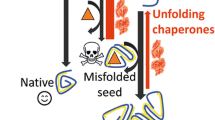
Abstract
Heavy metal ions such as Cd2+, Hg2+ and Pb2+ as well as metalloid arsenic(III) species very efficiently inhibit the refolding of chemically denatured proteins (IC50 values in nanomolar range). In their presence, the proteins misfold and aggregate. Denatured proteins appear to be much more susceptible to form high-affinity pluridentate complexes with heavy metals and metalloids than native proteins. In a denatured protein, the potential ligands of metal ions, the most important ones being cysteine and histidine residues, are more easily accessible for the toxic agents; moreover, denatured proteins with more flexible and motile backbones are more likely than folded native proteins to tolerate the formation of pluridentate protein–metal complexes with their defined geometry. In cells, the interference of metals with nascent and other non-native forms of proteins might manifest itself both in a quantitative deficiency of the affected proteins and the formation of proteotoxic aggregates. Possibly, the toxic effects of heavy metals and metalloids arise not only from their interaction with specific, particularly susceptible native proteins but also from a general derailing of protein folding . The toxic scope of heavy metals and metalloids thus could be more pleiotropic and extensive than assumed so far.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
Certain heavy metal ions and metalloids, e.g. iron, copper, manganese or zinc, act as cofactors of many proteins, enzymes in particular, and are essential components of living matter. However, these essential components in overdose as well as xenobiotic, i.e. non-essential, heavy metal ions and metalloids have proven to cause acute and chronic toxicoses in all forms of life (Hu 2005; Kosnett 2007), including carcinogenic effects (Waisberg et al. 2003) and prenatal and developmental defects (Bolin et al. 2006; Monnet-Tschudi et al. 2006; Wu et al. 2008). Environmental and occupational exposure of humans, in particular to cadmium , mercury , lead and arsenic , may entail severe health hazards. The very existence of this volume on Cellular effects of heavy metals testifies to the importance of heavy metal toxicity as a topic in medicine and public health. The interaction of heavy metal ions with living matter has also been medically exploited: organoarsenicals and mercury compounds were once used against syphilis and trypanosomiasis; calomel (Hg2Cl2) served as diuretic, laxative and antiseptic; sublimate (HgCl2) and organic mercury compounds as antiseptic in the treatment of wounds and as conserving agent in certain vaccines. While these applications are now considered obsolete, other metal-containing compounds are still in medical use: bismuth subgallate is employed as internal deodorant and in cosmetic formulations, and a platinum complex (cisplatin) serves as a well established cytostatic agent. Recently, arsenic trioxide has been approved as a cytostatic against acute promyelocytic leukemia (Wang and Chen 2008).
While the general toxicity of heavy metals and arsenic is undisputed and their remarkably pleiotropic toxic effects are known in detail, the underlying molecular mechanisms are mostly unclear. General consensus holds that proteins are the prime targets of heavy metal ions and arsenicals; only few metals such as chromium, nickel and platinum are known to interact directly with DNA. Proteins have as yet been considered to be affected by metal ions in two different ways: the toxic metal ions either bind to free thiol and other functional groups of certain native proteins, or replace essential metal ions in metal-dependent proteins (Gurd and Wilcox 1956; Vallee and Ulmer 1972; Kägi and Hapke 1984; Fraústo da Silva and Williams 1993). Here, we review a third mode of how heavy metals and metalloids may interact with and impair cellular proteins. Folding proteins have proven much more susceptible to heavy metal ions and arsenicals than proteins that have reached their native state (Sharma et al. 2008; Ramadan et al. 2009). Heavy metal ions (at nanomolar concentration) and arsenic(III) compounds (at micromolar concentrations) have been found to inhibit the refolding of chemically denatured proteins. Conceivably, nascent proteins and other forms of non-native proteins in cells are affected by heavy metals and metalloids in the same way.
