Abstract
Desulfarculaceae is the only family within the order Desulfarculales (Kuever et al. 2005a) that embraces a single genus Desulfarculus. Besides their 16S rRNA gene sequence phylogeny, the only member of the family is defined by a wide range of morphological and chemotaxonomic properties for the delineation to other genera and species. Strictly anaerobic, having a respiratory type of metabolism. Fermentative metabolism was not observed. Members are mesophilic sulfate-reducing bacteria. Members of the family are found in various habitats as indicated by clone sequences. The only described species is chemoorganoheterotroph and chemolithoheterotroph. The only member oxidizes organic substrates completely to carbon dioxide with sulfate as electron acceptor.
This contribution is a modified and updated version of a previous family description (Kuever et al. 2005b).
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
Keywords
- Fermentative Metabolism
- Serial Dilution Technique
- Respiratory Type
- Saturated Straight Chain Fatty Acid
- Distinct Phylogenetic Position
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Taxonomy, Historical and Current
Short Description of the Family
De.sul.far.cu.la’ce.ae. N.L. masc. n. Desulfarculus, type genus of the family; suff. -aceae, ending to denote family; N.L. fem. pl. n. Desulfarculaceae, the Desulfarculus family. The description is an emended version of the description given in Bergey´s Manual, 2nd edition (Kuever et al. 2005b, c).
The family belongs to the order Desulfarculales within the delta-Proteobacteria. The family Desulfarculaceae contains a single genus Desulfarculus (Kuever et al. 2006) which is the type genus of the family (Kuever et al. 2006). Gram-staining negative. Morphological forms are always vibrio-shaped cells. Spore formation is absent. The only member is motile by means of one or two polar flagella. Strictly anaerobic, having a respiratory type of metabolism. Fermentative metabolism was not observed. Formate and short- and long-chain fatty acids are used as electron donors. The only described species oxidizes organic substrates completely to carbon dioxide. The only member is mesophilic. The only described species is chemoorganoheterotroph and chemolithoheterotroph. Sulfate, sulfite, and thiosulfate are used as electron acceptor and reduced to sulfide. The only member has been isolated from a freshwater habitat. Similar organism might occur in brackish water and marine habitats.
Phylogenetic Structure of the Family
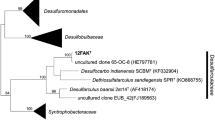
The phylogenetic structure of the family and its neighboring families is shown on Fig. 3.1 . The borders of the family are primarily based on the phylogenetic tree as framework and their unique properties (Physiology, chemotaxonomic markers) which are present in all members (see Table 3.1 ).
Phylogenetic reconstruction of the family Desulfarculaceae based on the neighbor-joining algorithm with the Jukes-Cantor correction. Sequence dataset and alignments according to the All-Species Living Tree Project release LTPs108 (Yarza et al. 2010). The tree topology was stabilized with the use of a representative set of around 750 high-quality-type strain sequences proportionally distributed among the different bacterial and archaeal phyla. In addition, a 40 % maximum frequency filter was applied to remove hypervariable positions from the alignment. Scale bar indicates estimated sequence divergence 2005
Molecular Analyses
DNA-DNA Hybridization Studies
Are absent.
Bioprinting and Ribotyping
Are absent.
MALDI-TOF
Are absent.
Genome Comparison
The complete genome sequence of the only described species of the genus Desulfarculus which is also the only member of this family which has been analyzed (Sun et al 2010). The genome of the type strain of Desulfarculus baarsii 2st14 is 3,655,731 bp long and contains 3,355 genes with 3,303 coding for proteins and 52 coding for RNA genes. The mol% G + C of the DNA is 65.7. In the genome single copies of the dissimilatory, adenylylsulfate reductase (AprBA) and the dissimilatory sulfite reduction (DsrAB) were found. The aprBA gene is similar to gram-positive aprBA genes and the Qmo complex is incomplete and consists only out of QmoAB. The QmoC is lacking which is also similar to gram-positive sulfate-reducing (e.g., Desulfotomaculum and Desulfosporosinus).
Phages
Phages might be present, but are not reported so far.
