Abstract
The family Halomonadaceae, within the class Gammaproteobacteria, consists mostly of marine and moderately halophilic microorganisms that are phenotypically rather diverse. As of January 2014, this family contains ten genera and 106 validly published species names and, therefore, constitutes the largest group of halophilic bacteria. In this chapter, the historical and current taxonomy have been reviewed along with molecular and phenotypic analyses. In addition, isolation and preservation procedures were considered, as well as the ecological habitats where the members of the family Halomonadaceae can be found and their clinical relevance as human pathogens. The increasing interest of this group of microorganisms due to its biotechnological and environmental applications has also been addressed.
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
The family Halomonadaceae, within the class Gammaproteobacteria, consists mostly of marine and moderately halophilic microorganisms that are phenotypically rather diverse. Because of this apparent lack of a core of differential phenotypic traits, many of its current species were previously assigned to other genera such as Deleya (now extinct), Alcaligenes, Pseudomonas, Halovibrio, Volcaniella, etc. Reorganizations among these species started by the mid 1990s with the aid of 16S rRNA gene sequence comparison. In the meanwhile, new genera and species descriptions within the family Halomonadaceae have been reported, and the increasing number of species led some authors to review its phylogeny (Arahal et al. 2002c; de la Haba et al. 2010a, 2012) and phenotypic features (Mata et al. 2002).
A Subcommittee on the Taxonomy of the Halomonadaceae, a member of the International Committee on Systematic of Prokaryotes, was constituted more than 10 years ago (Vreeland and Ventosa 2003) and can be taken as a sign of the increasing interest in this group of organisms.
Species of the genera Halomonas and Chromohalobacter have been largely studied as model organisms of halophilism. Some of their representatives are among the most halophilic bacteria (Ventosa et al. 1998) and are adapted to a wide range of saline concentrations, even wider than extreme halophiles. Another source of interest for the study of this group of organisms has been their potential in biotechnological applications. These include the production of compatible solutes, extracellular enzymes (adapted to saline stress), and exopolysaccharides among others.
Taxonomy, Historical and Current
Short Description of the Family
Ha.lo.mo.na.da’ce.ae. M.L. fem. n. Halomonas, type genus of the family; suff. -aceae, ending to denote a family; M.L. fem. pl. n. Halomonadaceae, the Halomonas family. The description of the family is identical to that given by Franzmann et al. (1988) and emended by Dobson and Franzmann (1996), Ntougias et al. (2007) and Ben Ali Gam et al. (2007).
The family Halomonadaceae belongs, together with the Alcanivoraceae, the Hahellaceae, the Litoricolaceae, the Oceanospirillaceae, the Oleiphilaceae and the “Saccharospirillaceae” to the order Oceanospirillales, within the class Gammaproteobacteria (Garrity et al. 2005a) that consists mainly of marine species.
The family Halomonadaceae was originally proposed by Franzmann et al. (1988) and it was later emended by Dobson and Franzmann (1996), Ntougias et al. (2007) and Ben Ali Gam et al. (2007). At the start of 2014, this family included ten recognized genera: Halomonas (type genus), Aidingimonas, Carnimonas, Chromohalobacter, Cobetia, Halotalea, Kushneria, Modicisalibacter, Salinicola, and Zymobacter (Parte 2014). Table 17.1 contains relevant taxonomic information on these genera and their species.
As mentioned before, some of the current species were isolated and described many years before the proposal of the genera Halomonas (Vreeland et al. 1980), Chromohalobacter (Ventosa et al. 1989) or Cobetia (Arahal et al. 2002a): Pseudomonas beijerinckii (basonym of Chromohalobacter beijerinckii), “Chromobacterium marismortui” (now Chromohalobacter marismortui), “Arthrobacter marinus” (earlier synonym of Cobetia marina), “Achromobacter aquamarinus” (Halomonas aquamarina), Flavobacterium halmophilum (basonym of Halomonas halmophila) and “Micrococcus halodenitrificans” (Halomonas halodenitrificans) are the oldest examples. In 1972, Baumann and coworkers published an extensive taxonomic study of Gram-negative, nonfermentative marine bacteria, including four organisms assigned at that time to the genus Alcaligenes, namely Alc. aestus, Alc. cupidus, Alc. pacificus and Alc. venustus (Baumann et al. 1972). About one decade later, Baumann et al. (1983) proposed the creation of the genus Deleya to accommodate those four marine species as well as Pseudomonas marina.