Principles of Protein Folding
The amino acid sequence of a protein determines its unique three-dimensional structure, which corresponds to the energetically most favorable spatial arrangement of the polypeptide chain (Anfinsen 1973). The refolding of chemically denatured proteins is initiated by abolishing the denaturing conditions, e.g. by dilution of the denaturing agent; thus, the total polypeptide chain simultaneously takes part in the refolding process. The intracellular folding of nascent proteins, however, appears to be co-translational, i.e. to start already in the first synthesized, amino-terminal segment before the synthesis of the polypeptide chain has been completed. Moreover, because of molecular crowding, the folding of many proteins in cells is assisted by molecular chaperones , specialized proteins that improve the yield of folding and, in many cases, driven by ATP hydrolysis, rescue proteins, which are misfolded because of heat and other types of cellular stress. All major classes of molecular chaperones comprise heat-inducible members, their expression being markedly enhanced at elevated temperature and other conditions of cellular stress (for reviews, see Georgopoulos and Welch 1993; Sharma et al. 2009). Despite the apparently more complex mechanisms of intracellular protein folding as compared to in vitro protein folding , the general principle that a folding polypeptide chain spontaneously seeks to attain the conformation of lowest free energy still holds true.
Interaction of Heavy Metals with Functional Groups of Proteins
Heavy metal ions form monodentate and pluridentate complexes with S, N, O atoms in proteins. The most important ligands are the thiol groups of cysteine residues and the imidazole groups of histidine residues because they produce the most stable complexes (Table 12.1). The values of the dissociation equilibrium constants K¢d given in Table 12.1 are those for monodentate complexes; pluridentate (multidentate) complexes, in which the metal ion coordinates with more than one ligand in the same protein, are much more stable, their K¢d values roughly equating to the product of the K¢d values of the monodentate complexes of the individual ligands. Metalloproteins form such highly stable pluridentate complexes with their essential metal ions, most complexes having tetrahedral (with four ligands) or octahedral (six ligands) geometry (Gurd and Wilcox 1956; Vallee and Ulmer 1972; Kägi and Hapke 1984). Engagement of a folding protein in highly stable pluridentate protein–metal complexes has recently been found to interfere gravely with the formation of the native protein structure (Sharma et al. 2008; Ramadan et al. 2009). The reasons for folding proteins being more susceptible to heavy metals than native proteins seem obvious: the side chains in unfolded proteins are not only more exposed to the solvent but also more flexible and motile than in native folded proteins and thus more prone to be incorporated as ligands in pluridentate metal complexes.
Interference of Heavy Metals with the Refolding of Chemically Denatured Proteins
Cd2+, Hg2+ and Pb2+ at nanomolar concentrations have been found to inhibit the spontaneous refolding of chemically denatured luciferase (Fig. 12.1). The metal ions affect the refolding of the protein without apparent delay. In contrast, the native protein is much less affected by the metal ions under the same conditions, being inactivated only to a limited degree in a slow time-dependent process (Fig. 12.2).
Inhibition of luciferase refolding by Cd2+, Hg2+ and Pb2+. Luciferase (17.5 mM) was chemically denatured in 6 M guanidine hydrochloride, 50 mM Tris acetate, 5 mM TCEP (Tris[2-carboxyethyl]phosphine, a non-thiol reducing agent), pH 7.5, for 30 min at 25°C. Spontaneous, unassisted refolding at 25°C was initiated through 1:50 dilution (final concentration of luciferase 350 nM) with refolding buffer (50 mM Tris acetate, 100 mM potassium perchlorate, 15 mM magnesium acetate, pH 7.5), containing the indicated concentrations of Cd2+ a Hg2+ b and Pb2+ c Luciferase activity was measured in samples of the refolding solution at the indicated times (for details, see Sharma et al. 2008). Error bars represent the SEM from three independent experiments
Effect of metal ions on the enzymic activity of native luciferase. The effect of Cd2+, Hg2+ and Pb2+ (100 nM) on the enzymic activity of native luciferase (350 nM) was tested in refolding buffer at 25°C under the same conditions as used for the refolding of chemically denatured luciferase (Fig. 12.1)
The dose–response curves for refolding inhibition (Fig. 12.3a) reveal IC50 values in the two-digit nanomolar range (Table 12.2), whereas the dose–response curves for inactivation of native luciferase show less than 50% inhibition even at the highest concentration (500 nM) of Cd2+ and Pb2+ (Fig. 12.3b).
