability compared to 26. A related analog, 26i, which contains a single methyoxy group on the central benzene ring, compared to two methoxy substituents on 27, exhibited significantly poorer cellular activity than the latter (EC50 = 15-27 μM). The additional methoxy group in 27 may allow the formation of an intra-hydrogen bond with the N-atom of the sulfonamide linker to facilitate better cell permeability. The same observation applies to the 8-quinolyl sulfonamide derivatives 29 and 29i (EC50 = 4–14 μM and 2–6 μM respectively). Inhibitors 26–29i were inactive against HCV replicon and human rhinoviruses (EC50 values >25 μM and >50 μM, respectively). No cytotoxicity was observed in five different cell lines tested (EC50 > 50 μM).
Additional information on this Section can be found in Lim et al. [15] and [26].
14.4 Profiling Mechanism of Action of N-Pocket Inhibitors
14.4.1 Mechanism Studies of Inhibitors in Biochemical Assays
In the standard assay format for DENV dnI FAPA assay, N pocket compounds 15, 27 and 29 were first exposed to enzyme alone followed by reaction initiations with ssRNA template and NTPs. Order-of-reagent addition experiments in which compounds were exposed to pre-formed enzyme-ssRNA complexes, followed by reaction initiation with NTPs, did not result in significant changes in their IC50 values (Table 14.2). Compounds exposed to elongated enzyme-dsRNA complexes, in which the active site was occupied by the ssRNA template and newly synthesized short RNA products AGAA or AGAACC reduced their inhibitory potencies by 8–15-fold. These findings suggest that the N-pocket underwent conformational changes when the DENV polymerase entered into the elongation phase, which reduced the binding affinities of the compounds. Compounds also demonstrated 10–23-folds weaker inhibitory properties in the DENV elongation FAPA assay compared to the standard dnI assay. Overall, the data indicate that N pocket inhibitors primarily inhibit DENV polymerase de novo initiation activity. Kinetics studies using Lineweaver-Burk plots further revealed that both 15 and 29 have uncompetitive inhibition profiles with respect to the viral ssRNA template. On the other hand, they display mixed non/uncompetitive inhibition with respect to GTP.
Additional information on this Section can be found in Lim et al. [15].
14.4.2 Biological Relevance of N-pocket for DENV Replication
Alanine mutations of RdRp residues interacting with 27 (PDB 5K5M, 5I3P and 5JJS) or 29 (PDB 5I3Q and 5JJR) and as well as residues lining the N-pocket were generated in the recombinant NS5 protein and the DENV replicon. Alanine mutations mostly negatively impacted DENV polymerase de novo initiation (particularly for residues S710 and R737), and have less influence on elongation activity. In agreement with the biochemical data, N pocket mutant DENV replicon R737A (as well as R729A mutant replicon) was non-replicative. R729A RdRp exhibited about 30–40% of both dnI and elongation activities whilst dnI activity of R737A was completely abolished. Mutant replicons Y766A, and W803A were also non-replicative, despite showing 55-98% de novo initiation and elongation activities in vitro. Mutant replicons with T794A and S796A with comparable levels of in vitro polymerase activities as WT NS5 polymerase, were weakly replicative. Overall, the N pocket conformation observed in the inhibitor-bound crystal structures is likely to correspond to the structural state adopted by the DENV RdRp during de novo initiation and they play important functions for virus replication.
Additional information on this Section can be found in Lim et al. [15] and [20].
14.4.3 Resistant DENV Replicons Raised to N-Pocket Inhibitors
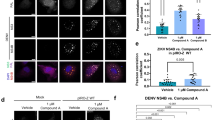
We propagated stable DENV-2-NGC EGFP replicon cells in 20 μM of 29 (≈1X EC90 value) for 5 weeks, and then increased the compound concentration to 25 μM. Two separate compound-resistant replicon clones harbored the same nucleotide change in NS5 (GAA → GAC), resulting in Glu802Asp mutation (note that residue 802 is Glu in DENV2-NGC and Gln in DENV3 RdRp). A third clone contained another nucleotide change in NS5 (CTG → GTG), resulting in Leu511Val mutation. A fourth clone contained a mixed profile in NS5, in the same position, with both the WT nucleotide (G) as well as mutation to C nucleotide present (GTG → G/CTG), giving rise to partial Leu511Val mutation. Similar efforts to raise resistant cells by exposure to high concentrations of 27 (14–20 μM; ≈2X EC90 value) were not successful. Stable DENV-2-NGC replicon cells were exposed to increasing concentrations of 27, starting from 1.5 μM (≈0.5X EC50), with media change every 2–3 days. After about 6 weeks, cells kept in 28 μM of 27 propagated robustly, at similar rates to WT DENV-2 replicon cells. RNA sequenced from the cells contained a partial Glu802Asp mutation profile (GAA → GAA/T). The crystal structures of DENV-3-RdRp bound to 29 (PDB 5I3Q, 5JJR) shows that the polar side chain of residue Gln802 (Glu802 in DENV2) hydrogens bond with the hydroxyl group of the propargyl alcohol of 29. E802D mutation results in the shortening of the amino acid side-chain by one methyl group and is likely to disrupt this H-bond formation. Residue Leu511 (in DENV-2 and -3) forms van der Waals interactions with the thiophene ring of 29. The loss of a methyl group from Val511 mutation, would weaken the interaction with the thiophene ring. Overall, these mutations lower the binding affinity of 29 in the N-pocket and verify that 27 and 29 inhibit DENV replication in cells by binding to the N-pocket in the DENV polymerase.