The genera Halomonas and Deleya served as the basis for the creation of the family Halomonadaceae (Franzmann et al. 1988). At that time these genera contained four and six species, respectively. A chemotaxonomic study (Franzmann and Tindall 1990) of members of the family Halomonadaceae concluded that on the basis of respiratory quinone, polar lipid, and fatty acid compositions, no clear distinction existed at the genus level. Additionally, it was concluded that Alcaligenes aquamarinus (currently Halomonas aquamarina) and Halovibrio variabilis (Halomonas variabilis) were members of the family Halomonadaceae and could perhaps be accomodated within existing genera of the family.
Only a few months earlier Ventosa et al. (1989) proposed Chromohalobacter as a new genus with a single species, C. marismortui, on the basis of a subculture of “Chromobacterium marismortui,” isolated from the Dead Sea (Elazari-Volcani 1940), and seven moderately halophilic isolates from a Mediterranean saltern in Spain that were found to be very closely related to it. Later, in the phylogenetic study of Mellado et al. (1995b) it was concluded that this genus belongs to the family Halomonadaceae.
The genus Zymobacter, with its single species Z. palmae, was created by Okamoto et al. (1993) and placed later in the family Halomonadaceae (Dobson and Franzmann 1996).
By then, 16S rRNA phylogenetic analyses were used as definitive evidence of the lack of correlation in the taxonomic arrangements within the family Halomonadaceae (Dobson et al. 1993). Mellado et al. (1995b) proposed the reclassification of Volcaniella eurihalina as Halomonas eurihalina and pointed out the heterogeneity of the Halomonas-Deleya complex. Dobson and Franzmann (1996) transferred all species of the genus Deleya to the genus Halomonas together with Halovibrio variabilis and Paracoccus halodenitrificans. In one way, this simplification stopped the confusion of the naming within the Halomonas-Deleya complex, but the resulting genus, Halomonas, contained (and still does) very different species and it is considered too heterogeneous. The genus Halomonas was expanded to 15 species, with few characters in common, while the only two other genera recognized at that time, Chromohalobacter and Zymobacter, contained one each. Meanwhile, the genus Carnimonas was created by Garriga et al. (1998) to accommodate one single species, C. nigrificans isolated from cured meat products, and later it was included into the family Halomonadaceae (Arahal et al. 2002c). The same year, the genus Cobetia was created by Arahal et al. (2002a) to accommodate the species Halomonas marina.
More recently, other five additional genera belonging to this family have been described: Halotalea (Ntougias et al. 2007), Modicisalibacter (Ben Ali Gam et al. 2007), Kushneria (Sánchez-Porro et al. 2009), Aidingimonas (Wang et al. 2009), and Salinicola (Anan’ina et al. 2007). Besides, the genera Halomonas, Chromohalobacter, and Cobetia were expanded to include new species since new descriptions were carried out. Moreover, several reclassifications have taken place among genera of the family Halomonadaceae: the species Halomonas canadensis and H. israelensis (Arahal et al. 2001a) were transferred to the genus Chromohalobacter; the species H. avicenniae, H. indalinina, and H. marisflavi to the genus Kushneria (Sánchez-Porro et al. 2009); and the species H. salaria and Chromohalobacter salarius to the genus Salinicola (de la Haba et al. 2010b). In 2013, Romanenko et al. (2013) classified H. halodurans as a later heterotypic synonym of Cobetia marina, invalidating the former species name.
As of January 2014 there are 106 validly published species names within the family Halomonadaceae, 79 belonging to the genus Halomonas, eight to Chromohalobacter, five to Cobetia, five to Kushneria, four to Salinicola, and the genera Aidingimonas, Carnimonas, Halotalea, Modicisalibacter, and Zymobacter with a single species each one. The list of articles in press of the International Journal of Systematic and Evolutionary Microbiology (http://ijs.sgmjournals.org/content/early/recent) includes the description of a new Halomonas species for which the name H. huangheensis has been proposed.
