Abstract
Plant Breeding is the art of selecting and discarding genetic material to achieve crop improvement. Favourable alleles resulting in quality improvement or disease resistance must be added, while unfavourable alleles must be removed. The source for novel alleles can be other varieties, landraces or crop wild relatives. The identification of allelic variation is referred to as allele mining. Before allelic variation can be used for breeding purposes several steps need to be taken. First of all an inventory is needed of the available genetic resources. Phenotypic screens are needed to uncover potential expected and even unanticipated alleles. Next, using genetic and molecular tools, the alleles responsible for the identified traits must be traced and distinguished in order to be introgressed into new varieties.
In this review we focus on the identification of novel disease resistance traits in the agronomically important genus Solanum. The fact that R genes are present in multigene clusters within the genome, which often include many paralogs necessitates thorough discussion on the distinction between alleles and paralogs. Often such a distinction cannot easily be made. An overview is given of how natural resources can be tapped, e.g. how germplasm can be most efficiently screened. Techniques are presented by which alleles and paralogs can be distinguished in functional and/or genetic screens, including also a specific tagging of alleles and paralogs. Several examples are given in which allele and paralog mining was successfully applied. Also examples are presented as to how allele mining supported our understanding about the evolution of R gene clusters. Finally an outlook is provided how the research field of allele mining might develop in the near future.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
1.1 General Introduction
Alleles are different forms of a gene and affect a particular process in different ways. Different combinations of alleles may result in different phenotypes. Plant breeders try to improve varieties by introducing new alleles, resulting in higher yields and better quality or resistance characteristics. Identifying new, promising alleles is not an easy task. In the post-genomics era, mining of a crop’s (wild) gene pool for novel and superior alleles for agronomically important traits is becoming more and more feasible. Genebanks all over the world contain huge untapped resources of distinct alleles that may have potential application in crop breeding programs. This hidden diversity, which can consist of naturally occurring sequence variation in coding or regulatory regions of genes, can be explored by allele mining (Ramkumar et al. 2010; Varshney et al. 2005, 2009). The variation includes single nucleotide polymorphisms (SNPs) as well as insertions and deletions (InDels), which have the possibility to change the resulting phenotype. This may be by altering the amount of protein or its structure and/or function (Ramkumar et al. 2010). The recent rapid advancements in the field of genomics leads to the accumulation of enormous amounts of sequence information and fast evolving bioinformatic tools which pave the road for identifying, characterizing, isolating, and deploying previously unknown or under-utilized sources of genetic variation.
In this chapter we consider allele mining as the research field that aims at unlocking the genetic diversity existing in genetic resource collections (genebanks) and artificially created mutant populations by identifying allelic variants of genes and loci. Since resistance genes occur in clusters, where allelic relationships are often not clear (Sanchez et al. 2006; Millett et al. 2007) and because paralogs in a cluster can have different functions, the scope of this chapter is broader than allele mining alone. To deal with this we introduce the concept of paralog mining. Paralog mining is the identification of a gene within a cluster of highly homologues genes with different, often unknown, functions. Paralog mining can be used as a tool to generate molecular markers and in combination with functional screens it can be used to identify new genes conferring resistance to a particular pathogen. In this review we discuss how allele and paralog mining can help to improve disease resistance in Solanum crops.
1.2 Solanaceae Resources
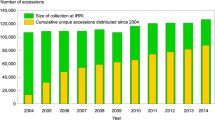
The family of Solanaceae is of high economic importance and is composed of more than 3,000 species which include important crop and model plants such as potato (Solanum tuberosum), tomato (Solanum lycopersicum) and eggplant (Solanum melongena) (Knapp 2002), but also wild species occurring in very different habitats (Spooner and Hijmans 2001; Spooner et al. 2004). About 15,000 wild potato accessions are being maintained in large collections worldwide and the establishment of core and mini collections that enable an effective use of the existing variation in gene banks while maintaining the variability, as has been proposed before (Frankel and Brown 1984; Hoekstra 2009). Allele mining requires the assembly of a reasonably sized core germplasm collection usually comprising ~ 1,000 accessions representative of genetic diversity existing in the global population (Hofinger et al. 2009). Such collections can effectively be constructed using the Focussed Identification of Germplasm Strategy (FIGS) approach (Mackay et al. 2004; Bhullar et al. 2009). About 15,000 wild potato accessions are being maintained in large collections worldwide and the establishment of core and mini collections that enable an effective use of the existing variation in gene banks while maintaining the variability, as has been proposed before (Frankel and Brown 1984; Hoekstra 2009).
The genome sequence of potato (Potato Genome Sequencing Consortium et al. 2011) and tomato (The Tomato Genome Consortium et al. 2012) will facilitate mining for novel alleles or paralogs of resistance® genes. These may be found in the largely untapped resources of crossable species within the genus Solanum allowing their exploitation in breeding programs. Also, insight into sequence diversity at the R gene loci in wild Solanum species with different resistance response against economically important diseases will result in a better understanding of the mechanism of R gene functionality and evolution but can also help to identify new alleles or paralogs with different race specificities, and develop allele-specific diagnostic markers for marker assisted breeding.
1.3 Resistance Genes
If a gene is responsible for the resistance of a particular plant to a particular pathogen, this gene is called a resistance ® gene. To date, more than 100 R genes which confer resistance to a diversity of pathogens including bacteria, fungi, oomycetes, viruses, insects and nematodes have been identified and/or cloned from various plants, by a wide variety of methods including map-based cloning, transposon tagging, and similarity based DNA library screening (Sanchez et al. 2006; Ingvardsen et al. 2008; Vleeshouwers et al. 2011a). An overview of mapped and cloned R genes from Solanaceae is given in Fig. 2.1.