Additional information on this Section can be found in Lim et al. [15].
14.4.4 Impact of Resistance Mutations on Compound Inhibition
We generated recombinant NS5 mutant proteins with N-pocket amino acid changes in DENV-2 (L511V and E802D) and DENV-4 (L512V and Q803N) and tested them against 27 and 29 (Table 14.3). Both compounds were significantly less active against mutant enzymes than WT protein in the dnI FAPA assay. Compared WT protein, IC50 values of 29 declined by 4–12-folds in DENV-2 and DENV-4, single mutants, and by 52–133-folds in double mutant enzymes. Likewise for 27, inhibitory potency fell by 5–14-folds in DENV-2 single mutants and by 88-folds in the double mutant. A complete loss of inhibitory activity (IC50 > 20 μM) was observed in DENV-4 single and double mutants. Furthermore, these compounds were less effective in stabilizing the mutant enzymes compared to the corresponding WT proteins [15]. Similar single and double (L511V and E802D) amino acid changes were also introduced into the DENV-2 (strain NGC) replicon and its infectious full length virus genome to examine their effects on the potencies of N pocket inhibitors (Table 14.3). Compared to WT replicon and virus, EC50 value of 29 was reduced by 3–6-folds in single and double mutant DENV-2 replicons and by 5–6-folds in virus mutants. On the other hand, EC50 values of 27 were reduced only by 2–4-folds in mutant DENV-2 replicons and viruses. The observed weaker EC50 shifts for 27 are puzzling as its binding mode is similar to 29 and involves non-covalent interaction of the thiophene ring with L511 and H-bond formation between the propargyl alcohol and E802D. Additional 27-resistant DENV-2 EGFP-replicons shifted the EC50 values of 27 and 29 shifted by 17- and 10-folds, respectively, in these cells, compared to control cells raised in DMSO. Full replicon genome sequence analyses revealed secondary mutations present in NS5 methyl-transferase and NS4B in the 27-resistant replicon cells, suggesting that N pocket inhibitors also affected the replication complex formation.
Additional information on this Section can be found in Lim et al. [15].
14.5 Summary and Conclusions
From fragment-based screening by X-ray crystallography with the DENV-3 apo-RdRp protein [19, 25], we identified a novel allosteric pocket at the DENV-3 RdRp thumb and palm interface [15, 20, 26]. This pocket which we term the “N pocket” is located near the priming loop (aa782-809) and is lined by residues highly conserved across DENV-1-4, as well as in other flaviviruses. Alanine mutagenesis studies indicate that this pocket is important for NS5 polymerase de novo initiation activity and virus replication. N-pocket inhibitors generated by rational design potently inhibited DENV-1-4 polymerase de novo initiation activities and virus replication in various cell types. They have a mixed non−/uncompetitive inhibition profile and bind with strong affinity to recombinant apo-enzyme as well as FL NS5 from DENV replicon cell lysates [15].
Resistant DENV raised against N pocket inhibitors harbored amino acid mutations (L511 V and E802D; DENV-2 numbering) that mapped to the N-pocket and reduced compound potencies in DENV RdRp enzyme and DENV cell-based assays. Based on results from order-of-reagent addition experiments, N pocket inhibitors block the enzyme de novo initiation activity better than the elongation activity. Compound potencies are reduced when the enzyme is pre-occupied with newly synthesized duplex RNA, and not by single-stranded viral RNA. Presumably, retraction of the priming loop (aa782-809) from the active site during enzyme elongation alters the conformation of the N-pocket, leading to weaker binding affinities of the RdRp for the compounds.