A valuable effort to address the taxonomy of the group from an entirely phenotypic point of view was the study of Mata et al. (2002). In that article, they presented a detailed phenotypic characterization of the type strains of all Halomonas species recognized at that time and the intraspecific variation of four of those species by studying 87 additional strains. The authors compared the reactions of 234 morphological, physiological, biochemical, nutritional and antimicrobial susceptibility tests. Part of the nutritional characterization was obtained by using a miniaturized (Biolog) identification system. It was the first time that such a method was employed so extensively among halomonads. In addition to the new data that were presented in their paper, some differences were observed between their results and those from the original species descriptions. Numerical analyses demonstrated the phenotypic heterogeneity of the Halomonas species (Mata et al. 2002). An important conclusion of the study of Mata et al. (2002) is that phenotypic traits can be selected according to their usefulness for distinguishing Halomonas species.
In 2007, the International Committee on Systematics of Prokaryotes–Subcommittee on the Taxonomy of the Halomonadaceae, following Recommendation 30b of the Bacteriological Code (1990 Revision) (Lapage et al. 1992), published the minimal standards for describing new taxa within this family (Arahal et al. 2007). This paper evaluates many different approaches to ensure that a rich polyphasic characterization is given and must be considered as guidelines for authors to prepare descriptions of novel taxa. Although a list of traits that are required and recommended is provided, the manuscript does not attempt to limit the characterization of new isolates to the features that are indicated in the text. Moreover, current or yet to be described species may show new and interesting characteristics not listed in the paper of Arahal et al. (2007) and such features may prove to be of taxonomic importance. More recently, the Subcommittee on the Taxonomy of Halomonadaceae, in an open meeting held in Storrs, Connecticut, USA (June 2013), agreed that the current minimal standards document are still adequate in combination with Notes on the characterization of prokaryote strains for taxonomic purposes (Tindall et al. 2010), but the following additions were suggested: (i) It was stressed that it is highly desirable to base the description of new species on more than one strain. (ii) A good database of sequences of suitable genes for multilocus sequence analysis is now available (de la Haba et al. 2012), and inclusion of multilocus sequence analysis in the minimal standards is highly desirable. (iii) Fatty acid analysis should be added as required rather than recommended in the recommended standards for describing new taxa of the family Halomonadaceae (Oren and Ventosa 2013).
Phylogenetic Structure of the Family and Its Genera
Seven main phylogenetic studies have been conducted on the Halomonadaceae. In the first one (Franzmann et al. 1988), which was the basis for the proposal of this family, the method employed was the 16S rRNA oligonucleotide cataloguing technique. Later, Dobson et al. (1993) obtained the 16S rRNA sequences of Deleya aquamarina (now Halomonas aquamarina), Deleya halophila (Halomonas halophila), Deleya marina (Cobetia marina), Halomonas elongata, Halomonas meridiana, Halovibrio variabilis (Halomonas variabilis) and Halomonas subglaciescola and analyzed them together with the sequences of Halomonas halmophila and other species belonging to the Gammaproteobacteria. They showed that the level of 16S rRNA sequence similarity among members of the family Halomonadaceae is 100–92.6 % and that the phylogenetic grouping did not correspond to the taxonomic assignment of the species analyzed, suggesting the unification into a single genus. They also proposed a number of characteristic sequence signatures of the members of the family Halomonadaceae that have been readapted in other studies (Dobson and Franzmann 1996; Arahal et al. 2002c; Ntougias et al. 2007; Ben Ali Gam et al. 2007), as new members have been described. Currently, the 16S rRNA sequence signature characteristics of the family Halomonadaceae are defined by the following positions: 484 (A or G), 486 (C or U), 640 (A or G), 660 (A), 668 (A), 669 (A), 737 (U), 738 (U), 745 (U), 776 (U), 1124 (U or G), 1297 (U), 1298 (C), 1423 (A), 1424 (C or U), 1439 (U or C), 1462 (A or G) and 1464 (C or U). However, fullsequence analyses are much more informative than signatures alone since the latter have to be redefined on the basis of present and future new species.