Genetic locations of disease resistance traits in Solanaceae. Twelve linkage groups are shown and the position of R genes is indicated. The R genes for potato are underlined and those for other species, mainly tomato, are not underlined. Map segments having QTL for resistance to Phytophthora infestans in potato are shown in black color
R genes often encode receptors for pathogen derived ligands and they are classified based on the combination of different domains (e.g. CC = coiled coil, TIR = toll interleukin receptor, Protein Kinase, NBS = nucleotide binding site, Lec (lectin), and LRRs = leucine rich repeats). Five classes can be identified, transmembrane proteins with extracelular LRRs (receptor like proteins, RLPs), transmembrane proteins with extracellular LRRs and intracellular protein kinase (receptor like kinases, RLKs), transmembrane proteins with extracellular “lectin like” domain and intracellular protein kinase (lectin receptor kinases, LecRKs), and intracellular NBS-LRR proteins which can be divided in CC-NBS-LRR and TIR-NBS-LRR (Dubery et al. 2012). The NBS-LRR class is the most abundant and has been extensively studied (Hulbert et al. 2001). Although NBS-LRR genes are assumed to cause pathogen race specific (or vertical) resistance, it has also been suggested that members of the NBS-LRR gene family are candidates for quantitative trait loci (QTL) that are responsible for horizontal resistance (Rietman et al. 2012; Sanz et al. 2012; Gebhardt and Valkonen 2001). Most characterized plant NBS-LRR genes are physically clustered in the plant genome. The homologous sequences in such a cluster are referred to as paralogs (Gebhardt and Valkonen 2001) and paralogs can confer resistance to different isolates of the same pathogen (Dodds et al. 2001; Li et al. 2011; Lokossou 2010) or to different pathogens (Dodds et al. 2001; van der Vossen et al. 2000). Some paralogs may also be considered as molecular fossils of evolution, whose activity is unclear or even absent, e.g. many pseudogenes have been found. In most R gene clusters the number of paralogs is very high and often an allelic relationship is hard to determine (Kuang et al. 2004). However, as the genome structure between species in the Solanaceae family is highly conserved, positional conservation of R gene clusters (synteny) is observed across Solanaceous species (Grube et al. 2000; Park et al. 2009, Fig. 2.1). Therefore, even when relatively unknown genetic sources are used, it is likely that the genes conferring resistance are linked to syntenic clusters of R genes known from well-studied species like potato and tomato.
Not just the 2006; Millett et al. identification of new alleles is important, also the functional characterisation of the identified alleles is extremely important to assess the added value of the new allele over alleles that are already present in crop plants. Many approaches have already been used and especially the currently booming research field of effector genomics, through which the identification of Avr genes is greatly accelerated, offers fast functional assays to distinguish the activity of newly identified R gene alleles and paralogs. So, allele mining approaches coupled with effector profiling enable the discovery of novel R genes at an unprecedented rate (Vleeshouwers et al. 2008, 2011a).
2 Functional Resistance Screens
2.1 Screening for Disease Resistant Accessions in Gene Bank Material
Several methods are available to carry out phenotypic screens for disease resistance in gene bank collections. Here we use the evaluation of potato germplasm for late blight resistance as an example. Inoculation of entire in vitro plantlets or inoculation of detached leaves can be used as high throughput screening methods (Vleeshouwers 2011b). In case of race specific resistance, the selection of the pathogen isolates is an important issue in the identification of major R genes. If the selected isolate happens to be compatible with the R gene (s) in a particular accession, these R genes may be overlooked. Multiple isolates can be used to distinguish the different R genes in a particular resistant accession (Huang et al. 2005; Verzaux 2010). Complementary to working with the entire pathogen, effector responsiveness can be used to identify and classify R gene alleles in a germplasm core collection (Rietman et al. 2010). In such an effectoromics approach, effectors or potential Avr genes from the pathogen, are expressed in the plant using agro-infiltration or through inoculation with recombinant Potato Virus X, referred to as agro-infection. Upon recognition of the effector by the R gene expressed by the plant, a defence reaction, referred to as hypersensitive response (HR), is initiated which is visible as a necrotic lesions in the infiltrated leaf (Fig. 2.2). The agro-infiltration test appears to be applicable and reliable for many genotypes and the variation in the HR in different genetic backgrounds is limited. The use of specific pathogen isolates and the use of specific effectors can also be employed to identify functional groups of R genes in germplasm of crop wild relatives. Within groups of functionally similar R genes, a true allele mining approach can be pursued in order to identify (sequence) variation. The functional grouping of R genes can also be employed to reduce the redundancy that is inevitably present in germplasm collections. Another virtue of the effectoromics approach was shown recently. Potato plants which have shown durable resistance to late blight contained stacks of different R genes (Verzaux 2010; Kim et al. 2012). The polygenic nature of the resistances could easily be characterised using the segregation patterns of the different effector responses. Effectors which displayed HR response in germplasm screens are potential Avr gene(s) recognized by the cognate R gene. These potential R-Avr interactions should be validated by additional genetic studies. Ideally, by cosegregation of responses to the effector with resistance to P. infestans isolates in segregating populations.
Effector induced hypersensitive response (HR) in Solanum tuberosum genotype MaR8. Available effectors from Phytophthora infestans were applied using agroinfiltration in leaves of the resistant plant MaR8. The dotted circles surround the infiltrated leaf area. The red dotted circles surround the effectors that elicited an HR. These effectors are selected for validation of R-Avr interactions by additional genetic analysis
2.2 QTL mapping/LD mapping
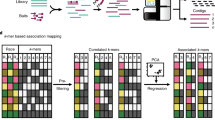
Plant pathogen resistance, at the phenotypic level, often does not behave as a single R gene but as a quantitative trait that is controlled by multiple genetic and environmental factors (Trognitz et al. 2002; Bai 2003). Understanding the molecular basis for quantitative traits will facilitate diagnosis and facilitate the combination of superior alleles in crop improvement programs. The possible approaches to mapping genes that underlie quantitative traits fall broadly into two categories: candidate gene studies, which use either association or resequencing approaches, and linkage studies, which include both QTL mapping and genome-wide association studies (GWAS). In this review, we do not discuss GWAS further because of the extensive review by Hirschhorn and Daly (2005). Linkage disequilibrium (LD) mapping, or association analysis based on candidate genes is also considered as an allele mining approach (Malosetti et al. 2007).
Some cases of close linkage between an R gene and quantitative trait loci (QTL) for pathogen resistance supports the hypothesis that qualitative and quantitative resistance have a similar molecular basis (Leonards-Schippers et al. 1994), thereby suggesting that genes showing sequence similarity to R genes are candidates for being factors underlying quantitative resistance (Rickert et al. 2003; Rietman et al. 2010). Candidate genes participating in the control of the quantitative resistance to pathogens are those involved in the disease response network; (i) R genes which recognize the pathogen and trigger the resistance response, (ii) genes which are involved in signal transduction pathways and (iii) the large group of pathogensis related (PR) genes which are expressed in response to pathogen attack and are involved in the execution phase of the defence response (reviewed by Gebhardt and Valkonen 2001).
