Abstract
Genomics and phenomics are two fundamentally important branches of biological sciences, and they stand at both ends of the multiple “omics” families. A central goal of current biology is to establish complete functional links between the genome and phenome, the so-called genotype–phenotype map. Recent advances in high-throughput and high-dimensional genotyping and phenotyping technologies enable us to uncover the casual networks inside the “black box” that lies between genotypes and phenotypes using the principles of genome-wide association studies (GWAS). Application of GWAS and analogous methodologies and incorporation of multiple omics data begin to unravel the contribution of genetic variation to phenotypic diversity. Integrating “omics” data at broad levels by using the systems-biology approach is paramount to further bridging the gaps between genomics and phenomics and eventually making accurate predictions of phenotypes based on genetic contribution.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- High-throughput phenotyping
- Next-generation sequencing (NGS)
- Genotype–phenotype map (G-P map)
- Phenomics
- Genome-wide association study (GWAS)
- Quantitative trait loci (QTL)
1 Introduction
With the rapid advances of high-throughput resequencing and marker genotyping, high-density genetic variation information (such as single-nucleotide polymorphisms, SNPs, and copy-number variants, CNVs) has been collected and need to be linked with functions. Over the past few years, a multitude of genome-wide association studies (GWAS) and related strategies have identified numerous genetic variants associated with complex diseases or other traits in humans and plants, providing valuable insights into their genetic architecture. These findings are definitely enriching our knowledge about the genetic basis of phenotypic variation and provide an opportunity for genetic testing. However, most variants identified so far explain only a small proportion of the causal genetic factors, leaving the remaining “missing” heritability to be explained [1]. Moreover, even with a complete understanding of the genetics of a complex phenotypic trait, it is still challenging to accurately predict phenotypic variation from individual genetic codes. Furthermore, the majority of these disease- or trait-related variants lie within noncoding regions of genomes, complicating their functional evaluation and offering the greatest challenge in the “post-GWAS” era [2].
Globally linking genetic variants to phenotypic diversity is one of the key goals of biology. Our understanding of such a genotype–phenotype map cannot be established without detailed phenotypic data [3]. However, our ability to characterise phenomes – the full set of phenotypes of an individual – largely lags behind our ability to characterise genomes. Hence, phenomics – high-throughput and high-dimensional phenotyping – is emerging as a suit of new technologies to accelerate progress in our understanding of the relationship between genotype and phenotype [3, 4].
In this chapter, we will first review the principle of dissecting genotypes and monitoring phenotypes, usually in high-throughput manners. We also highlight current approaches to obtaining phenomic data and the emerging applications of large-scale phenotyping approaches in the phenomics era. We then outline the current strategies, such as GWAS and analogous methodologies, for globally linking genetic variation to phenotypic diversity. We summarise insights about the complete “genotype–phenotype” map that could be established through integrating “omics” data at broad levels in terms of a systems-biology approach. Related phenome projects and phenomic tools are discussed. Please keep in mind that the results discussed here are mostly based on research in humans and/or plants and that only a subset of published information can be mentioned.
2 Defining the Genotype and Phenotype
In this section, we outline the state-of-the-art methods used for the assessment of genotypes and phenotypes and the corresponding mapping approaches for linking genotypes to phenotypes at global levels (Table 11.1). We also present phenomics-related projects that combine rich genomic data with data on quantitative variation in phenotypes and which have recently been launched in both humans and plants (Table 11.2). We highlight many emerging technologies developed for high-throughput phenotyping in plants (Table 11.3).
2.1 Genetic Variation: Genotyping
Genotyping technology is referred to as the set of methodologies and protocols used to elucidate the genetic makeup (genotype) of an individual, also known as genotypic assaying. Genotyping is essential in deciphering the genetic causes of complex phenomena, including health, disease, crop yields and evolutionary fitness. Human genetic mapping was initially performed based on restriction fragment length polymorphisms (RFLPs) [5, 6], amplified fragment length polymorphisms (AFLPs) [7] and microsatellite markers (also known as short tandem repeats or simple sequence repeats) [8]. More recently, SNPs, due to their high abundance, low mutation rates and amenability to high-throughput analysis, have become the markers of choice for linkage and linkage disequilibrium (LD) mapping [9, 10]. The usually binary SNP markers are well suited to automated, high-throughput typing. Indeed, it is now feasible to genotype SNPs with high density at the genome-wide scale by utilising array-based [11, 12] or sequencing-based [13, 14] technologies (Table 11.1). Although high-throughput SNP arrays avoid time-consuming cloning and primer design steps, they lack of the discovery process and show bias towards genotyping new populations. Now, with the advent of next-generation sequencing (NGS), new technologies such as reduced-representation libraries (RRLs) [15] or complexity reduction of polymorphic sequences (CRoPS) [16], restriction-site-associated DNA sequencing (RAD-seq) [17] and low-coverage genotyping, including multiplexed shotgun genotyping (MSG) [18] or genotyping by sequencing (GBS) [19], are capable of genome-wide marker discovery for both model organisms and non-model species. Although sequence-level variants have been catalogued more extensively, structural variations – including indels (insertions/deletions), CNVs and inversions – are now investigated for their contribution to complex traits, including many important common diseases [20]. CNVs can be identified with various genome analysis platforms, including array-based comparative genomic hybridisation (CGH), SNP genotyping platforms and NGS.
Our knowledge regarding human genetic variations is mostly derived from the international effort of the SNP Consortium [21] and the International HapMap Project [22] (Table 11.2). Recent advances in sequencing technology make it possible to comprehensively catalogue genetic variation in population samples. Projects such as the Personal Genome Project (PGP) (e.g. diploid personal genomes [23]), the 1000 Genomes Project (TGP) [24] and exome sequencing projects [25] are under way in an attempt to elucidate the full spectrum of human genetic variations as a foundation to investigate the relationship between genotype and phenotype. For example, the Phase 1 publication of TGP in 2012 included whole-genome sequences of 1,092 individuals from 14 populations. A total of 38 million SNPs, 1.4 million short indels and more than 14,000 larger deletions were identified [26]. Notably, the genome of any apparently healthy individual carries more than 2,500 nonsynonymous variants at conserved regions, 20–40 variants identified as damaging at conserved sites and ~150 loss-of-function (LoF) variants in protein-coding genes, some of which are known to cause Mendelian disease [26].
Meanwhile, genome-wide genotyping is extensively performed in plants in recent years (Table 11.2), such as in Arabidopsis thaliana [27], rice [28], maize [29, 30], sorghum [31] and barley [32]. These rich resources will ultimately help to explore the genetic basis of plant agriculture-related traits, such as flowering time, growth rate, yield and stress tolerance, and to improve crops and understand plant adaptation.
2.2 Phenomics: Multilevel and Multidimensional Assessment of Features
The term phenotype includes the composite of an organism’s observable traits or characteristics – such as its morphological, developmental, physiological, pathological or biochemical properties, phenology and behaviour – that can be monitored, quantified and/or visualised by some technical procedure. Phenomics is defined as the study of all the phenotypes of an organism (phenome) that are the result of genetic code (G), environmental factors (E) and their interactions (G × E). In contrast to genotypes, which are essentially single one-dimensional as merely determined by the linear DNA code, phenotypes are usually multi-dimensional and are frequently capricious in different spatial and temporal situations. An important field of research today is trying to improve, both qualitatively and quantitatively, the capacity to measure phenomes. In broad definition, phenome includes epigenomics, transcriptomics, proteomics, metabolomics and many other “omics” data regarding quantitative measurement of biochemical and cellular processes. We have relatively well-developed technologies of measurements, in vivo or in destructive manners, of physiological states and other “internal phenotypes” (endophenotypes), such as gene expression, protein and metabolite levels, whereas our ability to measure “external phenotypes” (exophenotypes) is rapidly evolving.
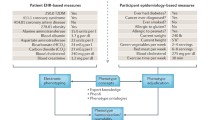
We will never be able to come even close to a complete characterisation of the phenome due to its highly dynamic and high-dimensional properties. However, increasing the quantitative information obtained by phenotypic measurements is an important goal for phenomics [3]. Phenotypic variation, a fundamental prerequisite and the perpetual force for evolution by natural selection, results from the complex interactions between genotype and environment (G × E). Phenomic-wide data are essential and necessary for enabling us to trace causal links in the genotype–phenotype map (G-P map [33]) as they define the space of all possible phenotypes (P space; Fig. 11.1).
The genotype–phenotype map (G-P map). The left panel shows the relationship of the genotype space (G space) and the phenotype space (P space) [3]. The corresponding information that transmits from G space to P space is shown in the right panel. Genotypes could gain mutation and recombination over generations. Phenotypes can be broadly classified into internal and external phenotypes. These internal phenotypes include properties from molecular, cellular or tissue levels, which in turn shape external phenotypes such as morphology and behaviour. Upon the environmental stimuli, the epigenetic process creates the phenotypes using genotype information. External phenotypes can in turn shape the environment that an individual occupies, creating complex feedback relationships between genes, environments and phenotypes. Natural selection act in the P space to change the average phenotype of parents away from the average phenotype of the generation. The importance of the environment suggests that we should explicitly broaden the G-P map to the genotype–environment–phenotype (G-E-P) map. g: genotype; p: phenotype; ip: internal phenotype
High-throughput automated imaging is the ideal tool for phenomic studies. Owing to the recent increased availability of high-precision robotic handling machinery, many imaging-based technologies that span molecular to organismal spatial scales have been or are being established and enable us to extract multiparametric phenotypic information in great detail. Various detectors using a broad range of the electromagnetic spectrum and magnetic resonance imaging (MRI) with different scales of resolution are widely used imaging techniques for phenotyping [34]. High-dimensional spatiotemporal data on many phenotype classes such as morphology, behaviour, physiological state and locations of proteins and metabolites can be captured by these imaging techniques and analysed via high-performance computing [3]. In recent years, systems for performing high-content microscopy-based assays have become available and are often used to investigate the effects of chemical (such as drugs and small molecules) and genetic (loss-of-function of genes using RNA interference [RNAi]) perturbations on cultured cells [35–42]. Such genome-wide RNAi screens enable us to discover novel gene functions and interrogate their functional relationships based on phenotypic similarity analysis [43, 44]. These screens produced huge amount of high-content image data that can be automatically processed using software tools such as ImageJ [45], EBImage [46], CellProfiler [47] or PhenoRipper [48]. Traditional microscopy is generally used in two-dimensional (2D) imaging. However, high-resolution and dynamic three-dimensional (3D) imaging data can be acquired by confocal laser scanning microscopy (CLSM), X-ray computerised tomography (CT) or MRI.