Mellado et al. (1995b) conducted a phylogenetic study on six new 16S rRNA sequences corresponding to Chromohalobacter marismortui (four strains), Volcaniella eurihalina (now Halomonas eurihalina), Deleya salina (Halomonas salina), and close relatives. They proposed the reclassification of Volcaniella eurihalina as Halomonas eurihalina but highlighted the need of a polyphasic approach to determine the natural taxonomic position of members of the Halomonadaceae, especially for the genus Halomonas since its heterogeneity was (and still is) too large for a single genus.
Further studies by Dobson and Franzmann (1996) determined another seven 16S rRNA sequences corresponding to the type strains of Halomonas subglaciescola, Deleya cupida (currently Halomonas cupida), Deleya pacifica (Halomonas pacifica), Deleya salina (Halomonas salina), Deleya venusta (Halomonas venusta), Halomonas halodurans (Cobetia marina), and Halomonas eurihalina. On the basis of their results, they proposed the unification of the genera Deleya, Halomonas and Halovibrio and the species Paracoccus halodenitrificans into the genus Halomonas.
Arahal et al. (2002c) evaluated the phylogenetic status of the family Halomonadaceae using 16S and 23S rRNA gene sequences. In addition to the new sequences determined in their study, 18 for the 23S rRNA and 7 for the 16S rRNA, the sequences were compared to more than 16,000 full or almost full rRNA sequences. By that time, the number of those sequences that could be ascribed to the family Halomonadaceae exceeded 70 (including many sequences from environmental clones and poorly characterized isolates). In addition, several treeing methods were used to elucidate the most stable branchings. A good agreement between the 16S rRNA- and the 23S rRNA-based trees was obtained. According to this study, the genus Halomonas was formed by two well-defined phylogenetic groups (containing five and seven species, respectively) as well as six species that could not be assigned to any of the above-mentioned groups. Group 1 comprised Halomonas elongata (type species of the genus), Halomonas eurihalina, Halomonas halmophila, Halomonas halophila, and Halomonas salina, all bearing a 98.2 % average sequence (16S rRNA or 23S rRNA) similarity. Group 2 included the species Halomonas aquamarina, Halomonas meridiana, Halomonas magadiensis, Halomonas variabilis, Halomonas venusta, Halomonas halodurans (now Cobetia marina), and Halomonas subglaciescola, and exhibited a 97.6 % mean 23S rRNA sequence similarity (97.4 % in the case of the 16S rRNA sequences). The species Halomonas pacifica, Halomonas halodenitrificans, Halomonas cupida, Halomonas desiderata, Halomonas campisalis, and Halomonas pantelleriensis, not only did not clearly fall into either of the two groups mentioned above but also shared relatively low values of sequence similarity with them or even between themselves (91.7–96.7 %; Arahal et al. 2002c). With respect to the genus Chromohalobacter, the four species described in those days within this genus formed a group closely related to Halomonas. The average rRNA sequence similarity of species of Chromohalobacter was 98.6 % (for the 23S rRNA) and 98.5 % (for the 16S rRNA). Within this group fell the sequence of Pseudomonas beijerinckii, which was later reclassified as Chromohalobacter beijerinckii (Peçonek et al. 2006). When the Chromohalobacter sequences were compared to those of other halomonads, values below 95 % (generally accepted as a good borderline for genus separation) were obtained in all cases. Similar low values were obtained for the sequence of Halomonas marina, which forms a deeper branch of the Halomonas-Chromohalobacter group. Indeed, according to this and other data, this organism was proposed as the type species of the new genus Cobetia (Arahal et al. 2002a). Finally, in this study, the sequences of Zymobacter palmae and Carnimonas nigrificans showed a deeper branching in the tree. Their 16S rRNA sequence similarity was 93.5 % and even lower values were obtained when comparing any of the two with the other members of the family (Arahal et al. 2002c).