The genetic dissection of complex plant traits in QTLs first became possible with the advent of DNA-based markers (Osborn et al. 1987). The first genes and their allelic variants underlying plant QTLs have been identified by positional cloning (reviewed in Salvi and Tuberosa 2005). Positional QTL cloning is a labor- and time-consuming process which requires the generation and analysis of large experimental mapping populations. An alternative to positional cloning of QTLs may be the allele mining approach, which is based on the knowledge of a gene’s function in controlling a characteristic of interest on the one hand, and genetic co-localization of a functional candidate gene with QTL of interest on the other (Pflieger et al. 2001; Faino et al. 2011). However, in this approach substantial a priori knowledge is required. DNA variation for genes fulfilling these criteria has been examined in natural populations of accessions related by descent for associations with positive or negative characteristic values (Li et al. 2005; Gonzalez-Martınez et al. 2007). Finding such associations indicates that DNA variation either at the candidate locus itself or at a physically linked locus is causal for the phenotypic variation, but defined prove for the involvement of the gene is still circumstantial.
3 Techniques for Allele Mining
Dependent on the research question but also dependent on genetic, genomic and financial resources available, several techniques can be used for allele mining, ranging from a rapid and inexpensive polymerase chain reaction (PCR) to next gen sequencing and everything in between. For some applications (partial) sequence information or only molecular polymorphism of the alleles is sufficient. For other applications actual cloning of the entire allele is required. Generally, all DNA based tools require the careful selection of target genes. The target gene model might require verification, and successively, a careful design of selective primers will allow the identification of novel alleles at candidate loci in the entire or core germplasm collection. In Fig. 2.3 a pipeline for novel allele discovery from germplasm collections is presented, including a combination of different approaches.
3.1 Molecular Tools for Allele Tagging
All molecular marker techniques include a PCR amplification of one or multiple alleles or paralogs. In order to identify polymorphisms between amplified alleles, single-strand specific nucleases could be applied. Using this technique, that is often used in TILLING approaches, nicking of heteroduplexes of PCR products can be easily detected. A recent development is the use of high resolution melting point analysis in order to screen for mismatches between amplified alleles in a high throughput fashion. Especially suitable for the highly polymorphic and duplicated R genes, is the NBS profiling technique (van der Linden et al. 2004). It is a powerful tool to identify specific fragments of candidate R genes or R gene homologs throughout the genome by using degenerated primers that anneal to conserved sequences in the NBS domain of the NBS-LRR class of R genes. A high throughput application of this technique is to study fragment length polymorphisms as molecular markers. Also, PCR amplification of specific R genes is possible if primers are located in unique regions in order to target the specific paralog. The results may be visible as a DNA fragment of a specific size on an agarose gel. However, when gene specific markers are used in different germplasm material, often a-specific annealing of the primers can occur and therefore it will always be necessary to sequence the resulting PCR fragments to confirm their identity and homogeniety.
3.1.1 NBS Profiling
Many plant R genes are a member of a multigene cluster composed of multiple copies with high sequence similarity (Song et al. 2003). The NBS region of (NBS-LRR) R genes and their analogs (RGAs) contain highly conserved common motifs like the P-loop, the kinase −2 motif and the GLPL motif (Meyers et al. 2003; Monosi et al. 2004). These conserved motifs within the NBS-LRR genes have been used successfully to sequence (parts of) NBS regions from various plant species (Collins et al. 1998; Pflieger et al. 1999; Zhang and Gassmann 2007). NBS profiling uses the conserved motifs for efficient tagging of NBS-LRR type of R genes and their analogs (Van der Linden et al. 2004, 2005). The technique involves three different steps. (1) Digestion of genomic DNA with a restriction enzyme and ligation of adaptors to compatible restriction ends. (2) PCR amplification of NBS containing fragments using an NBS primer and an adaptor primer. (3) Separation of amplified fragments by polyacrylamide gel electrophoresis. The technique produces a multilocus profile of the genome.
NBS profiling can easily be adapted to target other conserved gene families, which is referred to as motif-directed profiling (Van der Linden et al. 2004, 2005). Also NBS profiling can be adapted to target particular R gene clusters. R genes from the same cluster usually have similarities in their sequences not shared with other R genes (McDowell and Simon 2006; Meyers et al. 2005), allowing the design of specific primers for a particular R gene cluster. NBS profiling could therefore also be adapted to reach high fragment saturation in an R gene cluster of interest (Verzaux et al. 2011, 2012; Jo et al. 2011). This technique is referred to as cluster directed profiling.
3.1.2 (Eco)-tilling
Eco tilling is a molecular method to screen germplasm core and mini collections. This technique is distinct from the TILLING approach since TILLING screens identify novel alleles that are induced by mutagenesis (Till et al. 2003; Barkley and Wang 2008) whereas eco-tilling identifies naturally occurring alleles in germplasm (Barone et al. 2009). Both approaches employ a similar screening method to identify variation in alleles. Polymorphisms in PCR amplified DNA fragments are detected in hetroduplexes of the amplicons using single strand specific nucleases, high resolution melting point analysis or deep sequencing in next generation sequencing.