Phenotypic Analyses
The main features of the only member of the family Desulfarculaceae is listed in Table 3.1 .
All members are gram-negative vibrio. The only species is motile. All are strictly anaerobic, having a respiratory or fermentative metabolism.
The cellular fatty acid pattern of D. baarsii strain 2st14T is dominated by saturated straight chain fatty acids (43.0 % C 14:0, 9.9 % C 16:0, and 2.3 % C 18:0 ), followed by saturated iso- and anteiso-branched fatty acids (21.3 % i-C 14:0, 12.3 % ai-C 15:0, and 2.8 % i-C 15:0). A comparison of the fatty acid profiles of D. baarsii and various gram-negative sulfate reducers by cluster analysis indicated a separate position of D. baarsii (Vainshtein et al. 1992) [31], corroborating the distinct phylogenetic position of the species as shown based on the 16S rRNA sequence analysis (Fig. 3.1 ).
The dominant respiratory quinone is MK-7(H2) (Collins and Widdel 1986). Organic substrate oxidation is always complete. Typical electron donors for sulfate reduction are formate, short-chain and long-chain fatty acid, and short-chain branched fatty acids. Fermentation was never observed. Formate allows chemolithoautotrophic growth without acetate. Typical electron acceptors for growth are sulfate, sulfite, and thiosulfate.
Desulfoviridin is not present. The optimal growth is between 35 °C and 39 °C; NaCl is not required for growth.
Strains belonging to this genus might be isolated from various anoxic sediments.
Desulfarculus Kuever et al. 2006, 1VP (Effective Publication Kuever et al. 2005a, 1004)
De.sul.far’cu.lus. L. pref. de, from; L. n. sulfur, sulfur; N.L. masc. n. arculus, a small bow; N.L. masc. n. Desulfarculus, a bow-shaped sulfate-reducer.
The genus Desulfarculus contains a single validly described species. Its chemotaxonomic and physiological properties are listed in Table 3.1 .
In D. baarsii, a high activity of carbon monoxide dehydrogenase is observed, indicating the operation of the anaerobic C1-pathway (Wood-Ljungdahl pathway) for formate assimilation and CO2 fixation or complete oxidation of acetyl-CoA (Schauder et al. 1986). This is also reflected from genome data (Sun et al. 2010).
Taxonomic comment: The phylogenetic tree (Fig. 3.1 ) clearly indicates that Desulfarculus baarsii represents an isolated and deeply branching lineage. Therefore, the classification as a separate order and family is justified. This is also reflected by phylogenetic trees based on the dsrAB and aprBA genes (Zverlov et al. 2005; Meyer and Kuever 2007).
Biolog
Biolog date are not available for sulfate-reducing bacteria.
Isolation, Enrichment, and Maintenance Procedures
The only described member of the family Desulfarculaceae requires anoxic media for growth like all other sulfate-reducing bacteria. The media are prepared under specific conditions, and the addition of a reductant is required, in general sulfide to keep the medium oxygen-free. A detailed description is provided by Widdel and Bak (1992). A detailed description is provided in the genus description (Kuever et al. 2005c).
For enrichment the used electron donor should be highly selective and would have a strong influence on what kind of sulfate-reducing bacteria will grow in the medium. The most selective substrates for the isolation of Desulfarculus spp. might be long-chain fatty acids and branched short-chain fatty acids. One might succeed also with formate in the absence of acetate. As usual electron acceptor sulfate is used. All these substrates might enrich also for other sulfate-reducing bacteria belonging to other genera because there are several which have quite similar nutritional requirements.
Most sulfate-reducing bacteria of this family have been enriched using batch cultures; other options are serial dilution techniques of natural samples. For isolation in general, roll-tube techniques or deep agar serial dilution techniques are favored against plating techniques in combination with anoxic chambers (Widdel and Bak 1992; Kuever et al. 2005d).
For short-term preservation, stock cultures can be stored at 2–6 °C for 4–6 weeks. The transfer interval varies from strain to strain and depends on the tendency to lyse under suboptimal conditions. For long-term storage, cultures can be kept freeze dried, at −80 °C, or in liquid nitrogen.