Dose–response curves of Cd2+, Hg2+ and Pb2+ for inhibition of refolding and inactivation of native enzyme. a Inhibition of spontaneous refolding of luciferase by Cd2+, Hg2+ and Pb2+. Chemically denatured luciferase (350 nM) was refolded at 25°C in the presence of increasing concentrations of metal ions. Luciferase activity was measured as a function of time, and the final yield of activity after 120 min (expressed as percentage of the yield in the absence of metal ion) plotted vs metal concentration. b Inactivation of native luciferase by Cd2+, Hg2+ and Pb2+. Luciferase (350 nM) was incubated for 120 min at 25°C with the indicated concentrations of metal ions. The residual activity after 120 min is plotted
It is important to note that the concentration of luciferase was 350 nM (20 mg/ml) in all experiments; at lower concentrations, the reproducibility of the measurements had proven unsatisfactory. Therefore, the IC50 values had to be determined at metal ion concentrations considerably lower than the 350 nM concentration of the target protein and thus cannot serve for quantitatively estimating the stability of the protein–metal complexes underlying the folding inhibition. The IC50 values measured under these conditions (Table 12.2) perforce underestimate the folding-inhibitory effect of the metal ions, particularly that of Hg2+.
Reduced glutathione and the chelating agent EDTA attenuate the inhibitory effect of Cd2+. However, neither agent rescues protein that has become misfolded in the presence of cadmium ions (Sharma et al. 2008). The ATP-dependent Hsp70 molecular chaperone system (DnaK/DnaJ/GrpE of Escherichia coli) significantly reduces the refolding inhibitory effect of Cd2+ (Table 12.2). The cyclic action of this chaperone system includes the following steps: ATP-DnaK with fast binding and release kinetics binds the substrate, i.e. the non-native protein; DnaJ stimulates the hydrolysis of DnaK-bound ATP, thus converting ATP-DnaK to ADP-DnaK with slow kinetics and high affinity for the substrate (Palleros et al. 1993; Schmid et al. 1994); pulling action of tightly bound DnaK disentangles the misfolded substrate (De Los Rios et al. 2006; Sharma et al. 2009); GrpE exchanges DnaK-bound ADP with ATP, thus triggering the release of the substrate or its re-entry into the chaperone cycle (Siegenthaler and Christen 2006). Per cycle, one DnaK molecule hydrolyzes one ATP molecule, the rate of ATP hydrolysis thus corresponds to the rate of the chaperone cycle.
Measurement of ATP consumption clearly demonstrates an acceleration of the chaperone cycle, i.e. an increased engagement of the chaperone system, due to the metal-induced misfolding of luciferase (Fig. 12.4). The steady-state ATPase activity of DnaK/DnaJ/GrpE in the absence of denatured luciferase is relatively slow and not affected by Cd2+. Denatured luciferase increases the ATPase activity through cis-activation of the DnaK-ATPase by DnaJ in ternary (ATP-DnaK)-luciferase-DnaJ complexes (Han and Christen 2003). The additional presence of Cd2+ increases the ATPase activity even more. Apparently, the perturbation of luciferase refolding by the metal ion almost doubles the chaperone load (Fig. 12.4). The increased chaperone load is to be attributed to a higher incidence of misfolded polypeptide chains. An enhanced expression of cellular heat shock proteins, in particular of Hsp70, is indeed observed in cells exposed to heavy metal ions (Wagner et al. 1999; Han et al. 2007; Kusakabe et al. 2008; for reviews, see Hall 2002; Ahsan et al. 2009).
In addition to luciferase, we have tested with three other proteins whether their refolding was inhibited by metal ions (Table 12.2). Cysteine-containing lactate dehydrogenase proved as susceptible as cysteine-containing luciferase, while cysteine-containing malate dehydrogenase and cysteine-less glucose-6-phosphate dehydrogenase from Leuconostoc mesenteroides were somewhat less affected.