More recently, de la Haba et al. (2010a) updated the comparative analysis based on 23S and 16S rRNA gene sequences of Arahal et al. (2002c) including the 49 novel species that had been described since 2002. A total of 28 new complete 23S rRNA sequences were obtained in this study. Additionally, following the recommended minimal standards for the description of new members of the family Halomonadaceae, seven already-sequenced 16S rRNA genes of type strains were resequenced to resolve undetermined positions and to reach the established quality standards. In that sense, some suggestions were included in the paper about the recommended sequences to be used for future comparative phylogenetic analysis. In general, there was excellent agreement between the phylogenies based on both rRNA genes, but the 23S rRNA gene showed higher resolution in the differentiation of species of the family Halomonadaceae due to the slower evolutionary rate for the 16S rRNA gene. As previously reported by Arahal et al. (2002c), the genus Halomonas resulted to be not monophyletic and comprised two clearly separated phylogenetic groups that now contained larger numbers of species. Group 1, representing Halomonas sensu stricto, was formed by Halomonas elongata (the type species of the genus), H. eurihalina, H. caseinilytica, H. halmophila, H. sabkhae, H. almeriensis, H. halophila, H. salina, H. organivorans, H. koreensis, H. maura and H. nitroreducens. The mean 16S rRNA gene sequence similarity for this group was 97.8 %, whereas a lower value was obtained with the 23S rRNA gene sequences (97.0 %). Group 2, included the 16 species Halomonas aquamarina, H. meridiana, H. axialensis, H. magadiensis, H. hydrothermalis, H. alkaliphila, H. venusta, H. boliviensis, H. neptunia, H. variabilis, H. sulfidaeris, H. subterranea, H. janggokensis, H. gomseomensis, H. arcis and H. subglaciescola. This group displays mean similarities of 97.4 % and 97.5 % for the 16S and 23S rRNA gene sequences, respectively. Similarity values between groups 1 and 2 were low enough as to suggest that they could constitute two different genera, however, neither chemotaxonomic nor more general phenotypic studies have permitted their separation. The other 27 species at that time assigned to the genus Halomonas did not appear to be included clearly in either of these phylogenetic groups. One of these species, H. salaria, formed a separate cluster with the species Chromohalobacter salarius and Salinicola socius, which was confirmed by further studies proposing the transference of the first two species to the genus Salinicola (de la Haba et al. 2010b). An important finding in the paper of de la Haba et al. (2010a) is the fact that the type species of Halomonas halodurans and Cobetia marina shared 100 % sequence similarity (16S and 23S rRNA) and, according to their data, there was not sufficient evidence to determine whether they were members of the same or different species. A recent publication has demonstrated that, actually, they belong to the same genospecies (Romanenko et al. 2013). Concerning the genus Chromohalobacter, all the species described until then clustered together (the mean 16S and 23S rRNA gene sequence similarity of this group was 98.0 % and 97.8 %, respectively) with the only exception being Chromohalobacter salarius, as discussed previously (de la Haba et al. 2010a). Finally, the phylogenetic distinctness of the remaining genera at that time included in the family Halomonadaceae (Carnimonas, Cobetia, Halotalea, Kushneria, Modicisalibacter, Salinicola, and Zymobacter) was confirmed in the mentioned paper, being stable in the trees produced from all methods of analysis.
In the meanwhile, 26 new species have been proposed within the genus Halomonas, some of them belonging to the group 1 (H. beimenensis, H. sinaiensis, H. smyrnensis, and H. stenophila), others to the group 2 (H. alkaliantarctica, H. andesensis, H. cibimaris, H. hamiltonii, H. jeotgali, H. johnsoniae, H. nanhaiensis, H. olivaria, H. stevensii, H. titanicae, H. vilamensis, and H. zhanjiangensis) and others that cannot be assigned to any of these two groups (H. daqiaonensis, H. flava, H. fontilapidosi, H. ilicicola, H. qijiaojingensis, H. ramblicola, H. rifensis, H. xianhensis, H. xinjiangensis, and H. zincidurans) (Fig. 17.1 ). Besides, four novel species of the genus Cobetia, and one of the genera Kushneria and Salinicola, respectively, have been described, all of them forming a monophyletic branch with the other relatives of their respective genera (Fig. 17.1 ). Additionally, the new genus Aidingimonas, not included in the study of de la Haba et al. (2010a), clustered together with the genus Modicisalibacter (Fig. 17.1 ), but they share ≤95 % 16S rRNA similarity values among them and with respect to the other genera of the family Halomonadaceae.