In plants, the “phenotyping bottleneck” [4] needs to be addressed by high-throughput noninvasive technologies [49]. Thanks to developed new imaging sensors (e.g. high-resolution imaging spectrometers) and the advanced software for image analysis and feature extraction, a range of automated or semiautomated high-throughput plant phenotyping systems (Table 11.3) have been recently developed and applied to assess plant function and performance under controlled conditions [50–58]. One of the pioneer platforms, PHENOPSIS [51], was developed for the dissection of genotype × environment effects on different processes in Arabidopsis thaliana with reproducible phenotyping. TraitMill [50, 52], GROWSCREEN [53, 55, 59], LIMINA [54], HYPOTrace [56], HTPheno [57] and LeafAnalyser [58] provide general image-processing solutions for plant morphological measurements (such as plant height, length and width, shape, projected area and biovolume) and colorimetric analysis. Most recently, high-throughput phenotyping has been used for three-dimensional plant analysis [60–64], focusing on a specific organs (e.g. leaves, roots and aerials). However, most of these tools possess the inherent disadvantage that they are designed to address only very specific question [65]. Among the advancing solutions, the state-of-the-art phenotyping platform developed by LemnaTec (http://www.lemnatec.com/) is a robotic greenhouse system that uses non-destructive imaging to monitor plant growth under controlled environmental conditions (such as nutrition, water availability, irradiation and temperature) over a period of time. Several ingenious imaging cameras, such as visible/colour/RGB (red, blue and green) imaging, fluorescence, thermal and near-infrared imaging, have been adopted in this system to assess the physical and physiological status of plants, such us their geometric properties, pigment or fluorophore contents, canopy temperature and tissue water content. LemnaTec systems have now been deployed in growth champers or greenhouses (e.g. at the Leibniz Institute of Plant Genetics and Crop Plant Research [IPK; Germany], the Australian Centre for Plant Functional Genomics [ACPFG] at the University of Adelaide [Australia], the Aberystwyth University [UK] and the PhenoArch at Institut National de la Recherche Agronomique at Montpellier [France]) for high-throughput phenotyping in Arabidopsis [66], wheat [67], barley [57] and maize (unpublished data). The time-lapse phenotypic data from these large-scale phenotyping platforms provide an invaluable opportunity to model and predict plant growth [67, 68]. Also, these data can be used to map quantitative trait loci (QTL) for growth-related traits. Notably, a recent phenotyping application was developed for QTL mapping in pepper plants using phenotypic features such as leaf angle and leaf size from RGB images, resulting in heritabilities of 0.56 and 0.70, respectively [69]. At the same time, however, the huge amounts of imaging data generated from these platforms present a great challenge for data analysis. As one solution, the Integrated Analysis Platform (IAP; http://iap.ipk-gatersleben.de) [70] is being developed as a comprehensive framework for high-throughput phenotyping in plants, which enables us to extract a high-dimensional list of plant features from real-time images to quantify plant growth and performance.
2.3 Defining Genotype–Phenotype Relationships
Understanding the interplay between genotype and phenotype (G-P map; Fig. 11.1) is the ultimate goal in both genomics and phenomics research, which will yield insights that are important for predicting disease risk and individual therapeutic treatments in human population, for increasing the speed of selective breeding traits in agriculturally import crops and for predicting adaptive evolution [71]. The interactions between genotypes and phenotypes also inevitably involve the environmental factors [3]. Thus, the interaction between genotype and phenotype has often been conceptualised by the following relationship: genotype (G) + environment (E) + genotype × environment (G × E) → phenotype (P). Since individuals themselves may influence the environment and exert different effects depending on their characteristics, feedback of phenotypes needs to be considered in this concept. Furthermore, the response of a certain genotype to an environmental factor may depend strongly on the phenotypic status of the individual, which is the result of events that occurred in its preceding life history. Towards understanding, the G-P map will provide a framework for the development of personalised medicine and crop breeding [72, 73].
Genomics and other highly parallel technologies – including epigenomics, transcriptomics, proteomics, metabolomics and ionomics – have become the mainstay in biological research. These recently developed technologies commonly termed “omics” permit assessment of the entirety of the components of biological systems at broad levels (Table 11.1). Furthermore, the emerging high-throughput phenotyping technology is moving towards comprehensive, quantitative high-dimensional measurements of individuals (phenome). However, our current knowledge of the genetic basis of complex phenotypic traits probably represents only the tip of the iceberg. Why do even genetically identical twins often substantially differ in phenotypic traits such as disease risk and drug response? Indeed, it is now understood that the differences are to a large extent result of the epigenome and involve chromatin modifications as well as myriads of noncoding RNAs (ncRNAs) [74, 75]. The emerging task is to understand the complex relationships among the genome, the epigenome, the environment and the phenome. The goal of globally linking genotype to phenotype can only be achieved through integrating information from different levels into an integrative model in terms of systems-biology approaches, which makes prediction of phenotypes possible (Fig. 11.2). This model should also consider the complex environmental factors in the real world, which need to be very precisely defined. For example, it is now possible to model rice transcriptome dynamics under fluctuating field conditions [76], rising hopes to predict genome-wide transcriptional responses in the complex real-world settings [77].
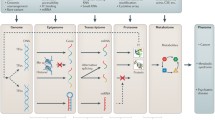
Chart flow of the assessment of gene function using quantitative trait locus (QTL) analyses. Genetic markers (DNA level) such as SNPs and CNVs can be genotyped using next-generation sequencing technology. Quantitative traits, such as DNA methylation level, transcript, protein or metabolite content and biomass can be analysed using different detection methods. The information flow is indicated with arrows. Environmental factors are also included. The data generated can be used for mapping to determine the genomic regions (QTLs) responsible for the observed variation. The identification of the causal genes underlying the QTL, and ultimately their functional characterisation, will be facilitated by the combined analysis of the data generated using different profiling techniques and additional information obtained using bioinformatics tools [78]. phQTLs: DNA methylation QTLs; eQTLs: expression QTLs; pQTLs: protein QTLs; mQTLs: metabolic QTLs; phQTLs: phenotypic QTLs; GWAS: genome-wide association studies; EWAS: epigenome-wide association studies; MWAS: metabolome-wide association studies
3 Approaches for Linking the Genome to the Phenome
3.1 QTL Detection Through Linkage and Association Mapping: Identifying the Genetic Basis of Complex Traits
Thanks to the advanced high-throughput experimental technologies such as microarray and sequencing, high-density genotyping arrays are available and are widely used recently to establish large-scale genome-wide maps of QTLs for various phenotypes such as human diseases and agricultural traits [20, 79–81]. Genome-wide association studies (GWAS, also called association mapping) are becoming the preferred method to relate genetic variation to phenotypic diversity in populations of unrelated individuals. The most common polymorphic markers used for GWAS are sequence polymorphisms such as SNPs and structural variants such indels and CNVs [20]. GWAS are now preferred over traditional family-based linkage studies (linkage-based QTL mapping; Fig. 11.3a) [82], which use interval mapping to estimate the map position and effect of each QTL.
Principle of quantitative trait locus (QTL) mapping. (a) Linkage-based mapping versus association mapping. The purpose of QTL mapping is to uncover the genetic basis of quantitative traits of interest. Linkage-based analyses seek to identify segregating genetic markers (M1, M2, M3 and M4) that predict the organismal phenotype, using a population that carries genetic mosaics derived from parental varieties, such as second generation (F2) plants or recombinant inbred lines (RILs). The relationships of individuals are known (P1 → F1 → F2 → RILs). It should be noted that RILs, rather than the F2 or F3 population, are needed to evaluate genotype-by-environment interactions. The region highlighted in yellow indicates the position of a causal locus or QTL. Association mapping (analogous to genome-wide association study [GWAS]) relies on correlations between genetic markers and a phenotype among collections of diverse germplasm. Thus, the recombination used in this strategy is historical. As shown in the figure, the association mapping population is separated by many generations from its progenitors. In linkage-based studies, the haplotype blocks in the mapping population may be large and, as a consequence, the causal locus might only be mapped to a large region. The haplotype blocks in an association mapping population tend to be much smaller, so it might be possible to localise the causal locus to a small genomic region. Within the QTL region, relevant genes may be identified for future studies or candidates may be suggested for targeted sequencing or experimental perturbation. (b) Conception of the intermediate phenotype used QTL mapping. The association of genetic variants is strongest with their closest intermediate phenotypes (IPs), such as variation of DNA methylation (methQTLs), transcript (eQTLs) or protein (pQTLs) content and metabolic traits (mQTLs). In some cases, the association of genetic variants with the organismal end point may not even be detectable at a level of genome-wide significance. (c) Relationships of GWAS and QTL mapping methodologies in integrative analyses (Part a is reproduced, with permission, from Mackay et~al. [98], Copyright 2009, Macmillan Publishers Ltd. Part b is adapted from Suhre and Gieger [136]. Part c is reproduced from Cookson et~al. [123])
GWAS use dense maps of genetic markers that cover the whole genome to look for allele-frequency differences between cases (e.g. patients with a specific disease or individuals with a certain trait) and controls. Several powerful statistical methods have been established to associate common complex trait with genomic variations, including efficient mixed-model association (EMMA) [83], EMMA expedited (EMMAX) [84], genome-wide EMMA (GEMMA) [85], mixed-model and regression (GRAMMAR) [86], fast linear mixed models (FaST-LMM) [87], general linear model and mixed linear model implemented in TASSEL (Trait Analysis by aSSociation, Evolution and Linkage) [88] and the EIGENSTRAT method [89]. In the past few years, intensive efforts in more than 1,500 GWAS have uncovered hundreds of genetic variants associated with hundreds of diseases and other traits [90], providing valuable insights into the complexities of genetic architecture of human diseases. Although disease-associated variants in protein-coding regions are expected to be more importantly related to trait/disease diversity, the vast majority (80 %) of variants are found to fall outside coding regions, highlighting the importance of noncoding regions in the search for disease-associatedvariants [1, 90]. However, the identified loci thus far explain only a small fraction of the phenotypic diversity in humans, raising questions regarding “the missing heritability” [1, 91]. An informative example is the investigation of height in humans, which is 80–90 % heritable, but a list of loci that has been detected in GWAS together accounts for less than 5 % of heritability for height [92]. Several explanations for this missing heritability have been proposed, including rare variants, allelic heterogeneity, epigenetic variation (see the next section), CNVs, gene–gene interactions and, perhaps most importantly, the environmental uncertainty [1, 91]. Intriguingly, GWAS have shown to be even more successful in plants than in humans [93], the key observation being that initial GWAS in plants (e.g. in Arabidopsis [94], maize [95, 96] and rice [28]) have explained a much greater proportion of the phenotypic variation. Perhaps the best example is a study in rice [28], in which the authors performed low-coverage resequencing of the genomes of a panel of about 500 rice landraces and identified 80 loci associated with 14 agronomic traits, explaining on average ~36 % of the phenotypic variance. Several of these loci matched previously characterised genes. The ongoing development of technologies in both genotyping for detection of CNVs and other structural variants and statistical methods for accurate association testing will help us to examine potential sources of missing heritability and to better illuminate the causality of complex traits/diseases.