Mechanism of Folding Inhibition by Heavy Metal Ions
The efficiency of folding inhibition as expressed by the reciprocal of the IC50 values was Hg2+ > Cd2+ > Pb2+ with all four proteins that have as yet been were tested (Table 12.2). This order correlates with the relative stability of the monodentate complexes of these metal ions with thiol, imidazole and carboxylate groups in proteins (Table 12.1). However, the IC50 values of Cd2+ and Pb2+ (very tight binding Hg2+, probably due to gross underestimating of its IC50 value as mentioned above, is an exception) are much lower than the dissociation equilibrium constants of the monodentate complexes. We infer from this discrepancy that the refolding-inhibitory protein–metal complexes are pluridentate rather than monodentate complexes, the metal ions being bound to several appropriately positioned liganding side chains of the denatured protein molecule. The possibility of metal ions interacting with two to six ligands and forming pluridentate complexes with their metal-specific geometry (Gurd and Wilcox 1956; Vallee and Ulmer 1972; Kägi and Hapke 1984; Fraústo da Silva and Williams 1993) is of course much higher in a denatured protein with its more flexible and motile polypeptide chain. The example of glucose-6-phosphate dehydrogenase (Table 12.2) shows that such chelate-like structures are even formed in proteins that are devoid of cysteine residues and apparently form stable pluridentate complexes exclusively with the more weakly binding imidazole and carboxylate ligands.
Xenobiotic heavy metals other than cadmium , mercury or lead as well as overdosed essential heavy metals are of course also to be expected to perturb protein folding . Depending on the affected protein and the type of metal ion or metalloid, differential effects on the kinetics and thermodynamics of the folding trajectory of the protein will ensue.
Interference of As(III) Species with Oxidative Refolding of Disulfide Bond-Containing Proteins
Results very similar to those obtained with heavy metal ions have been reported by Ramadan et al. (2009) with three different arsenic(III)compounds, such as arsenite (arsenous acid, As(OH)3) and monomethylarsenous acid (CH3As(OH)2) as inhibitors of oxidative protein refolding. Three different disulfide-bonded extracellular proteins were tested: lysozyme and ribonuclease A, each with four disulfide bridges, and riboflavin-binding protein with nine disulfide bridges. Low micromolar concentrations of the arsenicals efficiently inhibited the oxidative refolding of the chemically denatured proteins. The arsenicals bind rapidly and tightly to the cysteine residues of the reduced denatured proteins, three and two thiol groups coordinating with one molecule of arsenite and monomethylarsenous acid, respectively. Reduced glutathione (5 mM) weakens the inhibitory effect, which, however, still prevails. The interactions of the arsenic(III) compounds with the reduced proteins are complex and not amenable to quantitative analysis, a tentative estimate by the authors of this review suggests IC50 values in the one-digit micromolar range.
In comparison with heavy metal ions, arsenicals thus seem somewhat less efficient in disturbing protein folding and their mode of inhibition, i.e. preventing the oxidative formation of structurally indispensable disulfide bonds, is different from that of heavy metal ions, which form pluridentate complexes comprising also protein side chains other than thiols. Despite these differences, the consequences of heavy metal ions and arsenicals interfering with protein refolding are in fact very similar: in both cases, aggregates of inactive misfolded proteins are produced with an increased propensity for binding thioflavin-T (Sharma et al. 2008; Ramadan et al. 2009), indicative of β-structured protein aggregates (LeVine 1999). Similar to heavy metals , arsenic induces the expression of heat-shock genes (Johnston et al. 1980; Levinson et al. 1980) and causes an accumulation of ubiquitinated cellular proteins (Kirkpatrick et al. 2003; Stanhill et al. 2006).
Possible Sequels of Protein Folding Inhibition in Cells
The results reviewed here indicate that the toxic scope of heavy metals and metalloids like arsenic might be greater than assumed as yet. Both groups of toxic agents might interact not only with specific native proteins that are particularly susceptible, but also, at least in principle, with any protein in non-native state. All nascent polypeptide chains are at least transitorily potential targets, and any proteins in other non-native states, e.g. proteins under heat or other cellular stress as well as natively unfolded (intrinsically unstructured) proteins (for a review, see Fink 2005), might also be affected. Denatured and other non-native proteins are indeed well known to be much more susceptible to proteolytic attack and to chemical modification, because the cleavable bonds and functional groups that are buried in the native protein become exposed upon denaturation (Means and Feeney 1971).