DENV N-pocket compounds resemble HCV polymerase Site III (palm 1) non-nucleoside inhibitors [2]. The latter inhibitors bind at the interface of the HCV polymerase thumb and palm subdomains, with one side comprising the “primer grip” and the opposite side formed by the β-hairpin loop (equivalent to the priming loop in DENV RdRp). Inhibitor binding is promoted by interactions with both sides, in particular, with Y448 from the β-loop. This locks the HCV RdRp thumb subdomain in a conformation that prevents de novo initiation. Dasabuvir (ABT 333), a Site III inhibitor, has recently been approved for HCV therapy in combination with NS3/4A protease and NS5 inhibitors [7].
There is, however, no equivalent of the HCV RdRp primer grip wall for DENV N-pocket. In addition, unlike HCV RdRp where the C-terminal loop penetrates the active site and participates in enzyme activity, the C-terminal end of flavivirus RdRp is disordered in most reported crystal structures. We speculate that the absence of both regions prevent formation of additional contacts with inhibitors, and is the reason for the weaker binding affinities of N-pocket compounds, compared to HCV site III inhibitors. The availability of a co-crystal structure of DENV RdRp with RNA would hopefully, facilitate new design strategies to further enhance inhibitor binding affinity and block elongation activity.
High clearance was observed for acyl-sulfonamide propargyl alcohol compounds in vitro which rendered them unsuitable for mouse efficacy studies. To develop N-pocket inhibitors, more stable functionalities with better pharmaco-kinetic properties, that retain key hydrogen bond interactions, are required.
Finally, compounds 27 and 29 were inactive when tested on the WNV replicon cell-based assay. Previous comparisons revealed that the WNV RdRp priming loop is closer to the i-1 site, and prevents formation of a similar N-pocket [16]. Whilst DENV N-pocket residues are mostly conserved across the flavivirus family, residues 799-802, which accommodate the propargyl alcohol arm, are more divergent. This may explain the lack of activity of 27 and 29 on WNV. Interestingly, residues 799-802 are more similar amongst JEV, MVEV WNV, YFV and ZIKV compared to DENV-1-4. In this light, it may not be plausible to develop pan-active N-pocket inhibitors that work on all flaviviruses. Rather, designing N-pocket inhibitors that specifically target different subgroups of the flavivirus family may be a more attainable goal.
References
Bollati M, Alvarez K, Assenberg R, Baronti C, Canard B, Cook S, Coutard B, Decroly E, de Lamballerie X, Gould EA, Grard G, Grimes JM, Hilgenfeld R, Jansson AM, Malet H, Mancini EJ, Mastrangelo E, Mattevi A, Milani M, Moureau G, Neyts J, Owens RJ, Ren J, Selisko B, Speroni S, Steuber H, Stuart DI, Unge T, Bolognesi M (2010) Structure and functionality in flavivirus NS-proteins: perspectives for drug design. Antivir Res 87:125–148
Caillet-Saguy C, Simister PC, Bressanelli S (2011) An objective assessment of conformational variability in complexes of hepatitis C virus polymerase with non-nucleoside inhibitors. J Mol Biol 414:370–384
Caillet-Saguy C, Lim SP, Shi P-Y, Lescar J, Bressanelli S (2014) Polymerases of hepatitis C viruses and flaviviruses: structural and mechanistic insights and drug development. Antivir Res 105:8–16
Chung KY, Dong H, Chao AT, Shi P-Y, Lescar J, Lim SP (2010) Higher catalytic efficiency of N-7-methylation is responsible for processive N-7 and 2′-O methyltransferase activity in dengue virus. Virology 402:52–60
Davidson AD (2009) Chapter 2. New insights into flavivirus nonstructural protein 5. Adv Virus Res 74:41–101
Egloff MP, Benarroch D, Selisko B, Romette JL, Canard B (2002) An RNA cap (nucleoside-2′-O-)-methyltransferase in the flavivirus RNA polymerase NS5: crystal structure and functional characterization. EMBO J 21:2757–2768
Flisiak R, Janczewska E, Wawrzynowicz-Syczewska M, Jaroszewicz J, Zarębska-Michaluk D, Nazzal K, Bolewska B, Bialkowska J, Berak H, Fleischer-Stępniewska K, Tomasiewicz K, Karwowska K, Rostkowska K, Piekarska A, Tronina O, Madej G, Garlicki A, Lucejko M, Pisula A, Karpińska E, Kryczka W, Wiercińska-Drapało A, Mozer-Lisewska I, Jabłkowski M, Horban A, Knysz B, Tudrujek M, Halota W, Simon K (2016) Real-world effectiveness and safety of ombitasvir/paritaprevir/ritonavir ± dasabuvir ± ribavirin in hepatitis C: AMBER study. Aliment Pharmacol Ther 44(9):946–956