3.1.3 Amplification of Specific Allelic Variants
Family members with very similar sequences may have dispersed around the genome into non synthenous loci or may have remained within a genetic locus but has multiplicated resulting in tandem or inverted repeats. In general, sequences in coding regions will be more conserved than primers in flanking sequences. Dependent of the downstream application (sequence comparison, in plant expression), primers are chosen in- or outside coding sequence to amplify the entire gene, or part of the gene or only the open reading frame. Because even single nucleotide polymorphisms can be relevant differences between alleles, preferably the DNA polymerase will contain proofreading activity. Also, because often long stretches of the target gene are amplified, a long range polymerase chain reaction (LR-PCR) polymerase is preferred. Examples of enzymes that harbour both characteristics are Pfu-Turbo from Invitrogen or Phusion from Fermentas. One approach is to amplify the entire coding sequence of the R gene of interest using primers annealing to start and stop codon regions. Subsequently, the amplicon is sequenced and for expression studies it can be cloned in a vector that harbours heterologous regulatory sequences. For some accessions a possible lack of amplification can be expected due to absence of a coding gene or to low sequence homology at the primer annealing sites. A drawback of this approach is that the promotor and terminator regions of the novel alleles are missing, so variation in these regulatory regions are neglected. For ‘true’ allele mining, the use of primers matching the promotor and terminator regions is feasible when sequence conservation is sufficient. Accessions may also first be screened for the presence of the known R gene with a diagnostic molecular marker obtained from haplotype studies at the R gene locus and next for the presence of new alleles of the known R gene (Bhullar et al. 2009) to identify stronger alleles. Song et al. (2003) showed that allele mining could be used to clone the functional RB allele from a cluster with two highly similar paralogs. Also Wang et al. (2008) and Lokossou et al. (2010) could specifically amplify the target allele rather than paralogous genes in a Rpi-blb1 allele mining study. Latha et al. (2004) exploited allele mining to identify stress tolerance genes in Oryza species and related germplasm. A common feature of the three genes investigated was that they were members of multigene families. Primers based on the 5’ and 3ʹ untranslated region of genes were found to be sufficiently conserved over the entire range of germplasm in rice to which the concept of allelism is applicable, while the primers based on the start and stop codon amplified sequences from additional loci (Latha et al. 2004).
is the cloning of the Rpi-vnt1.1 gene (Pel et al. 2009). NBS profiling revealed a fragment that was co-segregating with resistance in a F1 population. The sequence of this NBS profiling band was similar to a known R gene (Tm-2 2). The mined allele had a different genetic position on the same chromosome as the Tm-2 2 gene. The entire coding sequence of the Rpi-vnt1.1 allele was found after sequence analysis of a BAC clone derived from the genomic locus (Foster et al. 2009).
3.2 Next Generation Sequencing in Allele Mining
Currently, most genome and transcriptome sequencing projects, which used Sanger sequencing methodology in the past, are being replaced by next generation sequencing (NGS) technologies. These NGS technologies are able to generate data inexpensively and at a rate that is several orders of magnitude faster than that of traditional technologies (reviewed in Ercolano et al. 2012). At present there are several next generation sequencers on the market (Voelkerding et al. 2009). Most of these systems have different underlying biochemistries but all of these technologies sequence populations of PCR-amplified DNA molecules. The Heliscope and the PacBio, which sequence single molecules, are the exceptions. The amount of sequence data and the length of the reads are increasing with the continued development of the technology. Now resequencing and de novo sequencing of transcriptomes and genomes is becoming more and more accessible for individual labs (Varshney et al. 2009). This will lead to the discovery of novel useful variation which has been limiting the application of sequence-based selection in plants in the pre NGS era (Henry 2011). The availability of large numbers of genetic markers that can facilitate linkage mapping and whole genome scanning (WGS)-based association genetics that are of practical use for MAS in marker-deficient crops (Varshney et al. 2009). Resequencing of several genomes (Cao et al. 2011) followed by the comparison of all candidate R genes is now feasible in Arabidopsis (Guo et al. 2011). Soon this type of analysis will also be applied for crops and their wild relatives. Resequencing of parts of the genome with duplicated sequences, like R gene clusters, will remain a challenge, especially in heterozygous species like potato (Potato Genome Sequencing Consortium 2012). Single molecule sequencing will offer great opportunities for this research field (Koren et al. 2012).
3.3 Functional Analysis of Newly Identified Alleles
If the candidate genes have been identified in allele mining studies, it is required to confirm their functionality. Transient and stable transformations are valid for that purpose. Agroinfiltration, an Agrobacterium tumefaciens-based method, is currently the best developed and most reliable method for transient expression in plants (Vleeshouwers et al. 2011). Using this method R gene alleles and the Avr genes can be coexpressed in N. benthamiana (Bos et al. 2006) or other plant species (Rietman et al. 2010). Consequently, a hypersensitive response (HR) occurs in the infiltrated leaf area. This approach is only applicable in cases where the cognate Avr gene is available. Agroinfiltration can also be carried out by expressing only the R gene in a host plant, followed by pathogen challenge inoculations (Lokossou et al. 2009; Pel et al. 2009). Another type of transient expression, which allows high throughput screenings is agroinfection. A gene of interest is cloned into a viral genome. Successively, the viral genome is introduced into plant cells through A. tumefaciens. Only a few cells need to be infected after which viral particles are formed that spread through the plant. Along with the virus, the gene of interest is expressed in the plant. There is, however, a limitation to the size of the gene to be expressed. Fragments over 500 bp in size will not express sufficiently.
The stable transformation into plant is still considered functional analysis which provides the most clear and definitive evidence. Transgenic plants can be tested for resistance in different developmental stages. For example, an in vitro inoculation assay was developed for routine high-throughput disease testing of Phytophthora infestans in potato (Huang et al. 2005).
4 Examples of Allele Mining in Solanum
As described in the previous section the technique of choice is highly dependent on the research question and application. Applications can be very diverse, ranging from very practical, like R gene mapping and cloning, to the identification of novel genetic resources, to more scientific applications like R gene geographic distribution and evolution. In this section examples of applications using the different techniques are presented.
4.1 Genetic Mapping
Mapping of R genes is strongly facilitated by allele mining through NBS profiling. Typical examples of R gene mapping approach are provided by Pel et al. (2009), Jacobs et al. (2010), Jo et al. (2011) and Verzaux et al. (2011, 2012). The first step in the approach consists of producing small (n = 20–100) populations segregating for P. infestans resistance, phenotyping the populations for resistance, and composing bulks of resistant and susceptible individuals. Then, the bulks are genotyped using NBS profiling to obtain markers that co-segregate with resistance. Next sequencing of co-segregating NBS fragments and BLAST analysis to identify the fragments is performed. Combining this information with literature and genome sequence data on mapping of resistance genes will suggest a putative map position. Finally, the map positions are confirmed using known flanking markers.
Jo et al. (2011) used NBS profiling and successive marker sequence comparison to the potato and tomato genome draft sequences to identify the genetic position of the late blight resistance gene R8. According to this work, R8 was located on the long arm of chromosome IX and not on the short arm of chromosome XI as was suggested previously by Huang et al. (2005). This is a first example where NBS markers could be directly landed in the sequenced (draft) genomes of potato and tomato. Through comparison of known markers in the tomato genetic map to the draft sequence, scaffolds were anchored to the tomato genetic map (anchored scaffold approach). Very recently, the R9 mediated-late blight resistance was also mapped near the R8 locus on chromosome IX using R gene cluster directed profiling approaches (Jo et al. in preparation).