The susceptibility of proteins to folding inhibition by heavy metal ions or arsenicals may be assumed to depend on various structural features: first, on the number of cysteine and histidine residues and the distribution of such residues along the polypeptide chain, which determines their accessibility and the steric feasibility of forming pluridentate complexes; and second, the relative rates of complex formation and of attaining the native structure of the protein. In the completely folded protein most potentially liganding groups will be buried; moreover, the formation of pluridentate complexes with their specific geometry would require at least partial unfolding of the protein. Importantly, formation of protein–metal complexes , in contrast to proteolysis or chemical modification, is extremely fast, the rate constants of divalent metal ions for substitution of inner-sphere water of aquo ions being in the range of 107–109 s−1 (Fraústo da Silva and Williams 1993). The fast rate of complex formation may explain that even the folding of a fast-folder protein like ribonucleaseA is impaired by arsenic(III) species (Ramadan et al. 2009).
In the case of heavy metal and metalloid poisoning, the derailing of protein folding in general might not only lead to a loss of function, i.e. to a quantitative shortage of the affected proteins, but might manifest itself in the formation of toxic protein aggregates . Under these circumstances, cellular protein homeostasis might become imbalanced as observed in folding diseases (Chiti and Dobson 2006; Gidalevitz et al. 2006) and possibly in aging (Cohen et al. 2006).
Conclusions
We have clear-cut experimental evidence that heavy metals and metalloids very efficiently interfere with in-vitro protein refolding, and there is irrefutable evidence that these agents are highly toxic to all forms of life. However, there is a missing link between the in-vitro observation on the molecular level and the vast toxicological data stock. The missing link is experimental evidence that heavy metals and metalloids interfere with protein folding and induce formation of toxic protein aggregates not only in vitro but also in cells. There is correlative, but not cogent, evidence for this cause-and-effect relationship. Cells exposed to heavy metals and arsenicals invariably respond with an induction of heat shock proteins and an accumulation of ubiquitinated proteins (Johnston et al. 1980; Levinson et al. 1980; Wagner et al. 1999; Kirkpatrick et al. 2003; Othumpangat et al. 2005; Stanhill et al. 2006; Han et al. 2007; Kusakabe et al. 2008; for reviews, see Hall 2002; Ahsan et al. 2009).
The biological defense mechanisms against the sequels of heavy metal poisoning indeed are, in the order of their employment, reduced glutathione, the intracellular concentration of which being 5 mM or higher (Bánhegyi et al. 2007); ubiquitous metal-binding metallothioneins (Kägi and Schäffer 1988; Klaassen et al. 2009) and, additionally, in plants the enzymically synthesized phytochelatins (Freisinger 2008); the cellular chaperone network, in particular Hsp70 and Hsp60; and finally the gated proteases. If all these lines of defense fail, the deposition and compaction by aggresomes in less toxic inclusions, which may be degraded by lysosomal autophagy, provide a last resort (for reviews, see Hinault et al. 2006; Sharma et al. 2009). The in-vivo Unfolded Protein Response to heavy metal or metalloid poisoning might thus relate to the in-vitro observations that the refolding of proteins in the presence of a heavy metal ion results in an increased chaperone load (Fig. 12.4) and that folding inhibition by both heavy metals and arsenicals results in an accumulation of thioflavinT-binding aggregates (Sharma et al. 2008; Ramadan et al. 2009).
Future experimental efforts should focus on in-vivo experiments aimed at assessing the extent of the interference of heavy metals and metalloids with intracellular non-native proteins. The perturbation of the folding of cellular proteins in general, if existing, could contribute to explaining the pleiotropic, yet metal-specific, symptomatology of heavy metal poisoning (Waisberg et al. 2003; Hu 2005; Kosnett 2007). This mode of toxic action might not only be important in the pathogenesis of classic heavy metal poisoning, but also underlie so far unknown or inexplicable consequences of exposure of living organisms to heavy metals , including certain protein folding diseases (Barnham et al. 2004; Chiti and Dobson 2006; Wu et al. 2008), autoimmune responses (Rowley and Monestier 2005), and subtle chronic impairments of health that are still undefined (Hu 2005; Cohen et al. 2006).