Ecology
Habitat
Members of this family might be isolated from various habitats including freshwater, brackish, and marine systems. There are several clone sequences based on the 16S rRNA gene described which would fell into this family. So members of this family might be widespread, although isolates are lacking. The type strain of the genus Desulfarculus, strain 2st14, was isolated with stearate as only electron donor and sulfate as electron acceptor from anoxic sediment of a freshwater ditch close to the University of Konstanz, Germany.
Pathogenicity, Clinical Relevance
There is no pathogenic relevance known.
References
Collins MD, Widdel F (1986) Respiratory quinones of sulphate-reducing and sulphur-reducing bacteria: a systematic investigation. Syst Appl Microbiol 8:8–18
Kuever J, Rainey FA, Widdel F (2005a) Order IV. Desulfarcales ord. nov. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology, 2nd edn, vol 2, (The Proteobacteria), part C, (The Alpha-, Beta-, Delta-, and Epsilonproteobacteria). Springer, New York, p 1003
Kuever J, Rainey FA, Widdel F (2005b) Family I. Desulfarculaceae fam. nov. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology, 2nd edn, vol 2, (The Proteobacteria), part C, (The Alpha-, Beta-, Delta-, and Epsilonproteobacteria). Springer, New York, p 1003
Kuever J, Rainey FA, Widdel F (2005c) Genus I. Desulfarculus gen. nov. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology, 2nd edn, vol 2, (The Proteobacteria), part C, (The Alpha-, Beta-, Delta-, and Epsilonproteobacteria). Springer, New York, pp 1004–1005
Kuever J, Rainey FA, Widdel F (2005d) Genus 1. Desulfobacter. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology, 2nd edn, vol 2, (The Proteobacteria), part C, (The Alpha-, Beta-, Delta-, and Epsilonproteobacteria). Springer, New York, pp 961–964
Meyer B, Kuever J (2007) Pyhlogeny of the alpha and beta subunits of the dissimilatory adenosine-5´-phosphosulfate (APS) reductase from sulfate-reducing prokaryotes –origin and evolution of the dissimilatory sulfate-reduction pathway. Microbiology 153:2026–2044
Schauder R, Eikmanns B, Thauer RK, Widdel F, Fuchs G (1986) Acetate oxidation to CO2 in anaerobic bacteria via a novel pathway not involving reactions of the citric acid cycle. Arch Microbiol 145:162–172
Sun H, Spring S, Lapidus A, Davenport K, Del Rio TG, Tice H et al (2010) Complete genome sequence of Desulfarculus baarsii type strain (2st14 T ). Stand Genomic Sci 3:276–284
Vainshtein M, Hippe H, Kroppenstedt RM (1992) Cellular fatty acid composition of Desulfovibrio species and its use in classification of sulfate-reducing bacteria. Syst Appl Microbiol 15:554–566
VALIDATION LIST NO 107. 2006. Int J System Evol Microbiol 56, 1–6
Widdel F (1980) Anaerober Abbau von Fettsäuren und Benzoesäure durch neu isolierte Arten sulfat-reduzierender Bakterien. PhD thesis. Georg-August-Universität zu Göttingen. Lindhorst/Schaumburg-Lippe, Göttingen
Widdel F, Bak F (1992) Gram-negative mesophilic sulfate-reducing bacteria. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer KH (eds) The prokaryotes. Springer, New York, pp 3352–3378, Chapter 183
Yarza P, Ludwig W, Euzeby J, Amann R, Schleifer K-H, Glöckner FO, Rossello-Mora R (2010) Update of the all-species living tree project based on 16S and 23S rRNA sequence analyses. Syst Appl Microbiol 33:291–299
Zverlov V, Klein M, Lücker S, Friedrich MW, Kellermann J, Stahl DA, Loy A, Wagner M (2005) Gene transfer of dissimilatory (bi)sulfite reductase revisited. J Bacteriol 187:2203–2208
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this entry
Cite this entry
Kuever, J. (2014). The Family Desulfarculaceae. In: Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F. (eds) The Prokaryotes. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-39044-9_270
Download citation
DOI: https://doi.org/10.1007/978-3-642-39044-9_270
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-39043-2
Online ISBN: 978-3-642-39044-9
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences