Abstract
Precise replication of genetic material and its equal distribution to daughter cells are essential to maintain genome stability. In eukaryotes, chromosome replication and segregation are temporally uncoupled, occurring in distinct intervals of the cell cycle, S and M phases, respectively. Cyclin E accumulates at the G1/S transition, where it promotes S phase entry and progression by binding to and activating CDK2. Several lines of evidence from different models indicate that cyclin E/CDK2 deregulation causes replication stress in S phase and chromosome segregation errors in M phase, leading to genomic instability and cancer. In this chapter, we will discuss the main findings that link cyclin E/CDK2 deregulation to genomic instability and the molecular mechanisms by which cyclin E/CDK2 induces replication stress and chromosome aberrations during carcinogenesis.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Cell cycle
- Cyclin E
- CDK2
- FBW7
- Replication stress
- Chromosome aberration
- Genomic instability
- Fragile sites
- Cancer
22.1 Introduction
Progression through the cell cycle is regulated by association of cyclin-dependent kinases (CDKs) with specific regulatory subunits known as cyclins. Oscillations in cyclin levels primarily dictate oscillations in CDK activity, ensuring the order and timing of cell cycle phases (Hochegger et al. 2008; Malumbres and Barbacid 2009). The E-type cyclin family is composed of two proteins, cyclin E1 and cyclin E2, which exhibit high sequence similarity and are functionally redundant (Lew et al. 1991; Koff et al. 1991; Gudas et al. 1999; Lauper et al. 1998; Zariwala et al. 1998). Cyclin E levels are tightly regulated during normal cell cycles, accumulating at the G1/S transition and being completely degraded by the end of S phase (Koff et al. 1992; Dulic et al. 1992). Consistent with its expression pattern, cyclin E binds to and activates CDK2 to control S phase entry and progression (Koff et al. 1992; Dulic et al. 1992; Ohtsubo and Roberts 1993; Resnitzky et al. 1994). Cyclin E mRNA levels are mostly induced by E2F transcription factors (Ohtani et al. 1995; Geng et al. 1996), whereas cyclin E protein is degraded by the SCFFbw7 ubiquitin ligase complex in a phosphorylation-dependent manner (Won and Reed 1996; Clurman et al. 1996; Strohmaier et al. 2001; Moberg et al. 2001; Koepp et al. 2001). Cyclin E/CDK2 activity is also controlled by the CDK inhibitors p21Cip1 and p27Kip, and potentially other CDK-inhibitory proteins, which are able to bind to and inactivate the cyclin E/CDK2 complex (Harper et al. 1993; Gu et al. 1993; Xiong et al. 1993; Polyak et al. 1994; Toyoshima and Hunter 1994; Reynaud et al. 1999).
Once activated, the cyclin E/CDK2 complex promotes the G1/S transition largely through phosphorylation and inactivation of the RB protein and the subsequent release of E2F transcription factors (Chellappan et al. 1991; Hinds et al. 1992; Dyson 1998; Harbour and Dean 2000). E2F proteins then promote S phase entry by regulating the expression of numerous genes required for DNA replication, such as the pre-replication complex components ORC1, CDC6, CDT1, and MCMs (Ohtani et al. 1996, 1998, 1999; Yan et al. 1998; Yoshida and Inoue 2004); the enzymes required for nucleotide and DNA synthesis, such as dihydrofolate reductase (DHFR), thymidine kinase (TK), and DNA polymerase α (Blake and Azizkhan 1989; Dou et al. 1994; Pearson et al. 1991); and the histone H2A (Oswald et al. 1996). Besides the RB protein, cyclin E/CDK2 directly phosphorylates and regulates other substrates required for S phase entry and progression, such as the DNA replication factors CDT1 and CDC6 (Liu et al. 2004; Mailand and Diffley 2005); the replication initiator Treslin (Kumagai et al. 2011); the activator of histone expression NPAT (Zhao et al. 2000; Ma et al. 2000); the transcription factors CBP/p300, E2F5, SMAD3, and MYC (Ait-Si-Ali et al. 1998; Morris et al. 2000; Matsuura et al. 2004; Hydbring et al. 2010); the centrosome proteins NPM, MPS1, and CP110 (Okuda et al. 2000; Tokuyama et al. 2001; Fisk and Winey 2001; Chen et al. 2002); and the DNA repair protein BRCA1 (Ruffner et al. 1999).
Regulation of E-type cyclins and the function of cyclin E/CDK2 in normal and aberrant cell cycles have been extensively reviewed elsewhere (Hwang and Clurman 2005; Caldon and Musgrove 2010; Siu et al. 2012). Here, we will focus on how deregulation of cyclin E/CDK2 causes replication stress and chromosome aberrations that may lead to genomic instability in cancer.
22.2 Cyclin E-Mediated Chromosome Instability
Several lines of evidence support the notion that cyclin E/CDK2 deregulation causes genomic instability. The initial finding that linked cyclin E to chromosome instability was the observation that constitutive cyclin E overexpression induced chromosome gains and losses in non-transformed rodent fibroblasts and human mammary epithelial cells, leading to aneuploidy (Spruck et al. 1999). Importantly, constitutive overexpression of cyclin D1 or cyclin A2 had no effect on the number of chromosomes in these cells. Later, it was shown that deletion of FBXW7, the gene encoding the F-box protein FBW7 involved in cyclin E recognition and degradation by the SCFFbw7 E3 ubiquitin ligase complex, resulted in increased frequency of micronucleus formation, multipolar spindles, and eventually chromosome instability in colorectal cancer cells (Rajagopalan et al. 2004). Even though FBW7 is involved in the degradation of other oncoproteins, such as c-MYC, c-JUN, and NOTCH (Davis et al. 2014), downregulation of cyclin E was sufficient to revert micronucleus formation in FBW7-depleted cells (Rajagopalan et al. 2004). More recently, generation of a hyperactive CDK2 knockin allele in a human colorectal cancer cell line that expresses high cyclin E-associated kinase activity also showed increased rates of micronucleus formation when compared to CDK2 wild-type cells (Hughes et al. 2013). Together, this evidence supports a causal role for cyclin E in chromosome instability during carcinogenesis.
One of the proposed mechanisms to explain chromosome instability in cancers is centrosome amplification, which leads to the formation of merotelic attachments and eventually chromosome segregation errors (Fig. 22.1) (Godinho and Pellman 2014). Normal cyclin E/CDK2 activity is required to ensure initiation of centrosome duplication in Xenopus egg extracts (Hinchcliffe et al. 1999; Lacey et al. 1999). In mammalian cells, it is also clear that CDK2 activity is necessary for centrosome duplication; however, it is still uncertain whether cyclin E or cyclin A plays a major role in CDK2-dependent centrosome duplication (Matsumoto et al. 1999; Meraldi et al. 1999; Hanashiro et al. 2008). Cyclin E overexpression alone does not efficiently induce centrosome amplification in mouse embryonic fibroblasts (MEFs), normal human fibroblasts, and epithelial cells (Spruck et al. 1999; Mussman et al. 2000; Kawamura et al. 2004). However, high levels of cyclin E synergize with loss of p53 function to induce centrosome amplification and chromosome instability in human cell lines and tumors (Mussman et al. 2000; Kawamura et al. 2004). Furthermore, MEFs from a hyperactive CDK2 knockin mouse model, which show elevated cyclin E- and cyclin A-associated kinase activities, had an elevated number of centrosomes when compared to wild-type MEFs (Zhao et al. 2012).
Mechanisms of cyclin E-induced genomic instability. Cyclin E/CDK2 deregulation may cause impaired assembly of pre-replication complex, increased origin initiation, deficiency of nucleotide biosynthesis pathway, collisions between replication and transcription machineries, formation of aberrant replication intermediates, such as fork reversal, centrosome amplification, and impairment of mitotic progression and checkpoint function
Cyclin E/CDK2 localizes to centrosomes (Matsumoto and Maller 2004), where it phosphorylates and dissociates NPM protein, initiating separation of paired centrioles and duplication of centrosomes (Okuda et al. 2000; Tokuyama et al. 2001). It is therefore possible that deregulation of cyclin E/CDK2 kinase activity, in combination with other insults, impairs NPM release from centrosomes, leading to centrosome amplification and chromosome instability. Indeed, deletion of NPM causes aberrant mitotic figures with multiple centrosomes and aneuploidy in MEFs (Grisendi et al. 2005), and alterations in NPM are frequently observed in human cancers (Grisendi et al. 2006). As discussed above, the centrosome proteins MPS1 and CP110 are also directly phosphorylated by cyclin E/CDK2 (Fisk and Winey 2001; Chen et al. 2002) and therefore may represent potential targets for cyclin E-induced chromosome instability as well.