Koonin EV (1991) The phylogeny of RNA-dependent RNA polymerases of positive-strand RNA viruses. J Gen Virol 72:2197–2206
Koonin EV (1993) Computer-assisted identification of a putative methyltransferase domain in NS5 protein of flaviviruses and lambda 2 protein of reovirus. J Gen Virol 74:733–740
Kuntz ID, Chen K, Sharp KA, Kollman PA (1999) The maximal affinity of ligands. Proc Natl Acad Sci U S A 96(18):9997–10002
Leslie AGW (1992) Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4-ESF-EAMCB newsletter on protein crystallography, no. 26
Lim SP, Wen D, Yap TL, Yan CK, Lescar J, Vasudevan SG (2008) A scintillation proximity assay for dengue virus NS5 2′-O-methyltransferase-kinetic and inhibition analyses. Antivir Res 80:360–369
Lim SP, Koh JH, Seh CC, Liew CW, Davidson AD, Chua LS, Chandrasekaran R, Cornvik TC, Shi PY, Lescar J (2013) A crystal structure of the dengue virus non-structural protein 5 (NS5) polymerase delineates interdomain amino acid residues that enhance its thermostability and de novo initiation activities. J Biol Chem 288(43):31105–31114
Lim SP, Noble CG, Shi PY (2015) The dengue virus NS5 protein as a target for drug discovery. Antivir Res 119:57–67
Lim SP, Noble CG, Seh CC, Soh TS, El Sahili A, Chan GK, Lescar J, Arora R, Benson T, Nilar S, Manjunatha U, Wan KF, Dong H, Xie X, Shi PY, Yokokawa F (2016) Potent Allosteric Dengue Virus NS5 Polymerase Inhibitors: Mechanism of Action and Resistance Profiling. PLoS Pathog 12(8):e1005737
Malet H, Massé N, Selisko B, Romette JL, Alvarez K, Guillemot JC et al (2008) The flavivirus polymerase as a target for drug discovery. Antivir Res 80:23–35
Murshudov GN, Skubak P, Lebedev AA, Pannu NS, Steiner RA, Nicholls RA, Winn MD, Long F, Vagin AA (2011) REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D Biol Crystallogr 67:355–367
Niyomrattanakit P, Wan KF, Chung KY, Abas SN, Seh CC, Dong H et al (2015) Stabilization of dengue virus polymerase in de novo initiation assay provides advantages for compound screening. Antivir Res 119:36–46
Noble CG, Lim SP, Chen YL, Liew CW, Yap L, Lescar J, Shi PY (2013) Conformational flexibility of the dengue virus RNA-dependent RNA polymerase revealed by a complex with an inhibitor. J Virol 87:5291–5295
Noble CG, Lim SP, Arora R, Yokokawa F, Nilar S, Seh CC, Wright SK, Benson TE, Smith PW, Shi PY (2016) A conserved pocket in the dengue virus polymerase identified through fragment-based screening. J Biol Chem 291(16):8541–8548
Potisopon S, Priet S, Collet A, Decroly E, Canard B, Selisko B (2014) The methyltransferase domain of dengue virus protein NS5 ensures efficient RNA synthesis initiation and elongation by the polymerase domain. Nucleic Acids Res 42:11642–11656
Ray D, Shah A, Tilgner M, Guo Y, Zhao Y, Dong H, Deas TS, Zhou Y, Li H, Shi PY (2006) West Nile virus 5′-cap structure is formed by sequential guanine N-7 and ribose 2′-O methylations by nonstructural protein 5. J Virol 80:8362–8370
Tarantino D, Cannalire R, Mastrangelo E, Croci R, Querat G, Barreca ML, Bolognesi M, Manfroni G, Cecchetti V, Milani M (2016) Targeting flavivirus RNA dependent RNA polymerase through a pyridobenzothiazole inhibitor. Antivir Res 134:226–235
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AG, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Overview of the CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr 67:235–242
Yap TL, Xu T, Chen YL, Malet H, Egloff MP, Canard B et al (2007) Crystal structure of the dengue virus RNA-dependent RNA polymerase catalytic domain at 1.85-angstrom resolution. J Virol 81:4753–4765
Yokokawa F, Shahul N, Noble CG, Lim SP, Rao R, Tania S et al (2016) Discovery of potent non-nucleoside inhibitors of dengue viral RNA-dependent RNA polymerase from a fragment hit using structure-based drug design. J Med Chem 59:3935–3952
Zhao Y, Soh S, Zheng J, Chan KWK, Phoo WW, Lee CC et al (2015) A crystal structure of the dengue virus NS5 protein reveals a novel inter-domain interface essential for protein flexibility and virus replication. PLoS Pathog 11(3):e1004682
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Discussion of Chapter 14 in Dengue and Zika: Control and Antiviral Treatment Strategies
Discussion of Chapter 14 in Dengue and Zika: Control and Antiviral Treatment Strategies
This discussion was held at the 2nd Advanced Study Week on Emerging Viral Diseases at Praia do Tofo, Mozambique.Transcribed by Hilgenfeld R and Vasudevan SG (Eds); approved by Dr. Siew Pheng Lim.