Phylogenetic reconstruction of the family Halomonadaceae based on 16S rRNA and created using the neighbour-joining algorithm with the Jukes-Cantor correction. The sequence datasets and alignments were used according to the All-Species Living Tree Project (LTP) database (Yarza et al. 2010; http://www.arb-silva.de/projects/living-tree). 1,000 resampling boostrap values over 50 % are shown. Scale bar indicates estimated sequence divergence
From the phylogenetic point of view, some well-defined relationships can be observed within members of the Halomonadaceae. These groups, as defined above, are stable regardless of the methodology employed. Other relations may become better defined once more in-between sequences become available.
The most recent phylogenetic study conducted within the family Halomonadaceae was carried out by de la Haba et al. (2012). A multilocus sequence analysis (MLSA) of 52 representative species was performed for the first time in a moderately halophilic bacterial group with the purpose of investigating in detail the phylogenetic relationships of species from the family Halomonadaceae and helping clarify the current classification of this complex and dynamic family. A total of six loci were selected for the analysis, the 16S and 23S rRNA and the following four protein-encoding genes (housekeeping genes): atpA (F1-ATP synthase, α subunit), gyrB (DNA gyrase, B subunit), rpoD (RNA polymerase, β subunit) and secA (protein translocase, SecA subunit). Different nucleotide substitution models and tree-constructing algorithms were compared. The average pairwise sequence similarity values for 16S rRNA, 23S rRNA, atpA, gyrB, rpoD and secA were 94.0 %, 93.1 %, 86.8 %, 79.7 %, 79.6 % and 76.6 %, respectively, indicating that the secA gene had the highest theoretical discriminatory power, although some halomonads secA gene sequences were identical, suggesting gene flow. In any case, the six genes studied were not always sufficient to correctly assign a new strain to the genus Halomonas since there was a large overlap between the intrageneric and intergeneric sequence similiarities, that is, halomonads sequences could be more similar to those from species belonging to other genera of the family than to those from species of the genus Halomonas. This overlap was mainly due to the enormous variability within the genus Halomonas, suggesting that the genus should be divided into two or more genera. Besides, the overlap problem also lies with the huge sequence similarity of the pair Halomonas halodurans-Cobetia marina, whose taxonomic status has been recently revised by Romanenko et al. (2013) and concluded that they are member of the same species. With respect to the phylogenetic trees, the different methods produced variable results, with those generated from the maximum-likelihood and neighbour-joining algorithms being more similar than those obtained by maximum-parsimony methods. Except atpA gene, the other five genes studied showed a consistent evolutionary history (with some exceptions probably due to lateral gene transfer events, and other intrinsic and extrinsic factors, such as the size of the dataset and the taxa included); therefore atpA gene may not be useful as an individual gene phylogenetic marker within the family Halomonadaceae. The gyrB-based tree was the only that formed monophyletic branches for the different genera of the family, including the genus Halomonas. Although there were some exceptions, in general, the two groups defined within this genus by Arahal et al. (2002c) and de la Haba et al. (2010a) (group 1 or Halomonas sensu stricto and group 2) could be clearly distinguished in the phylogenetic trees for each gene. Concatenation of the six loci enhanced the phylogenetic reconstruction and optimized the taxonomic resolution by adding more informative data and minimizing the weight of recombination events. Trees resulting from the six-gene concatenation demonstrated a monophyletic and well-supported separation of the different genera, including the genus Halomonas. The only exceptions were the pairs Halomonas halodurans-Cobetia marina (mentioned above, with concatenated sequence similarity of 99.7 %) and Halomonas muralis–M. tunisiensis (with concatenated sequence similarity of 90.8 %). With regard to the intrageneric groups, the six-gene concatenations resolved Halomonas groups 1 and 2 monophyletically, demonstrating to be a very good tool for the delineation of taxonomic relationships on a broad scale, including intrageneric and intergeneric relationships, at the level of the family Halomonadaceae. In order to simplify the MLSA approach within this family, de la Haba et al. (2012) attempted to reduce the number of genes to be analyzed but retaining the resolution obtained with the six concatenated gene sequences. With this idea the general use of the individual and concatenated 16S rRNA, gyrB and rpoD genes is suggested for future taxonomic studies using MLSA within the family Halomonadaceae. This proposal has been recently endorsed and recommended within the minimal standards by the ICSP-Subcommittee on the Taxonomy of the Halomonadaceaae (Oren and Ventosa 2013). Finally, the paper of de la Haba et al. (2012) analyze the phylogeny of this family within the domain Bacteria by comparing the sequences obtained in this study with those of 445 bacterial species with sequenced genomes available from the GenBank/EMBL/DDBJ databases. The results showed that the family Halomonadaceae constituted a robust and monophyletic branch within the domain Bacteria for three of the six analysed genes, 16S rRNA, gyrB and rpoD. Besides, according to Figs. 17.1, 17.2 , and 17.3 the family is related to families Saccharospirillaceae, Litoricolaceae, Oceanospirillaceae, and Hahellaceae.