Another mechanism that drives chromosome instability in tumorigenesis is impairment of mitotic checkpoint function and progression through mitosis, which may cause chromosome missegregation and aneuploidy (Fig. 22.1) (Varetti et al. 2014). It has been shown that cyclin E overexpression delays progression through early stages of mitosis, leading to accumulation of cells in prometaphase and unaligned metaphase (Keck et al. 2007). Impairment of mitotic progression was caused by cyclin E/CDK2-mediated phosphorylation and inactivation of the APC/C adaptor protein CDH1 and subsequent accumulation of the APC/CCdh1 ubiquitin ligase substrates cyclin B1 and securin, resulting in mitotic failure and polyploidy. In agreement, FBXW7-deficient cells, which have increased cyclin E-associated CDK2 activity, also exhibit increased levels of the APC/C substrates cyclin B1 and securin and accumulation of cells in prometaphase (Bailey et al. 2015). Interestingly, a genome-wide RNAi screen in these cells identified synthetic lethality with BUBR1, a spindle assembly checkpoint (SAC) protein, and high sensitivity to depletion of two other SAC components BUB1 and MPS1. These results suggest that cells with increased levels of cyclin E may depend on intact mitotic checkpoints for survival. Moreover, it has also been shown that cyclin E/CDK2 phosphorylates and prematurely activates the protein phosphatase CDC25C, leading to increased activity of the mitotic kinases cyclin B1/CDK1 and PLK1 and delayed mitotic progression (Bagheri-Yarmand et al. 2010).
22.3 Cyclin E-Mediated Replication Stress
Replication stress is characterized by the slowing or stalling of DNA replication forks, which may lead to fork collapse, DNA damage, and ultimately genomic instability (Zeman and Cimprich 2014). Activated oncoproteins and mutated tumor suppressors that drive sustained cellular proliferation cause replication stress and genomic instability, two events that are frequently observed in human cancers (Hills and Diffley 2014; Macheret and Halazonetis 2015). Indeed, this is the case for the oncoprotein cyclin E. Overexpression of cyclin E has been shown to cause replication stress, typified by slowed progression and premature termination of replication forks, DNA damage, and loss of heterozygosity at fragile sites (Bartkova et al. 2005, 2006; Bester et al. 2011). Cyclin E-mediated replication stress most likely is linked to elevated CDK2 kinase activity, as a CDK2 hyperactive knockin allele was sufficient to delay replication fork progression, induce DNA damage, and increase micronucleus formation without cyclin E overexpression (Hughes et al. 2013). In a seminal series of articles, it has been proposed that oncogene-induced replication stress, including the oncoprotein cyclin E, activates the DNA damage response (DDR) pathway and leads to cell cycle arrest, cell death, and senescence, acting as an inducible barrier to tumor progression (Bartkova et al. 2005, 2006; Gorgoulis et al. 2005; Di Micco et al. 2006). Disruption of the DDR pathway facilitates cell proliferation and increases replication stress, leading to genomic instability in preneoplastic lesions.
The primary mechanism underlying cyclin E/CDK2-induced replication stress and genomic instability is interference of the nucleotide biosynthesis pathway (Fig. 22.1). Nucleotides are structural components of nucleic acids and therefore essential for a wide variety of biological processes, such as cell growth, DNA replication, and transcription (Lane and Fan 2015). Cyclin E overexpression, through disruption of the RB/E2F pathway, enforces cell proliferation of human fibroblasts with insufficient nucleotide levels (Bester et al. 2011). Nucleotide deficiency induced by cyclin E overexpression slowed replication fork progression and caused double-strand DNA breaks. Importantly, either exogenous supplementation of nucleosides or upregulation of nucleotide metabolism genes attenuated cyclin E-mediated replication stress and DNA damage. Consistent with this, replication stress in the form of impaired fork progression has been shown to generate structural as well as numerical chromosome instability during mitosis (Burrell et al. 2013).
Collisions between DNA replication and transcription machineries are another important source of replication stress (Fig. 22.1). Transcription complexes represent natural obstacles to the progression of replication forks, especially at fragile sites that contain extremely long genes (>800 kb), where replication forks have a high probability of encountering transcription complexes during the period of one cell cycle (Helmrich et al. 2011). Transcription-replication collisions may generate increased DNA topological tension and formation of R-loops (RNA-DNA hybrid structures), inducing replication fork stalling, DNA damage, and fragile site instability (Bermejo et al. 2012; Helmrich et al. 2013). Oncogenic events that interfere with the timing and location of DNA replication and transcription may increase the probability of transcription-replication collisions. In cells overexpressing cyclin E, it has been shown that inhibition of transcription elongation attenuates replication stress and DNA damage (Jones et al. 2013). This study also showed that inhibition of replication initiation restores normal levels of fork progression in cyclin E-overexpressing cells, suggesting that increased replication initiation and transcription-replication collisions contribute to the replication stress upon high levels of cyclin E (Fig. 22.1). One potential consequence of transcription-replication collisions is the formation of aberrant replication intermediates, such as reversed replication forks (Neelsen and Lopes 2015). Consistently, it has been shown that cyclin E overexpression induces accumulation of reversed forks and chromosome breakage in human cells, suggesting that DNA topological stress also underlie cyclin E-mediated replication stress and genomic instability (Fig. 22.1) (Neelsen et al. 2013). Furthermore, it has been shown that cyclin E-induced collapsed forks may be processed and repaired by break-induced replication (BIR) repair, which generates copy number alterations, such as segmental genomic duplications (Costantino et al. 2014).
22.4 Genomic Instability in Cyclin E Mouse Models
Cyclin E is frequently overexpressed in human tumors, and its deregulation has been associated with poor prognosis and decreased survival of cancer patients (Scuderi et al. 1996; Porter et al. 1997; Iida et al. 1997; Erlanson et al. 1998; Fukuse et al. 2000; Muller-Tidow et al. 2001; Keyomarsi et al. 2002; Schraml et al. 2003). Overexpression of cyclin E in mouse models has been shown to induce mammary and lung carcinomas as well as hematopoietic malignancies, further supporting a causative role for cyclin E in carcinogenesis (Bortner and Rosenberg 1997; Karsunky et al. 1999; Geisen et al. 2003; Loeb et al. 2005; Smith et al. 2006; Ma et al. 2007; Minella et al. 2008; Siu et al. 2014).
Several tissue-specific transgenic and knockin mouse models have provided significant information on the role of cyclin E deregulation in genomic instability. A knockin mouse expressing a nondegradable form of cyclin E in MEFs showed increased chromosome breaks, translocations, and aneuploidy in a p21−/− background (Loeb et al. 2005). In this model, cyclin E overexpression also cooperated with p53 deficiency and RAS activation to cause cellular transformation, induce whole chromosome gains and losses, and accelerate lung carcinogenesis. Consistent with this, transgenic mice expressing either wild-type or degradation-resistant cyclin E in the lungs incurred multiple pulmonary adenocarcinomas with specific gains of chromosomes 4 and 6 (Ma et al. 2007). In mammary gland transgenic mouse models, cyclin E overexpression has been shown to induce p53 loss of heterozygosity and drastically increase tumor formation in a p53+/− background (Smith et al. 2006; Akli et al. 2007). Lastly, a knockin mouse with expression of nondegradable cyclin E in the hematopoietic stem cell compartment exhibited abnormal hematopoiesis, chromosome instability illustrated by chromosome gains and losses, and decreased latency of T-cell malignancies in a p53−/− background (Minella et al. 2008; Siu et al. 2014). Again, p53 and p21 deficiencies were synergistic with cyclin E deregulation in promoting chromosome instability. Indeed, it has been shown that cyclin E-associated genomic instability is restrained by the p53/p21 pathway (Bartkova et al. 2005; Minella et al. 2002, 2007). Disruption of the inducible barrier established by the p53/p21 pathway may allow cyclin E overexpression to trigger genomic instability through some of the mechanisms discussed above, such as centrosome amplification and replication stress. Therefore, current mouse models support the notion that cyclin E deregulation contributes to tumorigenesis by promoting genomic instability in vivo.
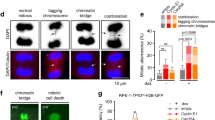
22.5 Cyclin E Deregulation Promotes Replication Failure at Targeted Sites
We have discussed above the relationship between cyclin E and replication stress. Since cyclin E overexpression promotes replication stress and therefore slows replication fork progression (Bester et al. 2011; Jones et al. 2013; Liberal et al. 2012), we hypothesized that cells experiencing cyclin E deregulation might enter mitosis with incompletely replicated chromosomes. This in turn would lead to abnormal segregation and chromosomal damage during anaphase. Consistent with this, we observed that cyclin E-overexpressing non-transformed cells exhibited high levels of anaphase chromosomal anomalies such as bridged chromosomes and nonattached chromosomal fragments up to the size of the entire chromosome arms (Teixeira et al. 2015). If this observed chromosomal damage is a result of incompletely replicated chromosomal segments impairing segregation of sister chromatids, there are two obvious models that could account for the under-replication. Cyclin E-mediated replication stress could promote under-replication without any regional or feature specificity, or under-replication could occur at specific sites or regions possessing features that might sensitize them. To distinguish between these alternatives, we harvested cells blocked in mitosis immediately following cyclin E overexpression and analyzed their DNA by comparative genomic hybridization (CGH) array analysis. Indeed, a relatively small number of specific regions varying in size from approximately 200 to 100,000 base pairs had frequently failed to complete replication prior to entry into mitosis (Teixeira et al. 2015). Presumably, these under-replicated regions were responsible for the anaphase segregation anomalies we had observed in real time after cyclin E overexpression. Based on these observations, one would predict that these under-replicated regions would be included in deleted chromosomal segments subsequent to anaphase. We interrogated both mixed populations and single cells after cyclin E overexpression and found that deletion of these specific loci did indeed occur at high frequency (Teixeira et al. 2015). However, it appears that most cells carrying such deletions were incapable of clonal expansion, suggesting that checkpoint barriers eliminate cells with severely damaged genomes, should surveillance mechanisms be intact. This presumably reduces the impact of cyclin E-mediated damage at the population level and is therefore protective from potentially oncogenic events (Bartkova et al. 2005). Indeed, the link between cyclin E deregulation and the p53 surveillance system has been discussed above.
22.6 Genomic Features Associated with Replication Failure
22.6.1 Late-Replicating Genomic Regions
The majority of under-replicated regions detected in our study have been annotated as large late-replicating domains (Teixeira et al. 2015; Weddington et al. 2008). These are, for the most part, heterochromatic regions with a paucity of replication origins. The fact that origins in these domains fire late during the replication cycle combined with the low density of origins provides a likely explanation for failure to complete replication under conditions of replication stress (Le Tallec et al. 2014; Ozeri-Galai et al. 2014). However, these properties alone cannot explain the specific locations and boundaries of the under-replicated regions, as they were relatively small compared to the larger domains and highly targeted to specific sites. Therefore, other features of these sites must be relevant.
22.6.2 Recombinational Hotspots/Translocation Breakpoints
A number of the sites have been annotated as recombinational hotspots or translocation breakpoints. A subset of these has been classified as fragile sites, as well. Both recombination and translocation are processes that are initiated by double-strand DNA breaks. Repair involving homologous sequences versus heterologous sequences containing microhomology determines the outcome (Berti and Vindigni 2016). Significantly, within this context, one characteristic of fragile sites is a tendency to experience double-strand breaks at abnormally high frequencies, presumably due to replication barriers within these sites (Le Tallec et al. 2014; Ozeri-Galai et al. 2014; Thys et al. 2015). These observations suggest that features of fragile sites might impede DNA replication under conditions of cyclin E-mediated replication stress leading to local replication failure. One feature of fragile sites suggested as causative for replication impairment and double-strand breaks is unusual DNA structures, such as palindromic sequences leading to formation of hairpins and loops (Thys et al. 2015; Ozeri-Galai et al. 2011). Such structured nonlinear DNA could easily explain why stressed replication forks might stall or collapse. However, this cannot completely explain the specificity of under-replicated sites under conditions of cyclin E overexpression, since only a small subset of fragile sites, recombinational hotspots, and translocation breakpoints is affected. In addition, two recent studies showing that cyclin E overexpression causes instability at genomic regions with fragile site characteristics have given somewhat different results (Teixeira et al. 2015; Miron et al. 2015). Interestingly, among the susceptible genomic regions identified on each study (16 and 26, respectively), only one chromosome band was coincident for both (3q26). Since one study was carried out in mammary epithelial cells and the other in fibroblasts, a possible interpretation of the data is that cyclin E-mediated fragility may also be cell type specific, as has been shown previously in cells from different tissues (Le Tallec et al. 2011, 2013; Hosseini et al. 2013). In addition, not every under-replicated site was associated with fragile site features, suggesting factors unique to cyclin E-mediated replication stress must come into play (see below).
22.6.3 Low Origin Density and Licensing
Interrogating a number of databases of origin distribution in human cells compiled using diverse methodologies, we found that most of the under-replicated sites were located in chromosomal regions characterized by extremely low origin density (Teixeira et al. 2015). Under conditions of replication stress leading to fork collapse, the probable lack of nearby functional forks is likely to eliminate the most common mechanism for rescue of localized replication failure: processing of unreplicated DNA by an adjacent replicon (Letessier et al. 2011; Kawabata et al. 2011). However, it is likely that replication stress caused specifically by cyclin E overexpression compounds the logistical problems of completing the replicative cycle. This is because, in addition to replication stress, cyclin E overexpression impairs assembly of the pre-replication complex (Ekholm-Reed et al. 2004). Specifically, high cyclin E/CDK2 activity at the M/G1 boundary inhibits chromatin loading of MCM proteins, which constitute the primary replicative helicase. Based on investigation of the impact of direct MCM protein depletion, it is unlikely that this effect of cyclin E overexpression would alter DNA replication during an unperturbed replicative cycle. Very high percentages of individual MCM proteins can be depleted via RNAi-mediated silencing with no apparent effect on unperturbed replication (Ge et al. 2007; Ibarra et al. 2008). However, MCM-depleted cells are extremely sensitive to replication stress, as they fail to assemble backup origins. These origins are competent but normally remain dormant except under conditions of replication stress, when they are mobilized to rescue collapsed and/or poorly functioning replication forks (McIntosh and Blow 2012). Since it is probable that cyclin E overexpression via impaired MCM loading leads to a deficiency of backup origins, but also simultaneously causes replication stress, the problem of rescuing collapsed forks in origin-sparse regions is likely exacerbated, leading to replication failure.
22.6.4 Transcription-Replication Collisions
As stated above, one possible source of cyclin E-mediated replication stress is collision between the replication and transcriptional machineries. These encounters would be predicted to occur most frequently at fragile sites containing very long genes (>300 kb). Consistent with this, cyclin E overexpression produced copy number losses at two very long genes, EPHA6 and NCAM2, in human mammary epithelial cells (approximate size of 935 kb and 545 kb, respectively) (Teixeira et al. 2015). In addition, transcriptional changes were observed at two other long genes, DAB1 and NRXN3, in human fibroblasts (approximate size of 430 kb and 1.7 Mb, respectively) experiencing deregulated levels of cyclin E (Miron et al. 2015).
22.6.5 Sensitive DNA Structures
As alluded to the above, a number of the sites under-replicated after cyclin E overexpression correspond to fragile sites, a characteristic of which is the presence of nonlinear DNA structures expected to pose barriers to replication fork progression (Thys et al. 2015). However, most fragile sites were not represented as under-replicated sites in our analysis. One fragile site, nevertheless, is likely to be informative: FRA11B/G (Fechter et al. 2007; Burrow et al. 2009). This site on chromosome 11 is particularly interesting because it is the breakpoint for rearrangements in mixed-lineage leukemia (MLL); hence, the locus is referred to as MLL (Muntean and Hess 2012). It is also noteworthy that this site is also frequently deleted in breast cancer (see below). The under-replicated segment detected in our study was 4,331 bp, which contains part of the breakpoint cluster region (BCR) for MLL translocations in leukemia (Muntean and Hess 2012). We therefore scanned this region for DNA sequences predictive of the ability to form energetically favorable hairpin loop structures. Interestingly, two such structures were detected near the center of the segment separated by approximately 500 base pairs (Teixeira et al. 2015). To determine whether these structured DNA elements posed a barrier to replication under conditions of cyclin E-mediated replication stress, we cloned the segment containing them, with and without the palindromic sequences, into an episomal vector that replicates autonomously in mammalian cells. While both the control and palindrome-containing plasmids were well maintained in the absence of cyclin E overexpression, only the control plasmid was maintained when cyclin E was overexpressed. These data are consistent with the hypothesis that palindromic structures pose a barrier to replication, specifically under conditions of cyclin E-mediated replication stress (Teixeira et al. 2015). Therefore, two structural barriers to replication in close proximity may represent a feature that promotes sensitivity to cyclin E-mediated replication stress, although such structures are likely to be sensitive to other sources of replication stress as well (see below). However, it is worth noting that cyclin E overexpression has been determined to be associated with MLL translocations in the context of B-cell precursor acute lymphoblastic leukemia (BCP-ALL) (Accordi et al. 2010).
22.7 Cyclin E and Genomic Instability in Breast Cancer
What does the observation that cyclin E overexpression promotes replication failure at a small subset of specific loci implicate for oncogenesis? The first question one might ask is are deletions in the chromosomal regions surrounding the under-replicated sites found in actual cancer? To address this, we interrogated a database of approximately 2000 breast tumors subjected to CGH array analysis (Teixeira et al. 2015; Curtis et al. 2012). As a surrogate for cyclin E overexpression, we employed copy number increases of the CCNE1 locus, presumably due to gene amplification. This undoubtedly represents an underestimation of tumors overexpressing cyclin E and likely introduces experimental noise, as several posttranslational mechanisms have been shown to elevate cyclin E levels. Nonetheless, when CCNE1 copy number increase was compared with copy number decrease at each of the under-replicated sites, a highly significant correlation was observed for many of them. This suggests that cyclin E overexpression is a driver of deletion at these sites. It should be noted that overall these specific sites experience deletions at relatively low frequency, and their detailed analysis is likely to yield clues concerning what characteristics constitute sensitivity specifically to cyclin E-mediated replication stress. On the other hand, some of the sites that were more frequently deleted in the data set did not show a significant correlation with cyclin E copy number increase. The MLL locus was one of these. Presumably, features of the MLL locus, specifically two likely hairpin loops in close proximity, render this site sensitive to multiple forms of replication stress, including but not exclusive to cyclin E. The second relevant question is whether these specific deletions have a direct role in oncogenesis. Unfortunately, at this point, we do not know the magnitudes or boundaries of deletions that occur surrounding these sites when they have not completed replication but are forced through anaphase. However, one might speculate that large deletions could drive oncogenesis by promoting loss of heterozygosity at tumor suppressor loci and deletions over fragile sites (Bignell et al. 2010). It is interesting to note that tumor suppressor genes have been identified on chromosomes 3q (Guo et al. 2002; Schwaenen et al. 2009; Thean et al. 2010) and 21q (Lee et al. 2003; Silva et al. 2003; Yamada et al. 2008), the arms where cyclin E-driven deletions occur in breast cancer.
In our study using immortalized non-transformed human mammary epithelial cells, it was clear that few cells that had sustained deletions after exposure to cyclin E overexpression were capable of expanding robustly and forming colonies. Although reassuring from a human health perspective, this observation raises the question of how cyclin E-driven deletions might get fixed in an expanding premalignant population. The answer probably lies in the fact that individual breast cancers when they present commonly possess more than 100 genetic modifications (Nik-Zainal et al. 2016). At least some of these are likely to have been selected to override checkpoint barriers, thereby allowing clonal expansion of chromosomally damaged cells.
22.8 Relevance to Other Sources of Replication Stress
The discussion above has focused on mechanisms of genomic instability associated with cyclin E overexpression/deregulation (Fig. 22.1). However, other oncogenic events have been associated with replication stress, e.g., overexpression of c-MYC (Dominguez-Sola et al. 2007; Srinivasan et al. 2013; Rohban and Campaner 2015) or mutation of RAS (Di Micco et al. 2006; Miron et al. 2015; Maya-Mendoza et al. 2015). Although the mechanisms whereby these overexpressed or mutant proteins cause replication stress are likely to differ, one can infer that in severe instances, incompletely replicated genomes will enter mitosis with the end result being genomic instability.
A number of model system experiments support this idea. A hypomorphic allele of the mouse MCM helicase component MCM4, designated Chaos3, which promotes instability of the pre-replication complex (Kawabata et al. 2011), causes anaphase aberrations similar to what we observed for cyclin E overexpression. Although MCM4Chaos3 does not appear to affect the number of origins fired, the number of dormant backup origins is reduced, and the number of stalled replication forks is increased, as it has been proposed for cyclin E overexpression. Presumably, intrinsic levels of replication stress during the normal replicative cycle require such backup origins in order to avoid under-replicated regions and the resulting aberrant anaphases. Interestingly, the MCM4Chaos3 mouse is cancer prone (Shima et al. 2007), indicating that this type of chromosomal damage is directly linked to oncogenesis.
In yeast, the absence of the cohesin-like complex Smc5-Smc6 causes replication impairment at loci that contain replication barriers, such as the rDNA cluster (Torres-Rosell et al. 2007). Yet, cells progress through mitosis resulting in elevated rates of chromosomal nondisjunction, even though all checkpoints are intact. These results confirm that no robust checkpoint exists that can detect and respond to small quantities of unreplicated DNA in eukaryotes ranging from yeast to human (Mohebi et al. 2015; Koundrioukoff et al. 2013).
22.9 Conclusions
Elevated cyclin E has been shown to be associated with aggressive disease and poor outcome in at least some human malignancies. The link between cyclin E overexpression and genomic instability suggests a mechanism whereby cyclin E might promote oncogenesis. Our recent work showing that cyclin E overexpression promotes replication failure at a small number of specific loci and, as a consequence, chromosomal damage following anaphase leaves some important unanswered questions relevant to oncogenesis. First and foremost is a detailed description of the genomic damage that occurs in individual cells. Such information will allow the assessment of whether the classic two-hit tumor suppressor model is applicable or whether more complex mechanisms apply such as amplifications and translocations. The second important issue to be resolved is whether cyclin E can serve as a prototype for other oncoproteins that cause replication stress and promote genomic instability. On the one hand, as outlined above, some of the modalities of cyclin E function in the context of the replication stress are likely to be unique. On the other, there are certain to be mechanistic commonalities. Only investigation of the pathways leading to replication stress and from replication stress to genomic alterations for other oncoproteins will resolve this question.
References
Accordi B, Espina V, Giordan M, VanMeter A, Milani G, Galla L, Ruzzene M, Sciro M, Trentin L, De Maria R, te Kronnie G, Petricoin E, Liotta L, Basso G (2010) Functional protein network activation mapping reveals new potential molecular drug targets for poor prognosis pediatric BCP-ALL. PLoS One 5(10):e13552. https://doi.org/10.1371/journal.pone.0013552
Ait-Si-Ali S, Ramirez S, Barre FX, Dkhissi F, Magnaghi-Jaulin L, Girault JA, Robin P, Knibiehler M, Pritchard LL, Ducommun B, Trouche D, Harel-Bellan A (1998) Histone acetyltransferase activity of CBP is controlled by cycle-dependent kinases and oncoprotein E1A. Nature 396(6707):184–186. https://doi.org/10.1038/24190
Akli S, Van Pelt CS, Bui T, Multani AS, Chang S, Johnson D, Tucker S, Keyomarsi K (2007) Overexpression of the low molecular weight cyclin E in transgenic mice induces metastatic mammary carcinomas through the disruption of the ARF-p53 pathway. Cancer Res 67(15):7212–7222. https://doi.org/10.1158/0008-5472.can-07-0599
Bagheri-Yarmand R, Nanos-Webb A, Biernacka A, Bui T, Keyomarsi K (2010) Cyclin E deregulation impairs mitotic progression through premature activation of Cdc25C. Cancer Res 70(12):5085–5095. https://doi.org/10.1158/0008-5472.can-09-4095
Bailey ML, Singh T, Mero P, Moffat J, Hieter P (2015) Dependence of human colorectal cells lacking the FBW7 tumor suppressor on the spindle assembly checkpoint. Genetics 201(3):885–895. https://doi.org/10.1534/genetics.115.180653
Bartkova J, Horejsi Z, Koed K, Kramer A, Tort F, Zieger K, Guldberg P, Sehested M, Nesland JM, Lukas C, Orntoft T, Lukas J, Bartek J (2005) DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434(7035):864–870. https://doi.org/10.1038/nature03482
Bartkova J, Rezaei N, Liontos M, Karakaidos P, Kletsas D, Issaeva N, Vassiliou LV, Kolettas E, Niforou K, Zoumpourlis VC, Takaoka M, Nakagawa H, Tort F, Fugger K, Johansson F, Sehested M, Andersen CL, Dyrskjot L, Orntoft T, Lukas J, Kittas C, Helleday T, Halazonetis TD, Bartek J, Gorgoulis VG (2006) Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 444(7119):633–637. https://doi.org/10.1038/nature05268
Bermejo R, Lai MS, Foiani M (2012) Preventing replication stress to maintain genome stability: resolving conflicts between replication and transcription. Mol Cell 45(6):710–718. https://doi.org/10.1016/j.molcel.2012.03.001
Berti M, Vindigni A (2016) Replication stress: getting back on track. Nat Struct Mol Biol 23(2):103–109. https://doi.org/10.1038/nsmb.3163
Bester AC, Roniger M, Oren YS, Im MM, Sarni D, Chaoat M, Bensimon A, Zamir G, Shewach DS, Kerem B (2011) Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 145(3):435–446. https://doi.org/10.1016/j.cell.2011.03.044
Bignell GR, Greenman CD, Davies H, Butler AP, Edkins S, Andrews JM, Buck G, Chen L, Beare D, Latimer C, Widaa S, Hinton J, Fahey C, Fu B, Swamy S, Dalgliesh GL, Teh BT, Deloukas P, Yang F, Campbell PJ, Futreal PA, Stratton MR (2010) Signatures of mutation and selection in the cancer genome. Nature 463(7283):893–898. https://doi.org/10.1038/nature08768
Blake MC, Azizkhan JC (1989) Transcription factor E2F is required for efficient expression of the hamster dihydrofolate reductase gene in vitro and in vivo. Mol Cell Biol 9(11):4994–5002
Bortner DM, Rosenberg MP (1997) Induction of mammary gland hyperplasia and carcinomas in transgenic mice expressing human cyclin E. Mol Cell Biol 17(1):453–459
Burrell RA, McClelland SE, Endesfelder D, Groth P, Weller MC, Shaikh N, Domingo E, Kanu N, Dewhurst SM, Gronroos E, Chew SK, Rowan AJ, Schenk A, Sheffer M, Howell M, Kschischo M, Behrens A, Helleday T, Bartek J, Tomlinson IP, Swanton C (2013) Replication stress links structural and numerical cancer chromosomal instability. Nature 494(7438):492–496. https://doi.org/10.1038/nature11935
Burrow AA, Williams LE, Pierce LC, Wang YH (2009) Over half of breakpoints in gene pairs involved in cancer-specific recurrent translocations are mapped to human chromosomal fragile sites. BMC Genomics 10:59. https://doi.org/10.1186/1471-2164-10-59
Caldon CE, Musgrove EA (2010) Distinct and redundant functions of cyclin E1 and cyclin E2 in development and cancer. Cell Div 5:2. https://doi.org/10.1186/1747-1028-5-2
Chellappan SP, Hiebert S, Mudryj M, Horowitz JM, Nevins JR (1991) The E2F transcription factor is a cellular target for the RB protein. Cell 65(6):1053–1061
Chen Z, Indjeian VB, McManus M, Wang L, Dynlacht BD (2002) CP110, a cell cycle-dependent CDK substrate, regulates centrosome duplication in human cells. Dev Cell 3(3):339–350
Clurman BE, Sheaff RJ, Thress K, Groudine M, Roberts JM (1996) Turnover of cyclin E by the ubiquitin-proteasome pathway is regulated by cdk2 binding and cyclin phosphorylation. Genes Dev 10(16):1979–1990
Costantino L, Sotiriou SK, Rantala JK, Magin S, Mladenov E, Helleday T, Haber JE, Iliakis G, Kallioniemi OP, Halazonetis TD (2014) Break-induced replication repair of damaged forks induces genomic duplications in human cells. Science 343(6166):88–91. https://doi.org/10.1126/science.1243211
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Langerod A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Borresen-Dale AL, Brenton JD, Tavare S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486(7403):346–352. https://doi.org/10.1038/nature10983
Davis RJ, Welcker M, Clurman BE (2014) Tumor suppression by the Fbw7 ubiquitin ligase: mechanisms and opportunities. Cancer Cell 26(4):455–464. https://doi.org/10.1016/j.ccell.2014.09.013
Di Micco R, Fumagalli M, Cicalese A, Piccinin S, Gasparini P, Luise C, Schurra C, Garre M, Nuciforo PG, Bensimon A, Maestro R, Pelicci PG, d’Adda di Fagagna F (2006) Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 444(7119):638–642. https://doi.org/10.1038/nature05327
Dominguez-Sola D, Ying CY, Grandori C, Ruggiero L, Chen B, Li M, Galloway DA, Gu W, Gautier J, Dalla-Favera R (2007) Non-transcriptional control of DNA replication by c-Myc. Nature 448(7152):445–451. https://doi.org/10.1038/nature05953
Dou QP, Zhao S, Levin AH, Wang J, Helin K, Pardee AB (1994) G1/S-regulated E2F-containing protein complexes bind to the mouse thymidine kinase gene promoter. J Biol Chem 269(2):1306–1313
Dulic V, Lees E, Reed SI (1992) Association of human cyclin E with a periodic G1-S phase protein kinase. Science 257(5078):1958–1961
Dyson N (1998) The regulation of E2F by pRB-family proteins. Genes Dev 12(15):2245–2262
Ekholm-Reed S, Mendez J, Tedesco D, Zetterberg A, Stillman B, Reed SI (2004) Deregulation of cyclin E in human cells interferes with prereplication complex assembly. J Cell Biol 165(6):789–800. https://doi.org/10.1083/jcb.200404092
Erlanson M, Portin C, Linderholm B, Lindh J, Roos G, Landberg G (1998) Expression of cyclin E and the cyclin-dependent kinase inhibitor p27 in malignant lymphomas-prognostic implications. Blood 92(3):770–777
Fechter A, Buettel I, Kuehnel E, Savelyeva L, Schwab M (2007) Common fragile site FRA11G and rare fragile site FRA11B at 11q23.3 encompass distinct genomic regions. Genes Chromosom Cancer 46(1):98–106. https://doi.org/10.1002/gcc.20389
Fisk HA, Winey M (2001) The mouse Mps1p-like kinase regulates centrosome duplication. Cell 106(1):95–104
Fukuse T, Hirata T, Naiki H, Hitomi S, Wada H (2000) Prognostic significance of cyclin E overexpression in resected non-small cell lung cancer. Cancer Res 60(2):242–244
Ge XQ, Jackson DA, Blow JJ (2007) Dormant origins licensed by excess Mcm2-7 are required for human cells to survive replicative stress. Genes Dev 21(24):3331–3341. https://doi.org/10.1101/gad.457807
Geisen C, Karsunky H, Yucel R, Moroy T (2003) Loss of p27Kip1 cooperates with cyclin E in T-cell lymphomagenesis. Oncogene 22(11):1724–1729. https://doi.org/10.1038/sj.onc.1206340
Geng Y, Eaton EN, Picon M, Roberts JM, Lundberg AS, Gifford A, Sardet C, Weinberg RA (1996) Regulation of cyclin E transcription by E2Fs and retinoblastoma protein. Oncogene 12(6):1173–1180
Godinho SA, Pellman D (2014) Causes and consequences of centrosome abnormalities in cancer. Philos Trans R Soc Lond B Biol Sci 369(1650):20130467. https://doi.org/10.1098/rstb.2013.0467
Gorgoulis VG, Vassiliou LV, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T, Venere M, Ditullio RA Jr, Kastrinakis NG, Levy B, Kletsas D, Yoneta A, Herlyn M, Kittas C, Halazonetis TD (2005) Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434(7035):907–913. https://doi.org/10.1038/nature03485
Grisendi S, Bernardi R, Rossi M, Cheng K, Khandker L, Manova K, Pandolfi PP (2005) Role of nucleophosmin in embryonic development and tumorigenesis. Nature 437(7055):147–153. https://doi.org/10.1038/nature03915
Grisendi S, Mecucci C, Falini B, Pandolfi PP (2006) Nucleophosmin and cancer. Nat Rev Cancer 6(7):493–505. https://doi.org/10.1038/nrc1885
Gu Y, Turck CW, Morgan DO (1993) Inhibition of CDK2 activity in vivo by an associated 20K regulatory subunit. Nature 366(6456):707–710. https://doi.org/10.1038/366707a0
Gudas JM, Payton M, Thukral S, Chen E, Bass M, Robinson MO, Coats S (1999) Cyclin E2, a novel G1 cyclin that binds Cdk2 and is aberrantly expressed in human cancers. Mol Cell Biol 19(1):612–622
Guo SS, Arora C, Shimoide AT, Sawicki MP (2002) Frequent deletion of chromosome 3 in malignant sporadic pancreatic endocrine tumors. Mol Cell Endocrinol 190(1–2):109–114
Hanashiro K, Kanai M, Geng Y, Sicinski P, Fukasawa K (2008) Roles of cyclins A and E in induction of centrosome amplification in p53-compromised cells. Oncogene 27(40):5288–5302. https://doi.org/10.1038/onc.2008.161
Harbour JW, Dean DC (2000) The Rb/E2F pathway: expanding roles and emerging paradigms. Genes Dev 14(19):2393–2409
Harper JW, Adami GR, Wei N, Keyomarsi K, Elledge SJ (1993) The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 75(4):805–816
Helmrich A, Ballarino M, Tora L (2011) Collisions between replication and transcription complexes cause common fragile site instability at the longest human genes. Mol Cell 44(6):966–977. https://doi.org/10.1016/j.molcel.2011.10.013
Helmrich A, Ballarino M, Nudler E, Tora L (2013) Transcription-replication encounters, consequences and genomic instability. Nat Struct Mol Biol 20(4):412–418. https://doi.org/10.1038/nsmb.2543
Hills SA, Diffley JF (2014) DNA replication and oncogene-induced replicative stress. Curr Biol 24(10):R435–R444. https://doi.org/10.1016/j.cub.2014.04.012
Hinchcliffe EH, Li C, Thompson EA, Maller JL, Sluder G (1999) Requirement of Cdk2-cyclin E activity for repeated centrosome reproduction in Xenopus egg extracts. Science 283(5403):851–854
Hinds PW, Mittnacht S, Dulic V, Arnold A, Reed SI, Weinberg RA (1992) Regulation of retinoblastoma protein functions by ectopic expression of human cyclins. Cell 70(6):993–1006
Hochegger H, Takeda S, Hunt T (2008) Cyclin-dependent kinases and cell-cycle transitions: does one fit all? Nat Rev Mol Cell Biol 9(11):910–916. https://doi.org/10.1038/nrm2510
Hosseini SA, Horton S, Saldivar JC, Miuma S, Stampfer MR, Heerema NA, Huebner K (2013) Common chromosome fragile sites in human and murine epithelial cells and FHIT/FRA3B loss-induced global genome instability. Genes Chromosom Cancer 52(11):1017–1029. https://doi.org/10.1002/gcc.22097