-
Paul Young: Great work! A perfect example of a great collaboration between biology and chemistry. The work you describe seems to have hit a little bit the end of the road. You have identified and derived reasonably active sompounds but they do not appear to be effectively bioavailable. So what’s the next step?
-
Siew Pheng Lim: So this is only one part of the story. Our goal is to design compounds that extend out of the pocket. In the parlance of chemists in the team these extension from the pocket are done to find the so-called sweet spot to further improve the potency. But despite a lot of effort, we did not find another sweet spot. I also talked a little about some similarity the N-pocket shares with HCV palm 1 site. However HCV palm 1 pocket has a another wall contributed by the C-terminal part of the protein which allows those inhibitors to bindto both sides. In the case of Dengue virus polymerase we appear to have only one wall, we are trying to find a way to reach a second wall and that has not been successful so far.
-
David Jans: I just wanted to know what EC50 you have been happy with. Several of the compounds have EC50 of around 3 micromolar. Do you think that would be potent enough? In other words what should be the target for an effective drug?
-
Siew Pheng Lim: If I compare against the phenotypic screens that my colleagues do, they typically find inhibitors that are double-digit nanomolar in EC50. There they have a different challenge because they have issues with solubility and clearance as the compounds are too greasy. So I think for us if we can hit below one micromolar, We set ourselves a benchmark of less than one micromolar which we consider a good threshold. So you can see we have not quite reach that point yet.
-
Subhash Vasudevan: In the case of HCV palm 1, the compound that targeted a similar pocket actually got to the clinic implying that it is possible to develop a non-nucleosidic inhibitor. The pocket that you are looking is also truly a hotspot for Dengue and other flaviviruses. So the question I guess is do we give up on that area based on your experience so far?
-
Siew Pheng Lim: For now, this work has been put on hold, and we published the work so that other researchers in the field can expand on the effort and take a fresh look at approaches to target this important pocket. The search may include using other flaviviruses and find something we have not yet found to extend the chemistry. As David Jans alluded yesterday, we primarily work with DENV3 NS5 RdRp for which we have structural data. So it would be nice to have structural studies on other serotypes and other flaviviruses to see if something maybe different.
-
Subhash Vasudevan: But your data clearly shows that Dengue 2 and Dengue 4 give broadly similar results. In the discovery process if you are trying to do DENV1 to 4 separately then that would be a huge challenge. But NS5 is a really important target and since the non-nucleoside approach is not quite ready yet for pan-serotype inhibitor, what about the nucleoside aproach?
-
Siew Pheng Lim: Nucleoside approach is still ongoing. We started with KAB-344, the adenine analog (also known as NITD008) that eventually failed in the two-week animal toxicity studies and could not be progressed further. Since then we are working on other scaffolds. I think the issue with nucleosides is the unpredictable toxicity. To mitigate this we have implemented additional cytotoxicity assays trying to weed out toxicity early in the flowchart, such as mitochondrial assays –longer cytotoxicity assays – to capture those compounds that are potentially cytotoxic before we go into two-week toxicity studies in rats and dogs. Through this process, we have a few candidates that we are evaluating. The challenge with nucleoside inhibitors is the potentially lengthy scale up synthesis which can be expensive – so the way forward remains to be seen.
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Lim, S.P., Noble, C.G., Nilar, S., Shi, PY., Yokokawa, F. (2018). Discovery of Potent Non-nucleoside Inhibitors of Dengue Viral RNA-Dependent RNA Polymerase from Fragment Screening and Structure-Guided Design. In: Hilgenfeld, R., Vasudevan, S. (eds) Dengue and Zika: Control and Antiviral Treatment Strategies. Advances in Experimental Medicine and Biology, vol 1062. Springer, Singapore. https://doi.org/10.1007/978-981-10-8727-1_14
Download citation
DOI: https://doi.org/10.1007/978-981-10-8727-1_14
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8726-4
Online ISBN: 978-981-10-8727-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)