Linkage-based QTL mapping approaches have proved to be enormously successful for plant breeding and have identified loci with large effects of genetic variants on complex traits, which include most agriculturally important traits [81, 97]. The primary advantages of QTL mapping in plants are the great feasibility of creating populations of segregating individuals showing measurable phenotypic variation. However, the generation of crosses is time-consuming, and there is the necessity to focus on traits that can be readily and accurately phenotyped. Furthermore, due to the low frequency of recombinations represented in biparental mapping populations, causal loci (QTLs) identified by linkage-based strategies can only be mapped to large chromosomal regions, and tedious fine mapping needs to be carried out to narrow down on candidate genes that can be subjected to targeted sequencing or experimental perturbation [97, 98].
The emergence of a next-generation of mapping populations [97] overcomes many of the limitations of biparental QTL mapping and association mapping. Such experimental designs combine association and linkage analysis as they involve the crossing of multiple parents and advance populations through several generations to increase allelic richness and to improve resolution in genetic mapping. Such designs include the nested association mapping (NAM) [95, 99, 100], the multiparent advanced generation intercross (MAGIC) [101, 102] and the recombinant inbred advanced intercross line (RIAIL) [103, 104] populations.
In a further aspect, it needs to be mentioned that genomic selection (GS) [105], a genomics-based strategy for predicting phenotypes by the use of genome-wide marker data, is receiving considerable attention among (animal and) plant breeders. Similar to linkage and association mapping methods, GS starts with the development of a prediction model on a training population with individuals characterised for genotype and phenotype. Unlike linkage and association mapping approaches, GS models consider all markers as predictors and can thus capture more of the variation due to small-effect QTLs. Most importantly, the training population used in GS is generally closely related to the breeding population under selection. This situation supports the use of GS models for most accurate predictions for breeding [106].
3.2 EWAS: Linking Epigenetic Variation and Complex Traits
In addition to genetic variability, epigenetic factors including DNA methylation, histone modifications and ncRNAs (e.g. small interfering RNAs [siRNAs], microRNAs [miRNAs] and large intergenic ncRNAs [lincRNAs]) are considered as the missing part of the underlying molecular control of phenotypic variation (Table 11.1) [71, 75]. DNA methylation is the most studied epigenetic modification, and its variation at a single CpG (cytosine–guanine dinucleotide) site (known as a methylation variable position, MVP), CHG (H = A, T or C) or CHH contexts or a differentially methylated region (DMR) can be considered as the epigenetic equivalent (heritable epigenetic polymorphism) of an SNP in the context of genome [107]. While the DNA-centric model (e.g. GWAS) has allowed scientists to uncover the molecular genetic origins of Mendelian traits and diseases successfully, many complex traits and diseases are non-Mendelian, making them hard to explain. Due to the elasticity and plasticity of epigenetic factors, epigenetics can provide a novel framework for the identification of aetiological factors in complex traits and diseases [108]. The direct evidence that epigenetics could “make the difference” comes from the remarkably different epigenetic profiles, including disease-associated epigenetic differences, in human monozygous (MZ) twins, who share an identical genotype [109–111]. Indeed, with the recent advances in genomic technologies, the large-scale, systematic epigenomic equivalents of GWAS, termed as epigenome-wide association studies (EWAS), are emerging as the promising tool to investigate human disease-associated epigenetic variation [71]. However, it is still challenging in EWAS to distinguish whether epigenetic variation is the cause or functional consequence of the identified effects. In this regard, the sample used in an EWAS should ideally consist of MZ twins, to eliminate the influence of genetic background on the identified epigenetic variation [71] and as recently demonstrated by several studies [112–115]. Analysis of epigenetic variation is likely to be most successful when integrating the analysis of genetic variants (i.e. QTL mapping), leading to the identification of the underlying genetic variants that influence epigenetic state (epigenotype). The loci that harbour genetic variants corresponding to methylation states (e.g. MVPs or DMRs) have thus been termed methylation QTLs (methQTLs) [116]. The most pronounced methQTLs influence epigenetic states in cis, and they reside less than 50 bp from the CpG site in question [112]. The notion of methQTLs provides a general idea for integrated GWAS and EWAS (Fig. 11.3) to explore genotypes that exert their function through epigenetic mechanisms, which can be maintained and propagated during cell division, resulting in permanent maintenance of the acquired phenotype [71, 108, 117].
At the same time, there is also evidence from plant research communities that naturally occurring epigenetic changes (i.e. DMRs) in a single gene locus (epiallele) can lead to heritable phenotypic variation [118–122]. The epialleles often show increased cytosine methylation of the promoter and can result in nearby gene expression changes that are sometimes transmitted across generations, thus contributing to heritable phenotypic variation independent of DNA sequence diversity. These outstanding resources will advance our understanding of the relative roles of genetic and epigenetic variation in controlling quantitative trait variation in plants.
3.3 Variation in Gene Expression: From eQTLs to Phenotypes
Variation in gene expression is an important mechanism underlying phenotypic variation such as disease susceptibility and drug response. DNA variants may alter transcript abundance and splicing patterns through modification of regulatory elements [123]. Genomic loci responsible for this genetic control are consequently termed expression QTLs (eQTLs). The combination of high-throughput phenotyping and transcriptional profiling has allowed the systematic identification of eQTLs (Fig. 11.3) [98]. In principle, eQTL mapping uses transcript abundance as a phenotypic trait and maps the genomic loci controlling the transcript level, as performed in the same manner of traditional QTL mapping of any other quantitative trait phenotype [124]. According to the genomic context of transcripts, eQTLs can be categorised into cis eQTLs if the molecular variants (e.g. SNPs) are mapped to the approximate location (within 100 kb upstream and downstream [112, 125]) of their gene-of-origin transcripts and trans eQTLs in other cases. Further statistical analysis revealed a strong enrichment of cis eQTLs around transcription start sites (TSSs) and within 250 bp upstream of transcription end sites (TESs) [126]. The cis-acting variants are more likely in exonic regions than in intronic regions. Given that genetic variation in the 3′UTR of a gene may create or destroy a miRNA binding site [127], the cis effects are likely mediated through miRNA-regulated pathways. Besides this, cis-acting variants in promoter or enhancer regions may influence the binding of transcription factors and thus promoter regulation. Nevertheless, it is still not known whether trans effects are mediated through transcription factor variants or through other mechanisms [123]. Generally, cis eQTLs tend to have stronger influence on target gene regulation than trans eQTLs. Moreover, there exist the so-called eQTL hot spots in which the expression levels of many transcripts are associated with the variation.
The resulting comprehensive eQTL maps provide potential insight into a biological basis for complex quantitative trait associations identified through GWAS [123]. Since the expression of transcripts is subject to intensive gene regulation, eQTL data should be interpreted further by the incorporation of additional biological information, such as results from GWAS and EWAS as discussed above, and analysis of regulatory networks, which are discussed below. This kind of integrated analyses has been utilised in several studies [112, 114, 115, 128, 129].
Proteins are mainly responsible for the biological phenotype; they thus should more accurately reflect the cellular physiological state or the changes induced by disease processes, drug treatment or other influences, compared with genetic, epigenetic or transcript variants. Various mechanisms of post-transcriptional regulation can lead to changes in protein abundance in the absence of a corresponding alteration of transcript levels, suggesting that the proteome is expected to provide important biological insights and disease biomarkers that cannot be captured through evaluation of the transcriptome alone [130]. We mention here that association mapping analysis could also be done at the protein level in terms of protein QTL (pQTL or PQL [131]) mapping, in which protein abundance or modification is treated as a phenotypic trait. pQTL mapping, complementary to eQTL mapping, is now becoming feasible with technical advances in mass spectrometry (MS)-based proteomics [130, 132, 133]. The little overlap between pQTLs and eQTLs from the same study [134] indicates that the proteome and the transcriptome give distinct insights into the diversity between different individuals and further highlights the implications for systems-biology approaches that utilise such high-throughput data into integrated analysis.