4.2 Cloning Functional Alleles
Several late blight R genes have been cloned from potato wild relatives using allele- and paralog mining (for reviews see: Vleeshouwers et al. 2011, Rietman et al. 2010). Sometimes there is no clear distinction between allele- and paralog mining because of the high similarity among genes. An example of true allele mining was shown by Vleeshouwers et al. (2008) who isolated the functional alleles of Rpi-blb1 present in S. stoloniferum, Rpi-sto1 and Rpi-pta1. The entire genes were isolated by long range PCR using primers up and downstream of the coding regions. Specificity of the cloned genes was shown with different P. infestans isolates and with effector IpiO −1 and 2, which is recognized by Rpi-blb1, Rpi-sto1 and Rpi-pta1. An allelic relationship between the three genes was also shown using marker (CT88) segregation studies (Wang et al. 2008). Sequence analyses showed that the putative functional homologs Rpi-sto1 and Rpi-pta1 are nearly identical to Rpi-blb1, with only 3 and 5 non-synonymous nucleotide substitutions inside the coding sequence, respectively.
A slightly different example of allele mining was provided by Lokossou et al. (2009), who described the map based cloning and functional characterization of Rpi-blb3 and Rpi-abpt, which are allelic variants R2 and R2-like. An allele mining strategy was employed using a start stopcodon approach. In this study a major technological improvement was made. The GatewayTM technology was used to clone the entire amplified coding sequences in a destination vector under the control of the Rpi-blb3 promotor and terminator. A combination of efficient cloning of candidate alleles was combined with transient complementation assays in Nicotiana benthamiana and allowed for the rapid cloning and identification of R2 and R2-like alleles.
Champouret used a similar technical approach to mine for R3a and R2 alleles. A start stopcodon approach was pursued and the R3a screen revealed alleles with identical activity in distantly related species. This is considered as true allele mining. Also the R2 screen revealed many genes with identical activity, however, also a few genes were identified which had slightly different recognition specificities, suggesting that not only alleles but also paralogs were mined. This is an example where allele mining and paralog mining are overlapping (Champouret 2010).
Paralog mining strategies can be pursued in order to facilitate map based cloning of novel R gene variants. An example of successful paralog mining came available using an R2 mining approach applied on S microdontum. This resulted in the isolation of Rpi-mcd1 which is functionally distinct from R2 since the Avr2 gene was not recognised (Lokossou 2010). Another example is the cloning of the Rpi-vnt1.1 gene (Pel et al. 2009). NBS profiling revealed a fragment that was co-segregating with resistance in a F1 population. The sequence of this NBS profiling band was similar to a known R gene (Tm-2 2). PCR amplification of Tm-2 2 homologs identified the functional Rpi-vnt1.1 gene. The mined allele had a different genetic position on the same chromosome as the Tm-2 2 gene. Also the biological activity was different and therefore this study followed a typical paralog mining approach. This study also illustrates a risk associated with paralog mining in multigene families (Pel et al. 2009). Using the start stop codon primer pair derived from Tm-2 2 only a part of the coding sequence was identified and a N-terminal extension, specific for the Rpi-vnt1 alleles were overlooked. The entire coding sequence of the Rpi-vnt1.1 allele was found after sequence analysis of a BAC clone derived from the genomic locus (Foster et al. 2009).
4.3 Uncovering Allelic Variation for Specific Genes
Allele mining can also be used to uncover genetic variation for a particular R gene and identify germplasm containing functional alleles from the same or different species. Nunziata et al. (2007) studied the variability of one cluster of genes at the Gro1 locus responsible for resistance to Globodera rostochiensis race Ro1 in several potato species. The cluster is known to comprise 10 different paralogs, among which only the Gro1–4 gene has been demonstrated to confer resistance against Globodera rostochiensis race Ro1. Using available sequence information, three primer pairs were designed that target different regions of the Gro1 sequence. The first was designed in a highly conserved region and allowed the presence of at least one member of the gene cluster to be identified in 16 wild species analysed. The second primer pair was designed on a Gro1–4 specific region and its use demonstrated that no gene identical to Gro1–4 was present in any wild potato species analysed. Finally, the major part of the LRR coding sequence of the Gro1 gene was amplified and sequenced in 16 wild species. In total, 409 SNPs were identified, varying between species from 12 SNPs in S. demissum to 35 in S. stoloniferum. These data could be used to identify evolutionary selection pressure since the non-synonymous/synonymous ratio (Ka/Ks) in most species was different from 1.
A similar type of screen was performed by Wang et al. (2008) and Lokossou et al. (2010). They analyzed the presence and allelic diversity of the late blight R genes Rpi-blb1, Rpi-blb2 and Rpi-blb3 in 196 different taxa of tuber-bearing Solanum species. The Rpi-blb1 gene is part of a resistance gene analog (RGA) cluster of four members on chromosome VIII, Rpi-blb2 resides in a locus harbouring at least 15 tomato Mi gene homologs on chromosome VI and Rpi-blb3 originates from a cluster on chromosome IV. For all genes primers were design that would allow amplification of a specific fragment of the gene. The genes were only present in some Mexican diploid as well as polyploid species closely related to S. bulbocastanum, although differences in the distribution existed among the 3 genes. The Rpi-blb1 gene was only found in S. bulbocastanum, S.cardiophyllum subsp. cardiophyllum, and S. stoloniferum, the Rpi-blb2 only in S. bulbocastanum, and the Rpi-blb3 gene in S. pinnatisectum, S. bulbocastanum (including some subspecies), S. hjertingii, S. nayaritense, S. brachistotrichum, and S. stoloniferum. Sequence analysis of part of the Rpi-blb1 and Rpi-blb3 gene suggests an evolution through recombination and point mutations. For Rpi-blb2 only sequences identical to the cloned gene were found, suggesting that it has emerged recently. The three R genes occurred in different combinations and frequencies in S. bulbocastanum accessions and their spread is confined to Central America (Lokossou et al. 2010). A practical outcome of the allele mining study by Wang et al. (2008) was the discovery of conserved homologues of Rpi-blb1 in an EBN 2 tetraploid potato species, e.g. S. stoloniferum. The Rpi-blb1 is present in the diploid tuber-bearing S. bulbocastanum, which is not directly crossable with the tetraploid S. tuberosum. Solanum stoloniferum can be directly crossed to cultivated potato, thus facilitating an easy transfer of a gene with exactly the same specificity and functionality as Rpi-blb1.
An allele mining approach to identify variation in the Avr9 recognizing Cf-9 alleles provided evidence for the presumed evolutionary mechanism driving R gene diversification. Subsequent intra- and intergenic unequal recombination events were held responsible for the sequence diversification of Cf-9 alleles. However, this diversification was not accompanied by a functional diversification since the Avr9 effector could still be recognized (Van der Hoorn et al. 2001; Kruijt et al. 2004).
4.4 Alleles in Natural Populations of Solanum
Knowledge on the evolution and distribution of disease resistance genes is important for a better understanding of the dynamics of these genes in nature. Caicedo (2008) studied geographic diversity cline of R gene homologs in natural populations of Solanum pimpinellifolium L., a wild relative of cultivated tomato, to determine the possible roles of demography and selection on R gene evolution. The patterns of diversity at the multigenic Cf-2 gene family were investigated which consisted of 26 closely related homologs, referred to as the Hcr2-p family (Caicedo and Schaal 2004). The 26 Hcr2-p homologs display length variation due primarily to variation in the number of LRR-coding units within each gene and can be classified into nine different size classes according to length; within size-classes, homologs differ from each other by one or a few single nucleotide polymorphisms (SNPs). Solanum pimpinellifolium individuals vary extensively in the number of Hcr2-p homologs they carry, with Southern blots results suggesting 1–5 genes per individual (Caicedo and Schaal 2004). Species-wide analyses of Hcr2-p sequence diversity suggest that selection has played a role in the evolution of the gene family. Patterns of amino acid substitution are consistent with purifying selection in the 5′ LRR-coding portion of the genes and positive selection on some amino acid residues in the 3′ region. Evolutionary relationships among homologs also suggest that balancing selection has shaped species-wide patterns of diversity. Studies on patterns of diversity at the multigenic Cf-2 gene family in S. pimpinellifolium populations along the northern coast of Peru showed that population diversity levels of Cf-2 homologs follow a latitudinal cline, consistent with the species’ history of gradual colonization of the Peruvian coast and population variation in outcrossing.
In another approach the wild tomato germplasm was screened for responsiveness to the Avr4 and Avr9 effectors from C. fulvum, which are recognized by the Cf-4 and Cf-9 R proteins respectively. Recognition and the presence of the matching R genes was ubiquitous throughout the screened germplasm. This allele mining approach showed that C. fulvum is an ancient pathogen of the genus Lycopersicon (Kruijt et al. 2005).
Several studies have now clearly shown the potential of allele mining in Solanum for the improvement of disease resistance. A large number of allelic variants of known disease resistance genes have been discovered and in several cases also functionality of the variants was shown. As was shown, allele mining can strongly facilitate the cloning of R genes by using comparative genomics approaches. Allele mining has also been shown to be useful for the identification of orthologous sequences in species that are more easily crossable with the cultivated material then the species the gene was originally discovered in, thus facilitating a more rapid deployment of genes in breeding programs. Furthermore allele mining was useful for the identification of novel, yet unknown R genes and shed light on evolutionary processes related to these genes. As more and more R genes are identified and cloned, the chances increase that new R genes reside at known and well-characterized loci, enabling the use of comparative genomics and, thus, the development of efficient allele mining strategies.
The availability of the potato and tomato genome sequences, together with a constant drop in the sequencing cost will boost allele mining even more. The fast (r)evolution in the high throughput sequencing technologies, especially the increase in read lengths expected from the third generation of single molecule sequencing platforms, will provide a complete survey of the distribution of R gene clusters in the Solanaceae family, enabling a dramatic acceleration in the process of identifying agronomically important genes like novel R genes.
We envisage that novel and efficient ‘mining’ strategies can give direct access to disease resistance genes of interest using next generation sequencing in combination with effector genomics. However, as not all effectors are known yet more effort should be made in that area. Another interesting research area relates to the durability of the R genes. At present it is unknown whether all allelic variants discovered for a particular gene are equally easily overcome by the pathogen. If not, this may be a way to identify more durable genes.
References
Bai Y, Huang CC, van der Hulst R et al (2003) QTLs for tomato powdery mildew resistance (Oidium lycopersici) in Lycopersicon parviflorum G1.1601 co-localize with two qualitative powdery mildew resistance genes. Mol Plant Microbe Interact 16:169–176
Barkley NA, Wang ML (2008) Application of TILLING and EcoTILLING as reverse genetic approaches to elucidate the function of genes in plants and animals. Curr Genomics 9:212–226
Barone A, Di Matteo A, Carputo D, Frusciante L (2009) High-throughput genomics enhances tomato breeding efficiency. Curr Genomics 10:1–9
Bhullar NK, Street K, Mackay M et al (2009) Unlocking wheat genetic resources for the molecular identification of previously undescribed functional alleles at the Pm3 resistance locus. Proc Natl Acad Sci U S A 106:9519–9524
Bos JIB, Kanneganti TD, Young C et al (2006) The C-terminal half of Phytophthora infestans RXLR effector AVR3a is sufficient to trigger R3a-mediated hypersensitivity and suppress INF1-induced cell death in Nicotiana benthamiana. Plant J 48:165–176
Caicedo AL (2008) Geographic diversity cline of R gene homologs in natural populations of Solanum pimpinellifolium (Solanaceae). Am J Bot 95:393–398
Caicedo AL, Schaal BA (2004) Heterogeneous evolutionary processes affect R gene diversity in natural populations of Solanum pimpinellifolium. Proc Natl Acad Sci U S A 101:17444–17449
Cao J, Schneeberger K, Ossowski S et al (2011) Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat Genet 43:956–963
Champouret (2010) Functional genomics of Phytophthora infestans effectors and Solanum resistance genes. PhD thesis, Wageningen University
Dodds PN, Lawrence GJ, Ellis JG (2001) Six amino acid changes confined to the leucine-rich repeat β-strand/β-turn motif determine the difference between the P and P2 rust resistance specificities in flax. Plant Cell 13:495–506
Dubery IA, Sanabria NM, Huang JC (2012) Self and nonself. Advances in experimental medicine biology 738:79–107. doi:10.1007/978-1-4614-1680-7_6
Ercolano MR, Sanseverino W, Carli P et al (2012) Genetic and genomic approaches for R-gene mediated disease resistance in tomato: retrospects and prospects. Plant Cell Rep 31:973–985
Faino L, Azizinia S, Hassanzadeh BH et al (2012) Fine mapping of two major QTLs conferring resistance to powdery mildew in tomato. Euphytica 184:223–234
Foster SJ, Park TH, Pel MA et al (2009) Rpi-vnt1.1, a Tm-2 homolog from Solanum venturii confers resistance to potato late blight. Mol Plant Microbe Interact 22:589–600
Frankel OH, Brown ADH (1984) Plant genetic resources today: a critical appraisal. In: Holden JHW, Williams JH (eds) Crop genetic resources: conservation & evaluation. G. Allen, London, pp. 249–257
Gebhardt C, Valkonen JPT (2001) Organization of genes controlling disease resistance in the potato genome. Annu Rev Phytopathol 39:79–102
Grube RC, Radwanski ER, Jahn M (2000) Comparative genetics of disease resistance within the Solanaceae. Genetics 155:873–887
Guo YL, Fitz J, Schneeberger K et al (2011) Genome-wide comparison of nucleotide-binding site-leucine-rich repeat-encoding genes in Arabidopsis. Plant Physiol 157:757–769
Gupta PK (2008) Single-molecule DNA sequencing technologies for future genomics research. Trends Biotechnol 26:602–611
Henry RJ (2011) Next-generation sequencing for understanding and accelerating crop domestication. Brief Funct Genomics [Epub. PMID:22025450]
Hoekstra R (2009) Exploring the natural biodiversity of potato late blight resistance. Potato Res 52:237–244
Hofinger BJ, Jing HC, Kim EHK, Kanyuka K (2009) High-resolution melting analysis of cDNA-derived PCR amplicons for rapid and cost-effective identification of novel alleles in barley. Theor Appl Genet 119:851–865
Huang S (2005) Discovery and characterization of the major late blight resistance complex in potato. PhD thesis, Wageningen University, Wageningen
Hulbert SH, Webb CA, Smith SM, Sun Q (2001) Resistance gene complexes: evolution and utilization. Annu Rev Phytopathol 9:285–312
Imelfort M, Batley J, Grimmond S, Edwards D (2009) Genome sequencing approaches and successes. In: Somers DJ et al (eds) Methods in molecular biology, plant genomics, vol 513. Humana Press, pp. 345–358
Jacobs MMJ, Vosman B, Vleeshouwers VGAA et al (2010) A novel approach to locate Phytophthora infestans resistance genes on the potato genetic map. Theor Appl Genet 120:785–796
Jo KR, Arens M, Kim TY et al (2011) Mapping of the S. demissum late blight resistance gene R8 to a new locus on chromosome IX. Theor Appl Genet 123:1331–1340
Kaur N, Street K, Mackay M et al (2008) Molecular approaches for characterization and use of natural disease resistance in wheat. Eur J Plant Pathol 121:387–397
Kim HJ, Lee HR, Jo KR et al (2012) Broad spectrum late blight resistance in potato differential set plants MaR8 and MaR9 is conferred by multiple stacked R genes. Theor Appl Genet 124:5
Knapp S (2002) Tobacco to tomatoes: a phylogenetic perspective on fruit diversity in the Solanaceae. J Exp Bot 53:2001–2022
Koren S, Schatz MC, Walenz BP et al (2012) Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat Biotechnol 30:693–700
Kuang H, Woo SS, Meyers BC et al (2004) Multiple genetic processes result in heterogeneous rates of evolution within the major cluster disease resistance genes in lettuce. Plant Cell 16:2870–2894
Kruijt M, Brandwagt BF, de Wit PJ (2004) Rearrangements in the Cf-9 disease resistance gene cluster of wild tomato have resulted in three genes that mediate Avr9 responsiveness. Genetics 168:1655–1663
Kruijt M, Kip DJ, Joosten MH et al (2005) The Cf-4 and Cf-9 resistance genes against Cladosporium fulvum are conserved in wild tomato species. Mol Plant Microbe Interact 18:1011–1021
Latha R, Rubia L, Bennett J, Swaminathan MS (2004) Allele mining for stress tolerance genes in Oryza species and related germplasm. Mol Biotech 27:101–108
Li G, Huang S, Guo X et al (2011) Cloning and characterization of R3b; members of the R3 superfamily of late blight resistance genes show sequence and functional divergence. Mol Plant Microbe Interact 24:1132–1142
Lokossou (2010) Dissection of the major late blight resistance cluster on potato linkage group IV. PhD thesis, Wageningen University
Lokossou AA, Park TH, van Arkel G et al (2009) Exploiting knowledge of R/Avr genes to rapidly clone a new LZ-NBS-LRR family of late blight resistance genes from potato linkage group IV. Mol Plant Microbe Interact 22:630–641
Lokossou AA, Rietman H, Wang M et al (2010) Diversity, distribution and evolution of Solanum bulbocastanum late blight resistance genes. Mol Plant Microbe Interact 23:1206–1216
McDowell JM, Simon SA (2006) Recent insights into R gene evolution. Mol Plant Pathol 7:437–448
Meyers BC, Kaushik S, Nandety RS (2005) Evolving disease resistance genes. Curr Opin Plant Biol 8:129–134
Meyers BC, Kozik A, Griego A et al (2003) Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 15:809–834
Millett BP, Bradeen JM (2007) Development of allele-specific PCR and RT-PCR assays for clustered resistance genes using a potato late blight resistance transgene as a model. Theor Appl Genet 114:501–513
Nunziata A, Ruggieri V, Frusciante L, Barone A (2007) Allele mining at the locus Gro 1 in Solanum wild species. In: VI International Solanaceae Conference 449–456
Park TH, Vleeshouwers VGAA, Jacobsen E et al (2009) Molecular breeding for resistance to Phytophthora infestans (Mont.) de Bary in potato (Solanum tuberosum L.): a perspective of cisgenesis. Plant Breed 128:109–117
Pel MA, Foster SJ, Park TH et al (2009) Mapping and cloning of late blight resistance genes from Solanum venturii using an interspecific candidate gene approach. Mol Plant Microbe Interact 22:601–615
Pflieger S, Lefebvre V, Caranta C et al (1999) Disease resistance gene analogs as candidates for QTLs involved in pepper/pathogen interactions. Genome 42:1100–1110
Pflieger S, Palloix A, Caranta C et al (2001) Defense response genes co-localize with quantitative disease resistance loci in pepper. Theor Appl Genet 103:920–929
Potato Genome Sequencing Consortium, Xu X, Pan S, Cheng S et al (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Ramkumar G, Sakthivel K, Sundaram RM et al (2010) Allele mining in crops: prospects and potentials. Biotechnol Adv 28:451–461
Rietman H, Bijsterbosch, Liliana MC et al (2012) Qualitative and quantitative late blight resistance in the potato cultivar Sarpo Mira is determined by the perception of five distinct RXLR effectors. Mol Plant Microb Interact 25:910–919
Rietman H, Champouret N, Hein I et al (2010) Plants and oomycetes, an intimate relationship: co-evolutionary principles and impact on agricultural practice. Hemming D (ed), CAB Reviews, CABI, UK. June 2011 5:1–17
Sanchez MJ, Bradeen JM (2006) Towards efficient isolation of R gene orthologs from multiple genotypes: optimization of long range-PCR. Mol Breed 17:137–148
Sanseverino W, Roma G, De Simone M et al (2010) PRGdb: a bioinformatics platform for plant resistance gene analysis. Nucl Acids Res 38:D814–D821
Sanz MJ, Loarce Y, Fominaya A et al (2012) Identification of RFLP and NBS/PK profiling markers for disease resistance loci in genetic maps of oats. Theor Appl Genet. doi:10.1007/s00122-012-1974-8
Sato S, Tabata S, Mueller LA et al (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Schneeberger K, Weigel D (2011) Fast-forward genetics enabled by new sequencing technologies. Trends Plant Sci 6:282–288
Song J, Bradeen JM, Naess SK et al (2003) Gene RB cloned from Solanum bulbocastanum confers broad spectrum resistance to potato late blight. Proc Natl Acad Sci U S A 100:9128–9133
Spooner DM, Hijmans RJ (2001) Potato systematics and germplasm collecting, 1989–2000. Amer J Potato Res 78:237–268
Spooner DM, Peralta IE, Knapp S (2005) Comparison of AFLPs with other markers for phylogenetic inference in wild tomatoes [Solanum L. section Lycopersicon (Mill.) Wettst.]. Taxon 54:43–61
Till BJ, Colbert T, Tompa et al (2003) High-throughput TILLING for functional genomics. In: Grotewold E (ed) plant functional genomics: methods and protocols. Methods in molecular biology, vol. 236. Human Press, Totowa, NJ, pp 205–220
Trognitz F, Manosalva P, Gysin R et al (2002) Plant defense genes associated with quantitative resistance to potato late blight in Solanum phureja × dihaploid S. tuberosum hybrids. Mol Plant Microbe Interact 15:587–597
Upadhyaya HD, Gowda CLL, Buhariwalla HK, Crouch JH (2006) Efficient use of crop germplasm resources: identifying useful germplasm for crop improvement through core and mini-core collections and molecular marker approaches. Plant Genetic Resour: Charac Util 4:25–35
Van derHRA, Kruijt M, Roth R et al (2001) Intragenic recombination generated two distinct Cf genes that mediate AVR9 recognition in the natural population of Lycopersicon pimpinellifolium. Proc Natl Acad Sci U S A 98:10493–10498
van der LCG, Wouters D, Mihalka V et al (2004) Efficient targeting of plant disease resistance loci using NBS profiling. Theor Appl Genet 109:384–393
van der Vossen EAG, van der Voort JNAMR, Kanyuka K et al (2000) Homologues of a single resistance-gene cluster in potato confer resistance to distinct pathogens: a virus and a nematode. Plant J 23:567–576
Varshney RK, Andreas GA, Sorrells ME (2005) Genomics assisted breeding for crop improvement. Trends Plant Sci 10:621–630
Verzaux E, Budding D, de Vetten N et al (2011) High resolution mapping of a novel late blight resistance gene Rpi-avl1, from the wild Bolivian species Solanum avilesii. Am J Potato Res 88:511–519
Verzaux E, van Arkel G, Vleeshouwers VGAA et al (2012) High-resolution mapping of two broad-spectrum late blight resistance genes from two wild species of the Solanum circaeifolium group. Potato Res 55:109–123
Vleeshouwers VGAA, Raffaele S, Vossen JH et al (2011a) Understanding and exploiting late blight resistance in the age of effectors. Annu Rev Phytopathol 49:507–531
Vleeshouwers VGAA, Finkers R, Budding D et al (2011b) SolRgene: an online database to explore disease resistance genes in tuber-bearing Solanum species. BMC Plant Biol 11:116
Vleeshouwers VGAA, Rietman H, Krenek P et al (2008) Effector genomics accelerates discovery and functional profiling of potato disease resistance and Phytophthora infestans avirulence genes. PLoS ONE 3:e2875
Voelkerding KV, Dames SA, Durtschi JD (2009) Next-generation sequencing: from basic research to diagnostics. Clin Chem 55:641–658
Wang M, Allefs S, van den Berg R et al (2008) Allele mining in Solanum: conserved homologues of Rpi-blb1 are identified in Solanum stoloniferum. Theor Appl Genet 116:933–943
Zhang XC, Gassmann W (2007) Alternative splicing and mRNA levels of the disease resistance gene RPS4 are induced during defense responses. Plant Phys 145:1577–1587
Acknowledgements
JV was supported by the DuRPh program funded by the Ministry of Agriculture in the Netherlands (now Ministry of EL&I). KRJ was financially supported by the international program BO −10–010-112 program of the Ministry of EL&I and the EuropeAid program 128275/C/ACT/KP2 project DCI-FOOD/2009/218–671.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Vossen, J., Jo, KR., Vosman, B. (2014). Mining the Genus Solanum for Increasing Disease Resistance. In: Tuberosa, R., Graner, A., Frison, E. (eds) Genomics of Plant Genetic Resources. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-7575-6_2
Download citation
DOI: https://doi.org/10.1007/978-94-007-7575-6_2
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-7574-9
Online ISBN: 978-94-007-7575-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)