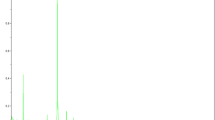
Phylogenetic distribution of the genus Halomonas based on 16S rRNA and created using the neighbor-joining algorithm with the Jukes-Cantor correction. The sequence datasets and alignments were used according to the All-Species Living Tree Project (LTP) database (Yarza et al. 2010; http://www.arb-silva.de/projects/living-tree). 1,000 resampling boostrap values over 50 % are shown. Scale bar indicates estimated sequence divergence
Phylogenetic distribution of genera within the family Halomonadaceae other than Halomonas based on 16S rRNA and created using the neighbor-joining algorithm with the Jukes-Cantor correction. The sequence datasets and alignments were usedaccording to the All-Species Living Tree Project (LTP) database (Yarza et al. 2010; http://www.arb-silva.de/projects/living-tree). 1,000 resampling boostrap values over 50 % are shown. Scale bar indicates estimated sequence divergence
Molecular Analyses
DNA-DNA Hybridization Studies
DNA–DNA hybridization data between type strains of species contained in the family Halomonadaceae are widely available. Actually, the vast majority of descriptions include results of DNA-DNA hybridization (DDH). As indicated in the recomended minimal standards for this family, DDH studies remains essential when novel species are described (Arahal et al. 2007), and only in those proposals based on a single isolate that possess less than 97 % 16S rRNA gene sequence similarity with its closest relative can DDH data be considered redundant.
With respect to DDH between strains belonging to the same species within the family Halomonadaceae data are very limited, mainly due to the fact that the majority of the species have been described based on a single isolate.
Multilocus Sequence Analysis (MLSA)
With the objective to overcome the limitations attached to DNA-DNA hybridization studies (time-consuming, expensive, lack of uniformity and reproducibility problems, etc.) and to 16S rRNA gene sequence based taxonomy (high levels of conservation, microheterogeneity among the different copies, lateral gene transfer, etc.), additional rRNA and protein-encoding genes (called housekeeping genes) have been suggested as phylogenetic markers (Stackebrandt et al. 2002; Zeigler 2003; Arahal et al. 2007; Tindall et al. 2010) to perform multilocus sequence analysis (MLSA).
Although some authors (Arahal et al. 2002c; Lee et al. 2005; de la Haba et al. 2010a; Okamoto et al. 2004; González-Domenech et al. 2010) had previously used some housekeeping genes (23S rRNA, gyrB, ectBC, narH, nirS, and nosZ) for studying the family Halomonadaceae, the first MLSA study was conducted by de la Haba et al. (2012) based on six phylogenetic markers: 16S rRNA, 23S rRNA, atpA, gyrB, rpoD, and secA genes. However, the correlation of DDH values against the MLSA data could not be determined and, therefore, this approach cannot be taken yet as an alternative to DDH for species circumscription.
DNA Fingerprinting Methods
Determination of inter- and intraspecies relatedness by rapid DNA typing methods (AFLP, RAPD, rep-PCR, BOX-PCR, PFGE, ribotyping of ribosomal ribonucleic operons, ARDRA) has not been widely applied to the Halomonadaceae except for the studies by Mellado et al. (1998) and Llamas et al. (2002), who applied PFGE, Garriga et al. (1998), who used RAPD, Heyrman et al. (2002), who reported rep-PCR, and Li et al. (2008) who carried out BOX-PCR genomic fingerprinting.