3.4 Genome-Wide Association Studies with Metabolomics: Metabolic QTL Analysis
In addition to genomics, epigenomics, transcriptomics and proteomics, metabolomics is emerging as a complementary approach for globally measuring ideally all endogenous small organic molecules (metabolic traits; normally below 1,500 Da) in a biological sample. However, unlike the transcriptome and to a lesser degree the proteome, the metabolome is much more amenable to variation. The metabolome is much more diverse in terms of chemical structure and function [135]. Metabolite profiles capture important information on the environment (diet, lifestyle, gut microbial activity and bacterial activity) that individuals experience and can give an instantaneous snapshot of the individual’s physiological state at that particular time under a particular set of conditions. Some changes in metabolite levels may be a consequence of the phenotypic diversity; therefore, a metabolic trait presents a functional intermediate trait or merely a correlated biomarker [136]. Noninvasive metabolic methodologies include nuclear magnetic resonance (NMR) spectroscopy [137], MS and high-performance liquid-phase chromatography (HPLC). Due to advances in these technologies, quantitative readouts for hundreds of small molecules that are detected in large scale can now be provided. Experimental design concerns the choice of which metabolites to study. While targeted methods provide precise measurements of specific (known) metabolites and are easy to replicate, nontargeted approaches are currently more promising as they provide the opportunity to discover novel associations including hitherto uncharacterised metabolites [136].
In the past few years, GWAS face the challenge that the effect of sizes of genetic association is generally small and information on the underlying biological processes is lacking [136]. These problems can be overcome, at least partially, by association with metabolic traits as functional intermediates [138]. There is the increased interest from the scientific community, and particularly plant biologists, in integrating metabolic approaches into research with the aim to unravel phenotypic diversity and its underlying genetic variation [78]. The combination of high-throughput metabolic phenotyping with general QTL analysis has thus given birth to the emerging field of metabolome-wide association studies (MWAS; Fig. 11.3).
The study of the chemical composition (i.e. the metabolite) of plants has always been of great interest in biological research, in part because metabolic phenotypes (metabotypes) largely reflect the developmental stage of the plant and its interactions with the environment. In plants, the first studies combining metabolic phenotyping with QTL analysis were performed in tomato [139–141] and successfully uncovered loci (metabolite QTLs, mQTLs) regulating plant metabolite composition. In Arabidopsis [142–147] and other crops, such as Brassica napus [148, 149], potato [150], rice [151] and maize [138, 152], mQTL mapping analyses have also been implemented using targeted and nontargeted metabolic profiling. Metabolite profiling-based approaches furthermore provide important steps towards the goal of hybrid performance prediction [152] and metabolomics-assisted crop breeding [153].
Similar MWAS were later performed in human studies [154–158]. Large panels of metabotypes have been analysed in association with genetic variants, disease-related phenotypes and lifestyle and environmental parameters, allowing dissection of the contribution of these factors to the aetiology of complex diseases [136]. These MWAS have identified genetic factors reliably that influence intermediate traits on phenotypes such as blood pressure [158], cardiometabolic disorder [157] and coronary heart disease [159]. In summary, incorporation of GWAS and metabolomics further refine the G-P map and eventually identify possible prognostic or diagnostic biomarkers of disease risk and biomarkers for predictive plant breeding.
3.5 Systems Biology: Genome-Scale Networks That Link Genes to Phenotypes
Associating sequence-level variation (such as SNPs and CNVs) with high-level variation in organismal phenotypes (such as disease susceptibility or crop yield) omits all of the intermediate steps in the chain of causation from genetic perturbation to phenotypic diversity. As mentioned above, intermediate molecular phenotypes (endophenotypes) such as epigenetic variation, transcript/protein abundance and metabolic traits vary genetically in populations and are themselves quantitative traits [98]. These endophenotypes functionally link genetic variation to disease-predisposing (for human) or biomass-predisposing (for plants) factors and then to complex phenotypic end points. Excitingly, the so-called “genetical genomics” approach [160] now enables us to integrate genetic variation, various endophenotypic variation and variation in organismal phenotypes in a linkage or association mapping population in both human [161] and plants [162], allowing to interpret quantitative genetic variation in terms of biologically meaningful causal networks of correlated transcripts.
However, it is becoming clear that each of the intermediate steps in translating biological information from genotype to phenotype does not stand alone [135]. The omics technologies now enable us to understand the biology inside the “black box” that lies between genotype and phenotype in terms of complex interacting networks [135, 163] (Fig. 11.4). Although we are still far away from a holistic understanding of the G-P map, systems biology is an emerging approach that aims to elucidate higher-level behaviour of biological systems and focuses on complex interactions within them, illuminating the path towards this ultimate goal – the complete G-P map. The integrative systems approach tries to link together the single-level omics data (e.g. genome, epigenome, transcriptome, proteome and metabolome) and, over time (if available [164]), to reveal and model the dynamic molecular regulatory networks or pathways from gene-to-function in order to bridge from genomics to phenomics. With the availability of increasingly powerful omics-based technologies, analytical and statistical tools and integrated knowledge bases, it has become possible to establish new links between genes, biological functions and a wide range of human diseases [165–179]. The comprehensive gene-disease associations present important insights that different disease modules (i.e. diseases share common genetic origins) could overlap and perturbations caused by one disease could affect other disease modules [180]. The identification of disease modules leads to the concept of the diseasome [165], which represents disease networks whose nodes are diseases and whose links represent the shared molecular relationships between the disease pairs. The underlying disease-associated cellular components are mostly investigated with protein-coding genes [165, 166, 168, 176, 177], though miRNAs [173, 178, 181], large intergenic noncoding RNAs (lincRNAs) [175] or metabolic pathways [171] are also investigated. Importantly, uncovering such diseasome networks provides hints on how different phenotypes are linked at the molecular level.
Schematic diagram depicting the strategy for integrated analysis of genetic and omic data. Large-scale genotyping and phenotyping are performed on segregating populations. Quantitative traits can be analysed on different levels to identify responsible loci (QTLs) based on QTL mapping approaches. Retrieved data can also be used in cluster analyses to identify gene-centred networks. The methodology of the combined used of genetic and omic technologies is commonly referred to as “genetical genomics” [160] and enables the elucidation of complex gene–phenotype networks (the G-P maps). This figure extends the work from Keurentjes [135]
Although GWAS and analogous methodologies have presented large numbers of disease-gene candidates, it still has the difficulty to identify the particular gene and the causal mutation [180]. A series of sophisticated strategies have recently been developed to predict potential disease genes (Fig. 11.5). These network-based tools include linkage methods [182], functional module-based or “guilt-by-association” methods [166, 176, 177] and diffusion-based methods [183, 184]. Furthermore, it is believed that genes tend to work in evolutionarily conserved pathways or modules; so the G-P maps can potentially be transferred between different species. Based on this assumption, orthologous phenotypes (phenologs) can be used to systematically predict genes associated nonobviously with diseases across different organisms using overlapping sets of orthologous genes [185]. In summary, the value of these tools is expected to increase with the wealth of disease gene candidates beyond GWAS. Although most of the initial studies based on these tools were performed in humans, similar strategies can also be applied to the plant biological research [186]. Indeed, networks for Arabidopsis [187], rice [188, 189] and maize [189] have been shown to connect thousands of genes accurately to phenotypes.
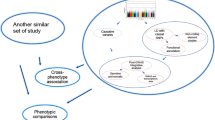
Methodologies for identifying trait-associated gene candidates. (a) Linkage methods. These methods combine both the linkage analysis (to determine the linkage interval of a specific trait) and protein–protein interaction (PPI) information. Genes (denoted as G1, G2 and so on) located in the linkage interval whose protein products interact with a known trait-associated protein are considered likely candidate genes. (b) Functional module-based or guilt-by-association methods. Function modules are identified from clustering analysis of genome-scale networks. The members of such modules are considered candidate genes linked to specific phenotypes. (c) Diffusion-based methods. Starting from proteins that are known to be associated with a phenotype, a random walker visits each node in the interactome with a certain probability. The outcome of this algorithm is a trait-association score that is assigned to each protein, that is, the likelihood that a particular protein is associated with the phenotype. (d) Phenologs (orthologous phenotypes). Phenologs is used to map phenotypes between organisms based on significantly overlapping sets of orthologous genes. Perturbation of overlapping modules of orthologous genes may result in one set of phenotypes in one organism but a different set of phenotypes in another organism. The genes in such modules are considered candidates associated with the corresponding phenotypes (Parts a–c are modified from Barabasi et~al. [180]. Part d is modified from McGary et~al. [185])
4 Perspectives and Future Challenges
The basic requirements for building an ideal phenomics realm are easy to imagine but still hard to realise. We are facing great opportunities but also great challenges in the areas of both genomics and phenomics. Although technically feasible, extensive and intensive measurement of genetic contents (such as epigenetic modification, gene expression, metabolite content) on large samples of genotypes across the full range of spatial and temporal scales is costly. Furthermore, the high density of genetic markers identified thus far yet awaits to be linked to their consequential phenotypic traits. On the phenomics side, the major challenge resides in the multitudes of phenotypic traits and environmental influences. The cost of a phenome project using current technology is extremely high [3]. High-throughput and high-resolution phenotyping technologies, for detection of both internal and external phenotypes, especially in plants, have started to open new horizons [3, 49]. Extracting as much quantitative information as possible from phenotyping data is a fundamental goal for phenomics. In other words, future phenomic efforts need to focus on comprehensive and quantitative measurements of phenotypes, rather than conventionally low-dimensional and qualitative phenotype categorisations [3]. Developments in phenomics will increase both the number of phenotypic traits that are quantitatively assessed and the sample sizes (number of individuals or genotypes characterised), resulting in major challenges with respect to data analysis. The available state-of-the-art methods, such as partial least squares (PLS) regression, principal component analysis (PCA), random forests (RF) and support vector machines (SVM), can be used to address the high-dimensional phenomic data. Another challenge in new analytics is automated analysis of phenotyping data, since navigating the huge imaging data sets manually is extremely tedious.
Regarding linking genotype to phenotype, many important challenges remain: (a) with respect to the problem of linking genes to traits, according to the observation of vast numbers of associated variants located within noncoding regions of the genome [90]; (b) with respect to epistatic interactions [190]; (c) with respect to gene-environment interactions [191]; (d) with respect to epigenetic influences on phenotypic variation; and (e) with respect to variation in the outcome of mutations among individuals [73]. One promising solution here is to combine data from multiple “omics” technologies in what may be termed “a genome-wide systems-biology approach”.