References
Ahsan N, Renaut J, Komatsu S (2009) Recent developments in the application of proteomics to the analysis of plant responses to heavy metals. Proteomics 9:2602–2621
Anfinsen CB (1973) Principles that govern the folding of protein chains. Science 181:223–230
Bánhegyi G, Benedetti A, Csala M, Mandl J (2007) Stress on redox. FEBS Lett 581:3634–3640
Barnham KJ, Masters CL, Bush AI (2004) Neurodegenerative diseases and oxidative stress. Nat Rev Drug Discov 3:205–214
Bolin CM, Basha R, Cox D, Zawia NH, Maloney B, Lahiri DK, Cardozo-Pelaez F (2006) Exposure to lead and the developmental origin of oxidative DNA damage in the aging brain. FASEB J 20:788–790
Chiti F, Dobson CM (2006) Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem 75:333–366
Cohen E, Bieschke J, Perciavalle RM, Kelly JW, Dillin A (2006) Opposing activities protect against age-onset proteotoxicity. Science 313:1604–1610
De Los Rios P, Ben-Zvi A, Slutsky O, Azem A, Goloubinoff P (2006) Hsp70 chaperones accelerate protein translocation and the unfolding of stable protein aggregates by entropic pulling. Proc Natl Acad Sci U S A 103:6166–6171
Fink AL (2005) Natively unfolded proteins. Curr Opin Struct Biol 15:35–41
Fraústo da Silva JJR, Williams RJP (1993) The biological chemistry of the elements: the inorganic chemistry of life. Clarendon Press, Oxford
Freisinger E (2008) Plant MTs-long neglected members of the metallothionein superfamily. Dalton Trans 47:6663–6675
Georgopoulos C, Welch WJ (1993) Role of the major heat shock proteins as molecular chaperones. Annu Rev Cell Biol 9:601–634
Gidalevitz T, Ben-Zvi A, Ho KH, Brignull HR, Morimoto RI (2006) Progressive disruption of cellular protein folding in models of polyglutamine diseases. Science 311:1471–1474
Gurd FR, Wilcox PE (1956) Complex formation between metallic cations and proteins, peptides and amino acids. Adv Protein Chem 11:311–427
Hall JL (2002) Cellular mechanisms for heavy metal detoxification and tolerance. J Exp Bot 53:1–11
Han SG, Castranova V, Vallyathan V (2007) Comparative cytotoxicity of cadmium and mercury in a human bronchial epithelial cell line (BEAS-2B) and its role in oxidative stress and induction of heat shock protein 70. J Toxicol Environ Health A 70:852–860
Han W, Christen P (2003) Mechanism of the targeting action of DnaJ in the DnaK molecular chaperone system. J Biol Chem 278:19038–19043
Hinault MP, Ben-Zvi A, Goloubinoff P (2006) Chaperones and proteases: cellular fold-controlling factors of proteins in neurodegenerative diseases and aging. J Mol Neurosci 30:249–265
Hu H (2005) Heavy metal poisoning. In: Kasper DL et al (eds) Harrison’s principles of internal medicine, 16th edn. McGraw-Hill, New York, pp 2577–2580
Johnston D, Oppermann H, Jackson J, Levinson W (1980) Induction of four proteins in chick embryo cells by sodium arsenite. J Biol Chem 255:6975–6980
Kägi JHR, Hapke H-J (1984) Biochemical interactions of mercury, cadmium and lead. In: Nriagu JO (ed) Changing metal cycles and human health. Springer, Berlin, pp 237–250
Kägi JHR, Schäffer A (1988) Biochemistry of metallothionein. Biochemistry 27:8509–8515
Kirkpatrick DS, Dale KV, Catania JM, Gandolfi AJ (2003) Low-level arsenite causes accumulation of ubiquitinated proteins in rabbit renal cortical slices and HEK293 cells. Toxicol Appl Pharmacol 186:101–109
Klaassen CD, Liu J, Diwan BA (2009) Metallothionein protection of cadmium toxicity. Toxicol Appl Pharmacol 238:215–220
Kosnett MJ (2007) Heavy metal intoxication and chelators. In: Katzung BG (ed) Basic and clinical pharmacology, 10th edn. McGraw-Hill, New York, pp 945–957