Hughes BT, Sidorova J, Swanger J, Monnat RJ Jr, Clurman BE (2013) Essential role for Cdk2 inhibitory phosphorylation during replication stress revealed by a human Cdk2 knockin mutation. Proc Natl Acad Sci U S A 110(22):8954–8959. https://doi.org/10.1073/pnas.1302927110
Hwang HC, Clurman BE (2005) Cyclin E in normal and neoplastic cell cycles. Oncogene 24(17):2776–2786. https://doi.org/10.1038/sj.onc.1208613
Hydbring P, Bahram F, Su Y, Tronnersjo S, Hogstrand K, von der Lehr N, Sharifi HR, Lilischkis R, Hein N, Wu S, Vervoorts J, Henriksson M, Grandien A, Luscher B, Larsson LG (2010) Phosphorylation by Cdk2 is required for Myc to repress Ras-induced senescence in cotransformation. Proc Natl Acad Sci U S A 107(1):58–63. https://doi.org/10.1073/pnas.0900121106
Ibarra A, Schwob E, Mendez J (2008) Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. Proc Natl Acad Sci U S A 105(26):8956–8961. https://doi.org/10.1073/pnas.0803978105
Iida H, Towatari M, Tanimoto M, Morishita Y, Kodera Y, Saito H (1997) Overexpression of cyclin E in acute myelogenous leukemia. Blood 90(9):3707–3713
Jones RM, Mortusewicz O, Afzal I, Lorvellec M, Garcia P, Helleday T, Petermann E (2013) Increased replication initiation and conflicts with transcription underlie cyclin E-induced replication stress. Oncogene 32(32):3744–3753. https://doi.org/10.1038/onc.2012.387
Karsunky H, Geisen C, Schmidt T, Haas K, Zevnik B, Gau E, Moroy T (1999) Oncogenic potential of cyclin E in T-cell lymphomagenesis in transgenic mice: evidence for cooperation between cyclin E and Ras but not Myc. Oncogene 18(54):7816–7824. https://doi.org/10.1038/sj.onc.1203205
Kawabata T, Luebben SW, Yamaguchi S, Ilves I, Matise I, Buske T, Botchan MR, Shima N (2011) Stalled fork rescue via dormant replication origins in unchallenged S phase promotes proper chromosome segregation and tumor suppression. Mol Cell 41(5):543–553. https://doi.org/10.1016/j.molcel.2011.02.006
Kawamura K, Izumi H, Ma Z, Ikeda R, Moriyama M, Tanaka T, Nojima T, Levin LS, Fujikawa-Yamamoto K, Suzuki K, Fukasawa K (2004) Induction of centrosome amplification and chromosome instability in human bladder cancer cells by p53 mutation and cyclin E overexpression. Cancer Res 64(14):4800–4809. https://doi.org/10.1158/0008-5472.can-03-3908
Keck JM, Summers MK, Tedesco D, Ekholm-Reed S, Chuang LC, Jackson PK, Reed SI (2007) Cyclin E overexpression impairs progression through mitosis by inhibiting APCCdh1. J Cell Biol 178(3):371–385. https://doi.org/10.1083/jcb.200703202
Keyomarsi K, Tucker SL, Buchholz TA, Callister M, Ding Y, Hortobagyi GN, Bedrosian I, Knickerbocker C, Toyofuku W, Lowe M, Herliczek TW, Bacus SS (2002) Cyclin E and survival in patients with breast cancer. N Engl J Med 347(20):1566–1575. https://doi.org/10.1056/NEJMoa021153
Koepp DM, Schaefer LK, Ye X, Keyomarsi K, Chu C, Harper JW, Elledge SJ (2001) Phosphorylation-dependent ubiquitination of cyclin E by the SCFFbw7 ubiquitin ligase. Science 294(5540):173–177. https://doi.org/10.1126/science.1065203
Koff A, Cross F, Fisher A, Schumacher J, Leguellec K, Philippe M, Roberts JM (1991) Human cyclin E, a new cyclin that interacts with two members of the CDC2 gene family. Cell 66(6):1217–1228
Koff A, Giordano A, Desai D, Yamashita K, Harper JW, Elledge S, Nishimoto T, Morgan DO, Franza BR, Roberts JM (1992) Formation and activation of a cyclin E-cdk2 complex during the G1 phase of the human cell cycle. Science 257(5077):1689–1694
Koundrioukoff S, Carignon S, Techer H, Letessier A, Brison O, Debatisse M (2013) Stepwise activation of the ATR signaling pathway upon increasing replication stress impacts fragile site integrity. PLoS Genet 9(7):e1003643. https://doi.org/10.1371/journal.pgen.1003643
Kumagai A, Shevchenko A, Shevchenko A, Dunphy WG (2011) Direct regulation of Treslin by cyclin-dependent kinase is essential for the onset of DNA replication. J Cell Biol 193(6):995–1007. https://doi.org/10.1083/jcb.201102003
Lacey KR, Jackson PK, Stearns T (1999) Cyclin-dependent kinase control of centrosome duplication. Proc Natl Acad Sci U S A 96(6):2817–2822
Lane AN, Fan TW (2015) Regulation of mammalian nucleotide metabolism and biosynthesis. Nucleic Acids Res 43(4):2466–2485. https://doi.org/10.1093/nar/gkv047
Lauper N, Beck AR, Cariou S, Richman L, Hofmann K, Reith W, Slingerland JM, Amati B (1998) Cyclin E2: a novel CDK2 partner in the late G1 and S phases of the mammalian cell cycle. Oncogene 17(20):2637–2643. https://doi.org/10.1038/sj.onc.1202477
Le Tallec B, Dutrillaux B, Lachages AM, Millot GA, Brison O, Debatisse M (2011) Molecular profiling of common fragile sites in human fibroblasts. Nat Struct Mol Biol 18(12):1421–1423. https://doi.org/10.1038/nsmb.2155
Le Tallec B, Millot GA, Blin ME, Brison O, Dutrillaux B, Debatisse M (2013) Common fragile site profiling in epithelial and erythroid cells reveals that most recurrent cancer deletions lie in fragile sites hosting large genes. Cell Rep 4(3):420–428. https://doi.org/10.1016/j.celrep.2013.07.003
Le Tallec B, Koundrioukoff S, Wilhelm T, Letessier A, Brison O, Debatisse M (2014) Updating the mechanisms of common fragile site instability: how to reconcile the different views? Cell Mol Life Sci 71(23):4489–4494. https://doi.org/10.1007/s00018-014-1720-2
Lee EB, Park TI, Park SH, Park JY (2003) Loss of heterozygosity on the long arm of chromosome 21 in non-small cell lung cancer. Ann Thorac Surg 75(5):1597–1600
Letessier A, Millot GA, Koundrioukoff S, Lachages AM, Vogt N, Hansen RS, Malfoy B, Brison O, Debatisse M (2011) Cell-type-specific replication initiation programs set fragility of the FRA3B fragile site. Nature 470(7332):120–123. https://doi.org/10.1038/nature09745
Lew DJ, Dulic V, Reed SI (1991) Isolation of three novel human cyclins by rescue of G1 cyclin (Cln) function in yeast. Cell 66(6):1197–1206
Liberal V, Martinsson-Ahlzen HS, Liberal J, Spruck CH, Widschwendter M, McGowan CH, Reed SI (2012) Cyclin-dependent kinase subunit (Cks) 1 or Cks2 overexpression overrides the DNA damage response barrier triggered by activated oncoproteins. Proc Natl Acad Sci U S A 109(8):2754–2759. https://doi.org/10.1073/pnas.1102434108
Liu E, Li X, Yan F, Zhao Q, Wu X (2004) Cyclin-dependent kinases phosphorylate human Cdt1 and induce its degradation. J Biol Chem 279(17):17283–17288. https://doi.org/10.1074/jbc.C300549200
Loeb KR, Kostner H, Firpo E, Norwood T, D Tsuchiya K, Clurman BE, Roberts JM (2005) A mouse model for cyclin E-dependent genetic instability and tumorigenesis. Cancer Cell 8(1):35–47. https://doi.org/10.1016/j.ccr.2005.06.010
Ma T, Van Tine BA, Wei Y, Garrett MD, Nelson D, Adams PD, Wang J, Qin J, Chow LT, Harper JW (2000) Cell cycle-regulated phosphorylation of p220NPAT by cyclin E/Cdk2 in Cajal bodies promotes histone gene transcription. Genes Dev 14(18):2298–2313
Ma Y, Fiering S, Black C, Liu X, Yuan Z, Memoli VA, Robbins DJ, Bentley HA, Tsongalis GJ, Demidenko E, Freemantle SJ, Dmitrovsky E (2007) Transgenic cyclin E triggers dysplasia and multiple pulmonary adenocarcinomas. Proc Natl Acad Sci U S A 104(10):4089–4094. https://doi.org/10.1073/pnas.0606537104
Macheret M, Halazonetis TD (2015) DNA replication stress as a hallmark of cancer. Annu Rev Pathol 10:425–448. https://doi.org/10.1146/annurev-pathol-012414-040424
Mailand N, Diffley JF (2005) CDKs promote DNA replication origin licensing in human cells by protecting Cdc6 from APC/C-dependent proteolysis. Cell 122(6):915–926. https://doi.org/10.1016/j.cell.2005.08.013
Malumbres M, Barbacid M (2009) Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer 9(3):153–166. https://doi.org/10.1038/nrc2602
Matsumoto Y, Maller JL (2004) A centrosomal localization signal in cyclin E required for Cdk2-independent S phase entry. Science 306(5697):885–888. https://doi.org/10.1126/science.1103544
Matsumoto Y, Hayashi K, Nishida E (1999) Cyclin-dependent kinase 2 (Cdk2) is required for centrosome duplication in mammalian cells. Curr Biol 9(8):429–432
Matsuura I, Denissova NG, Wang G, He D, Long J, Liu F (2004) Cyclin-dependent kinases regulate the antiproliferative function of Smads. Nature 430(6996):226–231. https://doi.org/10.1038/nature02650
Maya-Mendoza A, Ostrakova J, Kosar M, Hall A, Duskova P, Mistrik M, Merchut-Maya JM, Hodny Z, Bartkova J, Christensen C, Bartek J (2015) Myc and Ras oncogenes engage different energy metabolism programs and evoke distinct patterns of oxidative and DNA replication stress. Mol Oncol 9(3):601–616. https://doi.org/10.1016/j.molonc.2014.11.001
McIntosh D, Blow JJ (2012) Dormant origins, the licensing checkpoint, and the response to replicative stresses. Cold Spring Harb Perspect Biol 4(10):012955. https://doi.org/10.1101/cshperspect.a012955
Meraldi P, Lukas J, Fry AM, Bartek J, Nigg EA (1999) Centrosome duplication in mammalian somatic cells requires E2F and Cdk2-cyclin A. Nat Cell Biol 1(2):88–93. https://doi.org/10.1038/10054
Minella AC, Swanger J, Bryant E, Welcker M, Hwang H, Clurman BE (2002) p53 and p21 form an inducible barrier that protects cells against cyclin E-cdk2 deregulation. Curr Biol 12(21):1817–1827
Minella AC, Grim JE, Welcker M, Clurman BE (2007) p53 and SCFFbw7 cooperatively restrain cyclin E-associated genome instability. Oncogene 26(48):6948–6953. https://doi.org/10.1038/sj.onc.1210518
Minella AC, Loeb KR, Knecht A, Welcker M, Varnum-Finney BJ, Bernstein ID, Roberts JM, Clurman BE (2008) Cyclin E phosphorylation regulates cell proliferation in hematopoietic and epithelial lineages in vivo. Genes Dev 22(12):1677–1689. https://doi.org/10.1101/gad.1650208
Miron K, Golan-Lev T, Dvir R, Ben-David E, Kerem B (2015) Oncogenes create a unique landscape of fragile sites. Nat Commun 6:7094. https://doi.org/10.1038/ncomms8094
Moberg KH, Bell DW, Wahrer DC, Haber DA, Hariharan IK (2001) Archipelago regulates cyclin E levels in Drosophila and is mutated in human cancer cell lines. Nature 413(6853):311–316. https://doi.org/10.1038/35095068
Mohebi S, Mizuno K, Watson A, Carr AM, Murray JM (2015) Checkpoints are blind to replication restart and recombination intermediates that result in gross chromosomal rearrangements. Nat Commun 6:6357. https://doi.org/10.1038/ncomms7357
Morris L, Allen KE, La Thangue NB (2000) Regulation of E2F transcription by cyclin E-Cdk2 kinase mediated through p300/CBP co-activators. Nat Cell Biol 2(4):232–239. https://doi.org/10.1038/35008660
Muller-Tidow C, Metzger R, Kugler K, Diederichs S, Idos G, Thomas M, Dockhorn-Dworniczak B, Schneider PM, Koeffler HP, Berdel WE, Serve H (2001) Cyclin E is the only cyclin-dependent kinase 2-associated cyclin that predicts metastasis and survival in early stage non-small cell lung cancer. Cancer Res 61(2):647–653
Muntean AG, Hess JL (2012) The pathogenesis of mixed-lineage leukemia. Annu Rev Pathol 7:283–301. https://doi.org/10.1146/annurev-pathol-011811-132434
Mussman JG, Horn HF, Carroll PE, Okuda M, Tarapore P, Donehower LA, Fukasawa K (2000) Synergistic induction of centrosome hyperamplification by loss of p53 and cyclin E overexpression. Oncogene 19(13):1635–1646. https://doi.org/10.1038/sj.onc.1203460
Neelsen KJ, Lopes M (2015) Replication fork reversal in eukaryotes: from dead end to dynamic response. Nat Rev Mol Cell Biol 16(4):207–220. https://doi.org/10.1038/nrm3935
Neelsen KJ, Zanini IM, Herrador R, Lopes M (2013) Oncogenes induce genotoxic stress by mitotic processing of unusual replication intermediates. J Cell Biol 200(6):699–708. https://doi.org/10.1083/jcb.201212058
Nik-Zainal S, Davies H, Staaf J, Ramakrishna M, Glodzik D, Zou X, Martincorena I, Alexandrov LB, Martin S, Wedge DC, Van Loo P, Ju YS, Smid M, Brinkman AB, Morganella S, Aure MR, Lingjaerde OC, Langerod A, Ringner M, Ahn SM, Boyault S, Brock JE, Broeks A, Butler A, Desmedt C, Dirix L, Dronov S, Fatima A, Foekens JA, Gerstung M, Hooijer GK, Jang SJ, Jones DR, Kim HY, King TA, Krishnamurthy S, Lee HJ, Lee JY, Li Y, McLaren S, Menzies A, Mustonen V, O’Meara S, Pauporte I, Pivot X, Purdie CA, Raine K, Ramakrishnan K, Rodriguez-Gonzalez FG, Romieu G, Sieuwerts AM, Simpson PT, Shepherd R, Stebbings L, Stefansson OA, Teague J, Tommasi S, Treilleux I, Van den Eynden GG, Vermeulen P, Vincent-Salomon A, Yates L, Caldas C, van’t Veer L, Tutt A, Knappskog S, Tan BK, Jonkers J, Borg A, Ueno NT, Sotiriou C, Viari A, Futreal PA, Campbell PJ, Span PN, Van Laere S, Lakhani SR, Eyfjord JE, Thompson AM, Birney E, Stunnenberg HG, van de Vijver MJ, Martens JW, Borresen-Dale AL, Richardson AL, Kong G, Thomas G, Stratton MR (2016) Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534(7605):47–54. https://doi.org/10.1038/nature17676
Ohtani K, DeGregori J, Nevins JR (1995) Regulation of the cyclin E gene by transcription factor E2F1. Proc Natl Acad Sci U S A 92(26):12146–12150
Ohtani K, DeGregori J, Leone G, Herendeen DR, Kelly TJ, Nevins JR (1996) Expression of the HsOrc1 gene, a human ORC1 homolog, is regulated by cell proliferation via the E2F transcription factor. Mol Cell Biol 16(12):6977–6984
Ohtani K, Tsujimoto A, Ikeda M, Nakamura M (1998) Regulation of cell growth-dependent expression of mammalian CDC6 gene by the cell cycle transcription factor E2F. Oncogene 17(14):1777–1785. https://doi.org/10.1038/sj.onc.1202105
Ohtani K, Iwanaga R, Nakamura M, Ikeda M, Yabuta N, Tsuruga H, Nojima H (1999) Cell growth-regulated expression of mammalian MCM5 and MCM6 genes mediated by the transcription factor E2F. Oncogene 18(14):2299–2309. https://doi.org/10.1038/sj.onc.1202544
Ohtsubo M, Roberts JM (1993) Cyclin-dependent regulation of G1 in mammalian fibroblasts. Science 259(5103):1908–1912
Okuda M, Horn HF, Tarapore P, Tokuyama Y, Smulian AG, Chan PK, Knudsen ES, Hofmann IA, Snyder JD, Bove KE, Fukasawa K (2000) Nucleophosmin/B23 is a target of CDK2/cyclin E in centrosome duplication. Cell 103(1):127–140
Oswald F, Dobner T, Lipp M (1996) The E2F transcription factor activates a replication-dependent human H2A gene in early S phase of the cell cycle. Mol Cell Biol 16(5):1889–1895
Ozeri-Galai E, Lebofsky R, Rahat A, Bester AC, Bensimon A, Kerem B (2011) Failure of origin activation in response to fork stalling leads to chromosomal instability at fragile sites. Mol Cell 43(1):122–131. https://doi.org/10.1016/j.molcel.2011.05.019
Ozeri-Galai E, Tur-Sinai M, Bester AC, Kerem B (2014) Interplay between genetic and epigenetic factors governs common fragile site instability in cancer. Cell Mol Life Sci 71(23):4495–4506. https://doi.org/10.1007/s00018-014-1719-8
Pearson BE, Nasheuer HP, Wang TS (1991) Human DNA polymerase α gene: sequences controlling expression in cycling and serum-stimulated cells. Mol Cell Biol 11(4):2081–2095
Polyak K, Lee MH, Erdjument-Bromage H, Koff A, Roberts JM, Tempst P, Massague J (1994) Cloning of p27Kip1, a cyclin-dependent kinase inhibitor and a potential mediator of extracellular antimitogenic signals. Cell 78(1):59–66
Porter PL, Malone KE, Heagerty PJ, Alexander GM, Gatti LA, Firpo EJ, Daling JR, Roberts JM (1997) Expression of cell-cycle regulators p27Kip1 and cyclin E, alone and in combination, correlate with survival in young breast cancer patients. Nat Med 3(2):222–225
Rajagopalan H, Jallepalli PV, Rago C, Velculescu VE, Kinzler KW, Vogelstein B, Lengauer C (2004) Inactivation of hCDC4 can cause chromosomal instability. Nature 428(6978):77–81. https://doi.org/10.1038/nature02313
Resnitzky D, Gossen M, Bujard H, Reed SI (1994) Acceleration of the G1/S phase transition by expression of cyclins D1 and E with an inducible system. Mol Cell Biol 14(3):1669–1679
Reynaud EG, Pelpel K, Guillier M, Leibovitch MP, Leibovitch SA (1999) p57Kip2 stabilizes the MyoD protein by inhibiting cyclin E-Cdk2 kinase activity in growing myoblasts. Mol Cell Biol 19(11):7621–7629
Rohban S, Campaner S (2015) Myc induced replicative stress response: how to cope with it and exploit it. Biochim Biophys Acta 1849(5):517–524. https://doi.org/10.1016/j.bbagrm.2014.04.008
Ruffner H, Jiang W, Craig AG, Hunter T, Verma IM (1999) BRCA1 is phosphorylated at serine 1497 in vivo at a cyclin-dependent kinase 2 phosphorylation site. Mol Cell Biol 19(7):4843–4854
Schraml P, Bucher C, Bissig H, Nocito A, Haas P, Wilber K, Seelig S, Kononen J, Mihatsch MJ, Dirnhofer S, Sauter G (2003) Cyclin E overexpression and amplification in human tumours. J Pathol 200(3):375–382. https://doi.org/10.1002/path.1356
Schwaenen C, Viardot A, Berger H, Barth TF, Bentink S, Dohner H, Enz M, Feller AC, Hansmann ML, Hummel M, Kestler HA, Klapper W, Kreuz M, Lenze D, Loeffler M, Moller P, Muller-Hermelink HK, Ott G, Rosolowski M, Rosenwald A, Ruf S, Siebert R, Spang R, Stein H, Truemper L, Lichter P, Bentz M, Wessendorf S (2009) Microarray-based genomic profiling reveals novel genomic aberrations in follicular lymphoma which associate with patient survival and gene expression status. Genes Chromosom Cancer 48(1):39–54. https://doi.org/10.1002/gcc.20617
Scuderi R, Palucka KA, Pokrovskaja K, Bjorkholm M, Wiman KG, Pisa P (1996) Cyclin E overexpression in relapsed adult acute lymphoblastic leukemias of B-cell lineage. Blood 87(8):3360–3367
Shima N, Alcaraz A, Liachko I, Buske TR, Andrews CA, Munroe RJ, Hartford SA, Tye BK, Schimenti JC (2007) A viable allele of Mcm4 causes chromosome instability and mammary adenocarcinomas in mice. Nat Genet 39(1):93–98. https://doi.org/10.1038/ng1936
Silva FP, Morolli B, Storlazzi CT, Anelli L, Wessels H, Bezrookove V, Kluin-Nelemans HC, Giphart-Gassler M (2003) Identification of RUNX1/AML1 as a classical tumor suppressor gene. Oncogene 22(4):538–547. https://doi.org/10.1038/sj.onc.1206141
Siu KT, Rosner MR, Minella AC (2012) An integrated view of cyclin E function and regulation. Cell Cycle (Georgetown, Tex) 11(1):57–64. https://doi.org/10.4161/cc.11.1.18775
Siu KT, Xu Y, Swartz KL, Bhattacharyya M, Gurbuxani S, Hua Y, Minella AC (2014) Chromosome instability underlies hematopoietic stem cell dysfunction and lymphoid neoplasia associated with impaired Fbw7-mediated cyclin E regulation. Mol Cell Biol 34(17):3244–3258. https://doi.org/10.1128/mcb.01528-13
Smith AP, Henze M, Lee JA, Osborn KG, Keck JM, Tedesco D, Bortner DM, Rosenberg MP, Reed SI (2006) Deregulated cyclin E promotes p53 loss of heterozygosity and tumorigenesis in the mouse mammary gland. Oncogene 25(55):7245–7259. https://doi.org/10.1038/sj.onc.1209713
Spruck CH, Won KA, Reed SI (1999) Deregulated cyclin E induces chromosome instability. Nature 401(6750):297–300. https://doi.org/10.1038/45836
Srinivasan SV, Dominguez-Sola D, Wang LC, Hyrien O, Gautier J (2013) Cdc45 is a critical effector of myc-dependent DNA replication stress. Cell Rep 3(5):1629–1639. https://doi.org/10.1016/j.celrep.2013.04.002
Strohmaier H, Spruck CH, Kaiser P, Won KA, Sangfelt O, Reed SI (2001) Human F-box protein hCdc4 targets cyclin E for proteolysis and is mutated in a breast cancer cell line. Nature 413(6853):316–322. https://doi.org/10.1038/35095076
Teixeira LK, Wang X, Li Y, Ekholm-Reed S, Wu X, Wang P, Reed SI (2015) Cyclin E deregulation promotes loss of specific genomic regions. Curr Biol 25(10):1327–1333. https://doi.org/10.1016/j.cub.2015.03.022
Thean LF, Loi C, Ho KS, Koh PK, Eu KW, Cheah PY (2010) Genome-wide scan identifies a copy number variable region at 3q26 that regulates PPM1L in APC mutation-negative familial colorectal cancer patients. Genes Chromosom Cancer 49(2):99–106. https://doi.org/10.1002/gcc.20724
Thys RG, Lehman CE, Pierce LC, Wang YH (2015) DNA secondary structure at chromosomal fragile sites in human disease. Curr Genomics 16(1):60–70. https://doi.org/10.2174/1389202916666150114223205
Tokuyama Y, Horn HF, Kawamura K, Tarapore P, Fukasawa K (2001) Specific phosphorylation of nucleophosmin on Thr199 by cyclin-dependent kinase 2-cyclin E and its role in centrosome duplication. J Biol Chem 276(24):21529–21537. https://doi.org/10.1074/jbc.M100014200
Torres-Rosell J, De Piccoli G, Cordon-Preciado V, Farmer S, Jarmuz A, Machin F, Pasero P, Lisby M, Haber JE, Aragon L (2007) Anaphase onset before complete DNA replication with intact checkpoint responses. Science 315(5817):1411–1415. https://doi.org/10.1126/science.1134025
Toyoshima H, Hunter T (1994) p27, a novel inhibitor of G1 cyclin-Cdk protein kinase activity, is related to p21. Cell 78(1):67–74
Varetti G, Pellman D, Gordon DJ (2014) Aurea mediocritas: the importance of a balanced genome. Cold Spring Harb Perspect Biol 6(11):a015842. https://doi.org/10.1101/cshperspect.a015842
Weddington N, Stuy A, Hiratani I, Ryba T, Yokochi T, Gilbert DM (2008) Replication domain: a visualization tool and comparative database for genome-wide replication timing data. BMC Bioinformatics 9:530. https://doi.org/10.1186/1471-2105-9-530
Won KA, Reed SI (1996) Activation of cyclin E/CDK2 is coupled to site-specific autophosphorylation and ubiquitin-dependent degradation of cyclin E. EMBO J 15(16):4182–4193
Xiong Y, Hannon GJ, Zhang H, Casso D, Kobayashi R, Beach D (1993) p21 is a universal inhibitor of cyclin kinases. Nature 366(6456):701–704. https://doi.org/10.1038/366701a0
Yamada H, Yanagisawa K, Tokumaru S, Taguchi A, Nimura Y, Osada H, Nagino M, Takahashi T (2008) Detailed characterization of a homozygously deleted region corresponding to a candidate tumor suppressor locus at 21q11-21 in human lung cancer. Genes Chromosom Cancer 47(9):810–818. https://doi.org/10.1002/gcc.20582
Yan Z, DeGregori J, Shohet R, Leone G, Stillman B, Nevins JR, Williams RS (1998) Cdc6 is regulated by E2F and is essential for DNA replication in mammalian cells. Proc Natl Acad Sci U S A 95(7):3603–3608
Yoshida K, Inoue I (2004) Regulation of geminin and Cdt1 expression by E2F transcription factors. Oncogene 23(21):3802–3812. https://doi.org/10.1038/sj.onc.1207488
Zariwala M, Liu J, Xiong Y (1998) Cyclin E2, a novel human G1 cyclin and activating partner of CDK2 and CDK3, is induced by viral oncoproteins. Oncogene 17(21):2787–2798. https://doi.org/10.1038/sj.onc.1202505
Zeman MK, Cimprich KA (2014) Causes and consequences of replication stress. Nat Cell Biol 16(1):2–9. https://doi.org/10.1038/ncb2897
Zhao J, Kennedy BK, Lawrence BD, Barbie DA, Matera AG, Fletcher JA, Harlow E (2000) NPAT links cyclin E-Cdk2 to the regulation of replication-dependent histone gene transcription. Genes Dev 14(18):2283–2297
Zhao H, Chen X, Gurian-West M, Roberts JM (2012) Loss of cyclin-dependent kinase 2 (CDK2) inhibitory phosphorylation in a CDK2AF knock-in mouse causes misregulation of DNA replication and centrosome duplication. Mol Cell Biol 32(8):1421–1432. https://doi.org/10.1128/mcb.06721-11
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Teixeira, L.K., Reed, S.I. (2017). Cyclin E Deregulation and Genomic Instability. In: Masai, H., Foiani, M. (eds) DNA Replication. Advances in Experimental Medicine and Biology, vol 1042. Springer, Singapore. https://doi.org/10.1007/978-981-10-6955-0_22
Download citation
DOI: https://doi.org/10.1007/978-981-10-6955-0_22
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-6954-3
Online ISBN: 978-981-10-6955-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)