In a nutshell, however, phenomics lags largely behind genomics. In contrast to the situation in humans, in plant organisms it is relatively straightforward to carry out systematic genetic screens and large-scale phenotyping under various controlled environments. This provides unbiased assessment of the genetic complexity of phenotypic traits [73]. The G-P maps are therefore ultimately expected to be more complete and more systematic in plants than they may be in humans. Notably, many ongoing developing or developed phenomics tools will give plant scientists the power to unlock the information coded in genomes (Table 11.3). In the near future, the plant phenotypic landscape will be populated at a faster pace to accelerate research in model organisms and to bridge the gap between genomics and phenomics [3, 49].
References
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A et al (2009) Finding the missing heritability of complex diseases. Nature 461(7265):747–753
Freedman ML, Monteiro AN, Gayther SA, Coetzee GA, Risch A, Plass C, Casey G, De Biasi M, Carlson C, Duggan D et al (2011) Principles for the post-GWAS functional characterization of cancer risk loci. Nat Genet 43(6):513–518
Houle D, Govindaraju DR, Omholt S (2010) Phenomics: the next challenge. Nat Rev Genet 11(12):855–866
Furbank RT, Tester M (2011) Phenomics–technologies to relieve the phenotyping bottleneck. Trends Plant Sci 16(12):635–644
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Kan YW, Dozy AM (1978) Polymorphism of DNA sequence adjacent to human beta-globin structural gene: relationship to sickle mutation. Proc Natl Acad Sci U S A 75(11):5631–5635
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M et al (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23(21):4407–4414
Bodmer WF (1986) Human genetics: the molecular challenge. Cold Spring Harb Symp Quant Biol 51 Pt 1:1–13
Kruglyak L (1997) The use of a genetic map of biallelic markers in linkage studies. Nat Genet 17(1):21–24
Wang DG, Fan JB, Siao CJ, Berno A, Young P, Sapolsky R, Ghandour G, Perkins N, Winchester E, Spencer J et al (1998) Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science 280(5366):1077–1082
Gunderson KL, Steemers FJ, Lee G, Mendoza LG, Chee MS (2005) A genome-wide scalable SNP genotyping assay using microarray technology. Nat Genet 37(5):549–554
Syvanen AC (2005) Toward genome-wide SNP genotyping. Nat Genet 37(Suppl):S5–S10
Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat Rev Genet 12(7):499–510
Nielsen R, Paul JS, Albrechtsen A, Song YS (2011) Genotype and SNP calling from next-generation sequencing data. Nat Rev Genet 12(6):443–451
Van Tassell CP, Smith TP, Matukumalli LK, Taylor JF, Schnabel RD, Lawley CT, Haudenschild CD, Moore SS, Warren WC, Sonstegard TS (2008) SNP discovery and allele frequency estimation by deep sequencing of reduced representation libraries. Nat Methods 5(3):247–252
van Orsouw NJ, Hogers RC, Janssen A, Yalcin F, Snoeijers S, Verstege E, Schneiders H, van der Poel H, van Oeveren J, Verstegen H et al (2007) Complexity reduction of polymorphic sequences (CRoPS): a novel approach for large-scale polymorphism discovery in complex genomes. PLoS One 2(11):e1172
Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA, Selker EU, Cresko WA, Johnson EA (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS One 3(10):e3376
Andolfatto P, Davison D, Erezyilmaz D, Hu TT, Mast J, Sunayama-Morita T, Stern DL (2011) Multiplexed shotgun genotyping for rapid and efficient genetic mapping. Genome Res 21(4):610–617
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6(5):e19379
Frazer KA, Murray SS, Schork NJ, Topol EJ (2009) Human genetic variation and its contribution to complex traits. Nat Rev Genet 10(4):241–251
Thorisson GA, Stein LD (2003) The SNP Consortium website: past, present and future. Nucleic Acids Res 31(1):124–127
International HapMap C (2003) The International HapMap project. Nature 426(6968):789–796
Gonzaga-Jauregui C, Lupski JR, Gibbs RA (2012) Human genome sequencing in health and disease. Annu Rev Med 63:35–61
Genomes Project C, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, Hurles ME, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467(7319):1061–1073
Yi X, Liang Y, Huerta-Sanchez E, Jin X, Cuo ZX, Pool JE, Xu X, Jiang H, Vinckenbosch N, Korneliussen TS et al (2010) Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329(5987):75–78
Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491(7422):56–65
Atwell S, Huang YS, Vilhjalmsson BJ, Willems G, Horton M, Li Y, Meng D, Platt A, Tarone AM, Hu TT et al (2010) Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465(7298):627–631
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y, Li C, Zhu C, Lu T, Zhang Z et al (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42(11):961–967
Gore MA, Chia JM, Elshire RJ, Sun Q, Ersoz ES, Hurwitz BL, Peiffer JA, McMullen MD, Grills GS, Ross-Ibarra J et al (2009) A first-generation haplotype map of maize. Science 326(5956):1115–1117
Lai J, Li R, Xu X, Jin W, Xu M, Zhao H, Xiang Z, Song W, Ying K, Zhang M et al (2010) Genome-wide patterns of genetic variation among elite maize inbred lines. Nat Genet 42(11):1027–1030
Zheng LY, Guo XS, He B, Sun LJ, Peng Y, Dong SS, Liu TF, Jiang S, Ramachandran S, Liu CM et al (2011) Genome-wide patterns of genetic variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol 12(11):R114
Cockram J, White J, Zuluaga DL, Smith D, Comadran J, Macaulay M, Luo Z, Kearsey MJ, Werner P, Harrap D et al (2010) Genome-wide association mapping to candidate polymorphism resolution in the unsequenced barley genome. Proc Natl Acad Sci U S A 107(50):21611–21616
Waddington CH (1968) Towards a theoretical biology. Nature 218(5141):525–527
Walter T, Shattuck DW, Baldock R, Bastin ME, Carpenter AE, Duce S, Ellenberg J, Fraser A, Hamilton N, Pieper S et al (2010) Visualization of image data from cells to organisms. Nat Methods 7(3 Suppl):S26–S41
Perlman ZE, Slack MD, Feng Y, Mitchison TJ, Wu LF, Altschuler SJ (2004) Multidimensional drug profiling by automated microscopy. Science 306(5699):1194–1198
Neumann B, Held M, Liebel U, Erfle H, Rogers P, Pepperkok R, Ellenberg J (2006) High-throughput RNAi screening by time-lapse imaging of live human cells. Nat Methods 3(5):385–390
Bakal C, Aach J, Church G, Perrimon N (2007) Quantitative morphological signatures define local signaling networks regulating cell morphology. Science 316(5832):1753–1756
Loo LH, Wu LF, Altschuler SJ (2007) Image-based multivariate profiling of drug responses from single cells. Nat Methods 4(5):445–453
Jones TR, Carpenter AE, Lamprecht MR, Moffat J, Silver SJ, Grenier JK, Castoreno AB, Eggert US, Root DE, Golland P et al (2009) Scoring diverse cellular morphologies in image-based screens with iterative feedback and machine learning. Proc Natl Acad Sci U S A 106(6):1826–1831
Collinet C, Stoter M, Bradshaw CR, Samusik N, Rink JC, Kenski D, Habermann B, Buchholz F, Henschel R, Mueller MS et al (2010) Systems survey of endocytosis by multiparametric image analysis. Nature 464(7286):243–249
Fuchs F, Pau G, Kranz D, Sklyar O, Budjan C, Steinbrink S, Horn T, Pedal A, Huber W, Boutros M (2010) Clustering phenotype populations by genome-wide RNAi and multiparametric imaging. Mol Syst Biol 6:370
Mukherji M, Bell R, Supekova L, Wang Y, Orth AP, Batalov S, Miraglia L, Huesken D, Lange J, Martin C et al (2006) Genome-wide functional analysis of human cell-cycle regulators. Proc Natl Acad Sci U S A 103(40):14819–14824
Carpenter AE, Sabatini DM (2004) Systematic genome-wide screens of gene function. Nat Rev Genet 5(1):11–22
Pepperkok R, Ellenberg J (2006) High-throughput fluorescence microscopy for systems biology. Nat Rev Mol Cell Biol 7(9):690–696
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675
Pau G, Fuchs F, Sklyar O, Boutros M, Huber W (2010) EBImage–an R package for image processing with applications to cellular phenotypes. Bioinformatics 26(7):979–981
Carpenter AE, Jones TR, Lamprecht MR, Clarke C, Kang IH, Friman O, Guertin DA, Chang JH, Lindquist RA, Moffat J et al (2006) Cell Profiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol 7(10):R100
Rajaram S, Pavie B, Wu LF, Altschuler SJ (2012) PhenoRipper: software for rapidly profiling microscopy images. Nat Methods 9(7):635–637
Fiorani F, Schurr U (2013) Future scenarios for plant phenotyping. Annu Rev Plant Biol 64(1):267–291
Reuzeau C, Pen J, Frankard V, de Wolf J, Peerbolte R, Broekaert W (2005) TraitMill: a discovery engine for identifying yield-enhancement genes in cereals. Fenzi Zhiwu Yuzhong (Mol Plant Breed) 3:7534
Granier C, Aguirrezabal L, Chenu K, Cookson SJ, Dauzat M, Hamard P, Thioux JJ, Rolland G, Bouchier-Combaud S, Lebaudy A et al (2006) PHENOPSIS, an automated platform for reproducible phenotyping of plant responses to soil water deficit in Arabidopsis thaliana permitted the identification of an accession with low sensitivity to soil water deficit. New Phytol 169(3):623–635
Reuzeau C, Frankard V, Hatzfeld Y, Sanz A, Van Camp W, Lejeune P, De Wilde C, Lievens K, de Wolf J, Vranken E et al (2006) Traitmill™: a functional genomics platform for the phenotypic analysis of cereals. Plant Genet Resour 4(01):20–24
Walter A, Scharr H, Gilmer F, Zierer R, Nagel KA, Ernst M, Wiese A, Virnich O, Christ MM, Uhlig B et al (2007) Dynamics of seedling growth acclimation towards altered light conditions can be quantified via GROWSCREEN: a setup and procedure designed for rapid optical phenotyping of different plant species. New Phytol 174(2):447–455
Bylesjo M, Segura V, Soolanayakanahally RY, Rae AM, Trygg J, Gustafsson P, Jansson S, Street NR (2008) LAMINA: a tool for rapid quantification of leaf size and shape parameters. BMC Plant Biol 8:82
Jansen M, Gilmer F, Biskup B, Nagel KA, Rascher U, Fischbach A, Briem S, Dreissen G, Tittmann S, Braun S et al (2009) Simultaneous phenotyping of leaf growth and chlorophyll fluorescence via GROWSCREEN FLUORO allows detection of stress tolerance in Arabidopsis thaliana and other rosette plants. Funct Plant Biol 36(10–11):902–914
Wang L, Uilecan IV, Assadi AH, Kozmik CA, Spalding EP (2009) HYPOTrace: image analysis software for measuring hypocotyl growth and shape demonstrated on Arabidopsis seedlings undergoing photomorphogenesis. Plant Physiol 149(4):1632–1637
Hartmann A, Czauderna T, Hoffmann R, Stein N, Schreiber F (2011) HTPheno: an image analysis pipeline for high-throughput plant phenotyping. BMC Bioinform 12:148
Weight C, Parnham D, Waites R (2008) LeafAnalyser: a computational method for rapid and large-scale analyses of leaf shape variation. Plant J 53(3):578–586
Meyer RC, Kusterer B, Lisec J, Steinfath M, Becher M, Scharr H, Melchinger AE, Selbig J, Schurr U, Willmitzer L et al (2010) QTL analysis of early stage heterosis for biomass in Arabidopsis. Theor Appl Genet Theoretische und angewandte Genetik 120(2):227–237
Biskup B, Scharr H, Fischbach A, Wiese-Klinkenberg A, Schurr U, Walter A (2009) Diel growth cycle of isolated leaf discs analyzed with a novel, high-throughput three-dimensional imaging method is identical to that of intact leaves. Plant Physiol 149(3):1452–1461
Clark RT, MacCurdy RB, Jung JK, Shaff JE, McCouch SR, Aneshansley DJ, Kochian LV (2011) Three-dimensional root phenotyping with a novel imaging and software platform. Plant Physiol 156(2):455–465
Lobet G, Pages L, Draye X (2011) A novel image-analysis toolbox enabling quantitative analysis of root system architecture. Plant Physiol 157(1):29–39
Paproki A, Sirault X, Berry S, Furbank R, Fripp J (2012) A novel mesh processing based technique for 3D plant analysis. BMC Plant Biol 12:63
Wuyts N, Palauqui JC, Conejero G, Verdeil JL, Granier C, Massonnet C (2010) High-contrast three-dimensional imaging of the Arabidopsis leaf enables the analysis of cell dimensions in the epidermis and mesophyll. Plant Methods 6:17
Sozzani R, Benfey PN (2011) High-throughput phenotyping of multicellular organisms: finding the link between genotype and phenotype. Genome Biol 12(3):219
Arvidsson S, Perez-Rodriguez P, Mueller-Roeber B (2011) A growth phenotyping pipeline for Arabidopsis thaliana integrating image analysis and rosette area modeling for robust quantification of genotype effects. New Phytol 191(3):895–907
Golzarian MR, Frick RA, Rajendran K, Berger B, Roy S, Tester M, Lun DS (2011) Accurate inference of shoot biomass from high-throughput images of cereal plants. Plant Methods 7:2
Tardieu F, Tuberosa R (2010) Dissection and modelling of abiotic stress tolerance in plants. Curr Opin Plant Biol 13(2):206–212
van der Heijden G, Song Y, Horgan G, Polder G, Dieleman A, Bink M, Palloix A, van Eeuwijk F, Glasbey C (2012) SPICY: towards automated phenotyping of large pepper plants in the greenhouse. Funct Plant Biol 39(10–11):870–877
Klukas C, Pape JM, Entzian A (2012) Analysis of high-throughput plant image data with the information system IAP. J Integr Bioinform 9(2):191
Rakyan VK, Down TA, Balding DJ, Beck S (2011) Epigenome-wide association studies for common human diseases. Nat Rev Genet 12(8):529–541
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327(5967):818–822
Lehner B (2013) Genotype to phenotype: lessons from model organisms for human genetics. Nat Rev Genet 14(3):168–178
Wong AH, Gottesman II, Petronis A (2005) Phenotypic differences in genetically identical organisms: the epigenetic perspective. Hum Mol Genet 14(1):R11–R18
Meyer UA, Zanger UM, Schwab M (2013) Omics and drug response. Annu Rev Pharmacol Toxicol 53:475–502
Nagano AJ, Sato Y, Mihara M, Antonio BA, Motoyama R, Itoh H, Nagamura Y, Izawa T (2012) Deciphering and prediction of transcriptome dynamics under fluctuating field conditions. Cell 151(6):1358–1369
Jaeger PA, Doherty C, Ideker T (2012) Modeling transcriptome dynamics in a complex world. Cell 151(6):1161–1162
Carreno-Quintero N, Bouwmeester HJ, Keurentjes JJ (2013) Genetic analysis of metabolome-phenotype interactions: from model to crop species. Trends Genet 29(1):41–50
Altshuler D, Daly MJ, Lander ES (2008) Genetic mapping in human disease. Science 322(5903):881–888
Donnelly P (2008) Progress and challenges in genome-wide association studies in humans. Nature 456(7223):728–731
Takeda S, Matsuoka M (2008) Genetic approaches to crop improvement: responding to environmental and population changes. Nat Rev Genet 9(6):444–457
Ott J, Kamatani Y, Lathrop M (2011) Family-based designs for genome-wide association studies. Nat Rev Genet 12(7):465–474
Kang HM, Zaitlen NA, Wade CM, Kirby A, Heckerman D, Daly MJ, Eskin E (2008) Efficient control of population structure in model organism association mapping. Genetics 178(3):1709–1723
Kang HM, Sul JH, Service SK, Zaitlen NA, Kong SY, Freimer NB, Sabatti C, Eskin E (2010) Variance component model to account for sample structure in genome-wide association studies. Nat Genet 42(4):348–354
Zhou X, Stephens M (2012) Genome-wide efficient mixed-model analysis for association studies. Nat Genet 44(7):821–824
Aulchenko YS, Ripke S, Isaacs A, van Duijn CM (2007) GenABEL: an R library for genome-wide association analysis. Bioinformatics 23(10):1294–1296
Lippert C, Listgarten J, Liu Y, Kadie CM, Davidson RI, Heckerman D (2011) FaST linear mixed models for genome-wide association studies. Nat Methods 8(10):833–835
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23(19):2633–2635
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38(8):904–909
Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS, Manolio TA (2009) Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci U S A 106(23):9362–9367
Eichler EE, Flint J, Gibson G, Kong A, Leal SM, Moore JH, Nadeau JH (2010) Missing heritability and strategies for finding the underlying causes of complex disease. Nat Rev Genet 11(6):446–450
Maher B (2008) Personal genomes: the case of the missing heritability. Nature 456(7218):18–21
Brachi B, Morris GP, Borevitz JO (2011) Genome-wide association studies in plants: the missing heritability is in the field. Genome Biol 12(10):232
Li Y, Huang Y, Bergelson J, Nordborg M, Borevitz JO (2010) Association mapping of local climate-sensitive quantitative trait loci in Arabidopsis thaliana. Proc Natl Acad Sci U S A 107(49):21199–21204
Buckler ES, Holland JB, Bradbury PJ, Acharya CB, Brown PJ, Browne C, Ersoz E, Flint-Garcia S, Garcia A, Glaubitz JC et al (2009) The genetic architecture of maize flowering time. Science 325(5941):714–718
Li H, Peng Z, Yang X, Wang W, Fu J, Wang J, Han Y, Chai Y, Guo T, Yang N et al (2013) Genome-wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat Genet 45(1):43–50
Morrell PL, Buckler ES, Ross-Ibarra J (2011) Crop genomics: advances and applications. Nat Rev Genet 13(2):85–96
Mackay TF, Stone EA, Ayroles JF (2009) The genetics of quantitative traits: challenges and prospects. Nat Rev Genet 10(8):565–577
Yu J, Holland JB, McMullen MD, Buckler ES (2008) Genetic design and statistical power of nested association mapping in maize. Genetics 178(1):539–551
Kump KL, Bradbury PJ, Wisser RJ, Buckler ES, Belcher AR, Oropeza-Rosas MA, Zwonitzer JC, Kresovich S, McMullen MD, Ware D et al (2011) Genome-wide association study of quantitative resistance to southern leaf blight in the maize nested association mapping population. Nat Genet 43(2):163–168
Cavanagh C, Morell M, Mackay I, Powell W (2008) From mutations to MAGIC: resources for gene discovery, validation and delivery in crop plants. Curr Opin Plant Biol 11(2):215–221
Kover PX, Valdar W, Trakalo J, Scarcelli N, Ehrenreich IM, Purugganan MD, Durrant C, Mott R (2009) A multiparent advanced generation inter-cross to fine-map quantitative traits in Arabidopsis thaliana. PLoS Genet 5(7):e1000551
Rockman MV, Kruglyak L (2008) Breeding designs for recombinant inbred advanced intercross lines. Genetics 179(2):1069–1078
Balasubramanian S, Schwartz C, Singh A, Warthmann N, Kim MC, Maloof JN, Loudet O, Trainer GT, Dabi T, Borevitz JO et al (2009) QTL mapping in new Arabidopsis thaliana advanced intercross-recombinant inbred lines. PLoS One 4(2):e4318
Jannink JL, Lorenz AJ, Iwata H (2010) Genomic selection in plant breeding: from theory to practice. Brief Funct Genomics 9(2):166–177
Hamblin MT, Buckler ES, Jannink JL (2011) Population genetics of genomics-based crop improvement methods. Trends Genet 27(3):98–106
Rakyan VK, Hildmann T, Novik KL, Lewin J, Tost J, Cox AV, Andrews TD, Howe KL, Otto T, Olek A et al (2004) DNA methylation profiling of the human major histocompatibility complex: a pilot study for the human epigenome project. PLoS Biol 2(12):e405
Petronis A (2010) Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature 465(7299):721–727
Fraga MF, Ballestar E, Paz MF, Ropero S, Setien F, Ballestar ML, Heine-Suner D, Cigudosa JC, Urioste M, Benitez J et al (2005) Epigenetic differences arise during the lifetime of monozygotic twins. Proc Natl Acad Sci U S A 102(30):10604–10609
Javierre BM, Fernandez AF, Richter J, Al-Shahrour F, Martin-Subero JI, Rodriguez-Ubreva J, Berdasco M, Fraga MF, O’Hanlon TP, Rider LG et al (2010) Changes in the pattern of DNA methylation associate with twin discordance in systemic lupus erythematosus. Genome Res 20(2):170–179
Nguyen A, Rauch TA, Pfeifer GP, Hu VW (2010) Global methylation profiling of lymphoblastoid cell lines reveals epigenetic contributions to autism spectrum disorders and a novel autism candidate gene, RORA, whose protein product is reduced in autistic brain. Fed Am Soc Exp Biol 24(8):3036–3051
Gibbs JR, van der Brug MP, Hernandez DG, Traynor BJ, Nalls MA, Lai SL, Arepalli S, Dillman A, Rafferty IP, Troncoso J et al (2010) Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet 6(5):e1000952
Shoemaker R, Deng J, Wang W, Zhang K (2010) Allele-specific methylation is prevalent and is contributed by CpG-SNPs in the human genome. Genome Res 20(7):883–889
Bell JT, Pai AA, Pickrell JK, Gaffney DJ, Pique-Regi R, Degner JF, Gilad Y, Pritchard JK (2011) DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines. Genome Biol 12(1):R10
Bell JT, Tsai PC, Yang TP, Pidsley R, Nisbet J, Glass D, Mangino M, Zhai G, Zhang F, Valdes A et al (2012) Epigenome-wide scans identify differentially methylated regions for age and age-related phenotypes in a healthy ageing population. PLoS Genet 8(4):e1002629
Zhang D, Cheng L, Badner JA, Chen C, Chen Q, Luo W, Craig DW, Redman M, Gershon ES, Liu C (2010) Genetic control of individual differences in gene-specific methylation in human brain. Am J Hum Genet 86(3):411–419
Kilpinen H, Dermitzakis ET (2012) Genetic and epigenetic contribution to complex traits. Hum Mol Genet 21(R1):R24–R28
Vaughn MW, Tanurdzic M, Lippman Z, Jiang H, Carrasquillo R, Rabinowicz PD, Dedhia N, McCombie WR, Agier N, Bulski A et al (2007) Epigenetic natural variation in Arabidopsis thaliana. PLoS Biol 5(7):e174
Becker C, Hagmann J, Muller J, Koenig D, Stegle O, Borgwardt K, Weigel D (2011) Spontaneous epigenetic variation in the Arabidopsis thaliana methylome. Nature 480(7376):245–249
Eichten SR, Swanson-Wagner RA, Schnable JC, Waters AJ, Hermanson PJ, Liu S, Yeh CT, Jia Y, Gendler K, Freeling M et al (2011) Heritable epigenetic variation among maize inbreds. PLoS Genet 7(11):e1002372
Chodavarapu RK, Feng S, Ding B, Simon SA, Lopez D, Jia Y, Wang GL, Meyers BC, Jacobsen SE, Pellegrini M (2012) Transcriptome and methylome interactions in rice hybrids. Proc Natl Acad Sci U S A 109(30):12040–12045
Weigel D (2012) Natural variation in Arabidopsis: from molecular genetics to ecological genomics. Plant Physiol 158(1):2–22
Cookson W, Liang L, Abecasis G, Moffatt M, Lathrop M (2009) Mapping complex disease traits with global gene expression. Nat Rev Genet 10(3):184–194
Carlborg O, De Koning DJ, Manly KF, Chesler E, Williams RW, Haley CS (2005) Methodological aspects of the genetic dissection of gene expression. Bioinformatics 21(10):2383–2393
Dixon AL, Liang L, Moffatt MF, Chen W, Heath S, Wong KC, Taylor J, Burnett E, Gut I, Farrall M et al (2007) A genome-wide association study of global gene expression. Nat Genet 39(10):1202–1207
Veyrieras JB, Kudaravalli S, Kim SY, Dermitzakis ET, Gilad Y, Stephens M, Pritchard JK (2008) High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet 4(10):e1000214
Ryan BM, Robles AI, Harris CC (2010) Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer 10(6):389–402
Quon G, Lippert C, Heckerman D, Listgarten J (2013) Patterns of methylation heritability in a genome-wide analysis of four brain regions. Nucleic Acids Res 41(4):2095–2104
Emilsson V, Thorleifsson G, Zhang B, Leonardson AS, Zink F, Zhu J, Carlson S, Helgason A, Walters GB, Gunnarsdottir S et al (2008) Genetics of gene expression and its effect on disease. Nature 452(7186):423–428
Foss EJ, Radulovic D, Shaffer SA, Ruderfer DM, Bedalov A, Goodlett DR, Kruglyak L (2007) Genetic basis of proteome variation in yeast. Nat Genet 39(11):1369–1375
Damerval C, Maurice A, Josse JM, de Vienne D (1994) Quantitative trait loci underlying gene product variation: a novel perspective for analyzing regulation of genome expression. Genetics 137(1):289–301
Picotti P, Clement-Ziza M, Lam H, Campbell DS, Schmidt A, Deutsch EW, Rost H, Sun Z, Rinner O, Reiter L et al (2013) A complete mass-spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature 494(7436):266–270
Gstaiger M, Aebersold R (2009) Applying mass spectrometry-based proteomics to genetics, genomics and network biology. Nat Rev Genet 10(9):617–627
Ghazalpour A, Bennett B, Petyuk VA, Orozco L, Hagopian R, Mungrue IN, Farber CR, Sinsheimer J, Kang HM, Furlotte N et al (2011) Comparative analysis of proteome and transcriptome variation in mouse. PLoS Genet 7(6):e1001393
Keurentjes JJ (2009) Genetical metabolomics: closing in on phenotypes. Curr Opin Plant Biol 12(2):223–230
Suhre K, Gieger C (2012) Genetic variation in metabolic phenotypes: study designs and applications. Nat Rev Genet 13(11):759–769
Borisjuk L, Rolletschek H, Neuberger T (2012) Surveying the plant’s world by magnetic resonance imaging. Plant J Cell Mol Biol 70(1):129–146
Riedelsheimer C, Lisec J, Czedik-Eysenberg A, Sulpice R, Flis A, Grieder C, Altmann T, Stitt M, Willmitzer L, Melchinger AE (2012) Genome-wide association mapping of leaf metabolic profiles for dissecting complex traits in maize. Proc Natl Acad Sci U S A 109(23):8872–8877
Schauer N, Semel Y, Roessner U, Gur A, Balbo I, Carrari F, Pleban T, Perez-Melis A, Bruedigam C, Kopka J et al (2006) Comprehensive metabolic profiling and phenotyping of interspecific introgression lines for tomato improvement. Nat Biotechnol 24(4):447–454
Tieman DM, Zeigler M, Schmelz EA, Taylor MG, Bliss P, Kirst M, Klee HJ (2006) Identification of loci affecting flavour volatile emissions in tomato fruits. J Exp Bot 57(4):887–896
Schauer N, Semel Y, Balbo I, Steinfath M, Repsilber D, Selbig J, Pleban T, Zamir D, Fernie AR (2008) Mode of inheritance of primary metabolic traits in tomato. Plant Cell 20(3):509–523
Steinfath M, Strehmel N, Peters R, Schauer N, Groth D, Hummel J, Steup M, Selbig J, Kopka J, Geigenberger P et al (2010) Discovering plant metabolic biomarkers for phenotype prediction using an untargeted approach. Plant Biotechnol J 8(8):900–911
Meyer RC, Steinfath M, Lisec J, Becher M, Witucka-Wall H, Torjek O, Fiehn O, Eckardt A, Willmitzer L, Selbig J et al (2007) The metabolic signature related to high plant growth rate in Arabidopsis thaliana. Proc Natl Acad Sci U S A 104(11):4759–4764
Keurentjes JJ, Fu J, de Vos CH, Lommen A, Hall RD, Bino RJ, van der Plas LH, Jansen RC, Vreugdenhil D, Koornneef M (2006) The genetics of plant metabolism. Nat Genet 38(7):842–849
Lisec J, Meyer RC, Steinfath M, Redestig H, Becher M, Witucka-Wall H, Fiehn O, Torjek O, Selbig J, Altmann T et al (2008) Identification of metabolic and biomass QTL in Arabidopsis thaliana in a parallel analysis of RIL and IL populations. Plant J Cell Mol Biol 53(6):960–972
Lisec J, Steinfath M, Meyer RC, Selbig J, Melchinger AE, Willmitzer L, Altmann T (2009) Identification of heterotic metabolite QTL in Arabidopsis thaliana RIL and IL populations. Plant J Cell Mol Biol 59(5):777–788
Rowe HC, Hansen BG, Halkier BA, Kliebenstein DJ (2008) Biochemical networks and epistasis shape the Arabidopsis thaliana metabolome. Plant Cell 20(5):1199–1216
Lou P, Zhao J, He H, Hanhart C, Del Carpio DP, Verkerk R, Custers J, Koornneef M, Bonnema G (2008) Quantitative trait loci for glucosinolate accumulation in Brassica rapa leaves. New Phytol 179(4):1017–1032
Feng J, Long Y, Shi L, Shi J, Barker G, Meng J (2012) Characterization of metabolite quantitative trait loci and metabolic networks that control glucosinolate concentration in the seeds and leaves of Brassica napus. New Phytol 193(1):96–108
Carreno-Quintero N, Acharjee A, Maliepaard C, Bachem CW, Mumm R, Bouwmeester H, Visser RG, Keurentjes JJ (2012) Untargeted metabolic quantitative trait loci analyses reveal a relationship between primary metabolism and potato tuber quality. Plant Physiol 158(3):1306–1318
Matsuda F, Okazaki Y, Oikawa A, Kusano M, Nakabayashi R, Kikuchi J, Yonemaru J, Ebana K, Yano M, Saito K (2012) Dissection of genotype-phenotype associations in rice grains using metabolome quantitative trait loci analysis. Plant J Cell Mol Biol 70(4):624–636
Riedelsheimer C, Czedik-Eysenberg A, Grieder C, Lisec J, Technow F, Sulpice R, Altmann T, Stitt M, Willmitzer L, Melchinger AE (2012) Genomic and metabolic prediction of complex heterotic traits in hybrid maize. Nat Genet 44(2):217–220
Fernie AR, Schauer N (2009) Metabolomics-assisted breeding: a viable option for crop improvement? Trends Genet 25(1):39–48
Gieger C, Geistlinger L, Altmaier E, Hrabe de Angelis M, Kronenberg F, Meitinger T, Mewes HW, Wichmann HE, Weinberger KM, Adamski J et al (2008) Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum. PLoS Genet 4(11):e1000282
Illig T, Gieger C, Zhai G, Romisch-Margl W, Wang-Sattler R, Prehn C, Altmaier E, Kastenmuller G, Kato BS, Mewes HW et al (2010) A genome-wide perspective of genetic variation in human metabolism. Nat Genet 42(2):137–141
Suhre K, Shin SY, Petersen AK, Mohney RP, Meredith D, Wagele B, Altmaier E, CardioGram, Deloukas P, Erdmann J et al (2011) Human metabolic individuality in biomedical and pharmaceutical research. Nature 477(7362):54–60
Kettunen J, Tukiainen T, Sarin AP, Ortega-Alonso A, Tikkanen E, Lyytikainen LP, Kangas AJ, Soininen P, Wurtz P, Silander K et al (2012) Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat Genet 44(3):269–276
Holmes E, Loo RL, Stamler J, Bictash M, Yap IK, Chan Q, Ebbels T, De Iorio M, Brown IJ, Veselkov KA et al (2008) Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 453(7193):396–400
Shah SH, Bain JR, Muehlbauer MJ, Stevens RD, Crosslin DR, Haynes C, Dungan J, Newby LK, Hauser ER, Ginsburg GS et al (2010) Association of a peripheral blood metabolic profile with coronary artery disease and risk of subsequent cardiovascular events. Circ Cardiovasc Genet 3(2):207–214
Jansen RC, Nap JP (2001) Genetical genomics: the added value from segregation. Trends Genet 17(7):388–391
de Koning DJ, Haley CS (2005) Genetical genomics in humans and model organisms. Trends Genet 21(7):377–381
Fu J, Keurentjes JJ, Bouwmeester H, America T, Verstappen FW, Ward JL, Beale MH, de Vos RC, Dijkstra M, Scheltema RA et al (2009) System-wide molecular evidence for phenotypic buffering in Arabidopsis. Nat Genet 41(2):166–167
Keurentjes JJ, Koornneef M, Vreugdenhil D (2008) Quantitative genetics in the age of omics. Curr Opin Plant Biol 11(2):123–128
Chen R, Mias GI, Li-Pook-Than J, Jiang L, Lam HY, Chen R, Miriami E, Karczewski KJ, Hariharan M, Dewey FE et al (2012) Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell 148(6):1293–1307
Goh KI, Cusick ME, Valle D, Childs B, Vidal M, Barabasi AL (2007) The human disease network. Proc Natl Acad Sci U S A 104(21):8685–8690
Wu X, Jiang R, Zhang MQ, Li S (2008) Network-based global inference of human disease genes. Mol Syst Biol 4:189
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ, Lerner J, Brunet JP, Subramanian A, Ross KN et al (2006) The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313(5795):1929–1935
Schuster-Bockler B, Bateman A (2008) Protein interactions in human genetic diseases. Genome Biol 9(1):R9
Ramsey SA, Gold ES, Aderem A (2010) A systems biology approach to understanding atherosclerosis. EMBO Mol Med 2(3):79–89
Inouye M, Kettunen J, Soininen P, Silander K, Ripatti S, Kumpula LS, Hamalainen E, Jousilahti P, Kangas AJ, Mannisto S et al (2010) Metabonomic, transcriptomic, and genomic variation of a population cohort. Mol Syst Biol 6:441
Lee DS, Park J, Kay KA, Christakis NA, Oltvai ZN, Barabasi AL (2008) The implications of human metabolic network topology for disease comorbidity. Proc Natl Acad Sci U S A 105(29):9880–9885
Park J, Lee DS, Christakis NA, Barabasi AL (2009) The impact of cellular networks on disease comorbidity. Mol Syst Biol 5:262
Jiang Q, Hao Y, Wang G, Juan L, Zhang T, Teng M, Liu Y, Wang Y (2010) Prioritization of disease microRNAs through a human phenome-microRNAome network. BMC Syst Biol 4(Suppl 1):S2
Butte AJ, Kohane IS (2006) Creation and implications of a phenome-genome network. Nat Biotechnol 24(1):55–62
Ning S, Wang P, Ye J, Li X, Li R, Zhao Z, Huo X, Wang L, Li F, Li X (2013) A global map for dissecting phenotypic variants in human lincRNAs. Eur J Hum Genet 21(10):1128–1133
Lee I, Blom UM, Wang PI, Shim JE, Marcotte EM (2011) Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res 21(7):1109–1121
Lage K, Karlberg EO, Storling ZM, Olason PI, Pedersen AG, Rigina O, Hinsby AM, Tumer Z, Pociot F, Tommerup N et al (2007) A human phenome-interactome network of protein complexes implicated in genetic disorders. Nat Biotechnol 25(3):309–316
Iliopoulos D, Malizos KN, Oikonomou P, Tsezou A (2008) Integrative microRNA and proteomic approaches identify novel osteoarthritis genes and their collaborative metabolic and inflammatory networks. PLoS One 3(11):e3740
Pujana MA, Han JD, Starita LM, Stevens KN, Tewari M, Ahn JS, Rennert G, Moreno V, Kirchhoff T, Gold B et al (2007) Network modeling links breast cancer susceptibility and centrosome dysfunction. Nat Genet 39(11):1338–1349
Barabasi AL, Gulbahce N, Loscalzo J (2011) Network medicine: a network-based approach to human disease. Nat Rev Genet 12(1):56–68
Lu M, Zhang Q, Deng M, Miao J, Guo Y, Gao W, Cui Q (2008) An analysis of human microRNA and disease associations. PLoS One 3(10):e3420
Oti M, Snel B, Huynen MA, Brunner HG (2006) Predicting disease genes using protein-protein interactions. J Med Genet 43(8):691–698
Kohler S, Bauer S, Horn D, Robinson PN (2008) Walking the interactome for prioritization of candidate disease genes. Am J Hum Genet 82(4):949–958
Vanunu O, Magger O, Ruppin E, Shlomi T, Sharan R (2010) Associating genes and protein complexes with disease via network propagation. PLoS Comput Biol 6(1):e1000641
McGary KL, Park TJ, Woods JO, Cha HJ, Wallingford JB, Marcotte EM (2010) Systematic discovery of nonobvious human disease models through orthologous phenotypes. Proc Natl Acad Sci U S A 107(14):6544–6549
Bassel GW, Gaudinier A, Brady SM, Hennig L, Rhee SY, De Smet I (2012) Systems analysis of plant functional, transcriptional, physical interaction, and metabolic networks. Plant Cell 24(10):3859–3875
Lee I, Ambaru B, Thakkar P, Marcotte EM, Rhee SY (2010) Rational association of genes with traits using a genome-scale gene network for Arabidopsis thaliana. Nat Biotechnol 28(2):149–156
Lee I, Seo YS, Coltrane D, Hwang S, Oh T, Marcotte EM, Ronald PC (2011) Genetic dissection of the biotic stress response using a genome-scale gene network for rice. Proc Natl Acad Sci U S A 108(45):18548–18553
Ficklin SP, Feltus FA (2011) Gene coexpression network alignment and conservation of gene modules between two grass species: maize and rice. Plant Physiol 156(3):1244–1256
Lehner B (2011) Molecular mechanisms of epistasis within and between genes. Trends Genet 27(8):323–331
Thomas D (2010) Gene–environment-wide association studies: emerging approaches. Nat Rev Genet 11(4):259–272
Acknowledgement
This work was supported by grants from the Federal Office of Agriculture and Food (15/12-13, 530–06.01-BiKo CHN), the Robert Bosch Stiftung (32.5.8003.0116.0) and the Federal Ministry of Education and Research (BMBF – 0315958A).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
WWW List in This Chapter
WWW List in This Chapter
-
The NHGRI GWAS Catalogue: http://www.genome.gov/gwastudies/
A catalogue of published genome-wide association studies (GWAS)
-
LemnaTec: http://www.lemnatec.com/
High-throughput and high-content screening solutions for plant phenomics
-
IAP: http://iap.ipk-gatersleben.de/
An Integrated Analysis Platform (IAP) for plant high-throughput phenotyping data analysis
-
Note: Other useful links are listed in Tables 11.2 and 11.3.
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Chen, D., Chen, M., Altmann, T., Klukas, C. (2014). Bridging Genomics and Phenomics. In: Chen, M., Hofestädt, R. (eds) Approaches in Integrative Bioinformatics. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-41281-3_11
Download citation
DOI: https://doi.org/10.1007/978-3-642-41281-3_11
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-41280-6
Online ISBN: 978-3-642-41281-3
eBook Packages: Computer ScienceComputer Science (R0)