Kusakabe T, Nakajima K, Nakazato K, Suzuki K, Takada H, Satoh T, Oikawa M, Arakawa K, Nagamine T (2008) Changes of heavy metal, metallothionein and heat shock proteins in Sertoli cells induced by cadmium exposure. Toxicol In Vitro 22:1469–1475
LeVine H 3rd (1999) Quantification of beta-sheet amyloid fibril structures with thioflavin T. Methods Enzymol 309:274–284
Levinson W, Oppermann H, Jackson J (1980) Transition series metals and sulfhydryl reagents induce the synthesis of four proteins in eukaryotic cells. Biochim Biophys Acta 606:170–180
Means GE, Feeney RE (1971) Chemical modification of proteins. Holden-Day, San Francisco
Monnet-Tschudi F, Zurich MG, Boschat C, Corbaz A, Honegger P (2006) Involvement of environmental mercury and lead in the etiology of neurodegenerative diseases. Rev Environ Health 21:105–117
Othumpangat S, Kashon M, Joseph P (2005) Eukaryotic translation initiation factor 4E is a cellular target for toxicity and death due to exposure to cadmium chloride. J Biol Chem 280:25162–25169
Palleros DR, Reid KL, Shi L, Welch WJ, Fink AL (1993) ATP-induced protein-Hsp70 complex dissociation requires K+ but not ATP hydrolysis. Nature 365:664–666
Ramadan D, Rancy PC, Nagarkar RP, Schneider JP, Thorpe C (2009) Arsenic(III) species inhibit oxidative protein folding in vitro. Biochemistry 48:424–432
Rowley B, Monestier M (2005) Mechanisms of heavy metal-induced autoimmunity. Mol Immunol 42:833–838
Schmid D, Baici A, Gehring H, Christen P (1994) Kinetics of molecular chaperone action. Science 263:971–973
Sharma SK, Goloubinoff P, Christen P (2008) Heavy metal ions are potent inhibitors of protein folding. Biochem Biophys Res Commun 372:341–345
Sharma SK, Christen P, Goloubinoff P (2009) Disaggregating chaperones: an unfolding story. Curr Protein Pept Sci 10:432–446
Siegenthaler RK, Christen P (2006) Tuning of DnaK chaperone action by nonnative protein sensor DnaJ and thermosensor GrpE. J Biol Chem 281:34448–34456
Stanhill A, Haynes CM, Zhang Y, Min G, Steele MC, Kalinina J, Martinez E, Pickart CM, Kong XP, Ron D (2006) An arsenite-inducible 19S regulatory particle-associated protein adapts proteasomes to proteotoxicity. Mol Cell 23:875–885
Vallee BL, Ulmer DD (1972) Biochemical effects of mercury, cadmium, and lead. Annu Rev Biochem 41:91–128
Wagner M, Hermanns I, Bittinger F, Kirkpatrick CJ (1999) Induction of stress proteins in human endothelial cells by heavy metal ions and heat shock. Am J Physiol 277:L1026–1033
Waisberg M, Joseph P, Hale B, Beyersmann D (2003) Molecular and cellular mechanisms of cadmium carcinogenesis. Toxicology 192:95–117
Wang ZY, Chen Z (2008) Acute promyelocytic leukemia: from highly fatal to highly curable. Blood 111:2505–2515
Wu J, Basha MR, Brock B, Cox DP, Cardozo-Pelaez F, McPherson CA, Harry J, Rice DC, Maloney B, Chen D, Lahiri DK, Zawia NH (2008) Alzheimer’s disease (AD)-like pathology in aged monkeys after infantile exposure to environmental metal lead (Pb): evidence for a developmental origin and environmental link for AD. J Neurosci 28:3–9
Acknowledgements
This work was supported in part by a grant from the Swiss National Science Foundation (3100A0-109290) to PG. We thank J.H.R. Kägi for valuable discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media B.V.
About this chapter
Cite this chapter
Sharma, S.K., Goloubinoff, P., Christen, P. (2011). Non-native Proteins as Newly-Identified Targets of Heavy Metals and Metalloids. In: Banfalvi, G. (eds) Cellular Effects of Heavy Metals. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-0428-2_12
Download citation
DOI: https://doi.org/10.1007/978-94-007-0428-2_12
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-0427-5
Online ISBN: 978-94-007-0428-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)