Genes Sequenced and Characterized
Several genes from different members of the family Halomonadaceae have been sequenced and characterized, mainly from Halomonas elongata (Göller et al. 1998; Grammann et al. 2002; Kraegeloh et al. 2005; Schwibbert et al. 2011) and Chromohalobacter salexigens (Cánovas et al. 1998, 2000; Copeland et al. 2011), but also from Cobetia marina (Kraiwattanapong et al. 1999), Halomonas eurihalina (Llamas et al. 2003), Halomonas halodenitrificans (Sakurai and Sakurai 1998; Sakurai et al. 2006), Halomonas maura (Llamas et al. 2006), Halomonas meridiana (Coronado et al. 2000b), Halomonas organivorans (Moreno et al. 2011), Halomonas salina (Sripo et al. 2002), and Zymobacter palmae (Raj et al. 2002).
Genome Sizes and Plasmid
Before the genomic era, estimations of the genome size of 11 Halomonas and Chromohalobacter strains were carried out by using pulsed-field gel electrophoresis (Mellado et al. 1998; Llamas et al. 2002; Quesada et al. 2004). Additionally, the presence of plasmids (and megaplasmids) in strains of Halomonas, Chromohalobacter and Cobetia has been investigated (Fernández-Castillo et al. 1992; Vargas et al. 1995; Mellado et al. 1995a; Llamas et al. 1997; Argandoña et al. 2003).
Genome Comparison
As of January 2014, complete or draft genome sequences were available for type strains of the following species: Carnimonas nigrificans (JAGO00000000), Chromohalobacter salexigens (CP000285), Halomonas anticariensis (ASTJ00000000, AUAB00000000), Halomonas boliviensis (AGQZ00000000), Halomonas elongata (FN869568), Halomonas halocynthiae (AUDZ00000000), Halomonas jeotgali (AMQY00000000), Halomonas lutea (ARKK00000000), Halomonas smyrnensis (AJKS00000000), Halomonas stevensii (AJTS00000000), Halomonas titanicae (AOPO00000000), Halomonas zhanjiangensis (ARIT00000000), and Kushneria aurantia (ARNK00000000). Although they have not been sequenced yet, the type species of Halomonas halodenitrificans, Halotalea alkalilenta, and Zymobacter palmae are part of the Genomic Encyclopedia of Type Strains, Phase I: the 1,000 microbial genomes (KMG) project.
Additional sequenced strains are: Halomonas sp. 23_GOM-1509m (JADJ01000000), Halomonas sp. A3H3 (CBRE000000000), Halomonas sp. BJGMM-B45 (AVBC00000000), Halomonas sp. GFAJ-1 (AHBC00000000), Halomonas sp. HAL1 (AGIB00000000), Halomonas sp. HTNK1 (Gi05412), Halomonas sp. KM-1 (BAEU00000000), and Halomonas sp. TD01 (AFQW00000000).
A summary of the genome characteristics of the sequenced species of the family Halomonadaceae is presented in Table 17.2 .
Phenotypic Analyses
Halomonadaceae Franzmann et al. (1989), emend. Dobson and Franzmann (1996), Ntougias et al. (2007), and Ben Ali Gam et al. (2007)
Ha.lo.mo.na.da’ce.ae. M.L. fem. n. Halomonas, type genus of the family; suff. -aceae, ending to denote a family; M.L. fem. pl. n. Halomonadaceae, the Halomonas family.
The members of the family Halomonadaceae cannot be defined by a reasonably large number of common-to-all features. This phenotypic heterogeneity of the family is also a handicap for the identification at the genus or species level unless a sufficient number of characters are determined. Cells are Gram-negative, straight or curved, rod-shaped. They are either slight or moderate halophiles or halotolerant, except species of the genus Zymobacter (Okamoto et al. 1993), growing in the presence of high concentrations of sugars. Aerobic or facultatively anaerobic, chemoorganotrophs (Dobson and Franzmann 1996; Garrity et al. 2005b). Mata et al. (2002) reviewed in depth the phenotypic features of the genus Halomonas, and confirmed the enormous diversity among the species of this genus. In addition, they reported a large number of traits not analyzed previously for all strains and found tests that are useful for distinguishing the species of the genus Halomonas.
Genotypic diversity within the Halomonadaceae is also huge, as indicated for the genomic DNA G+C content, which ranges from 52.0 to 74.3 mol% (Martínez-Cánovas et al. 2004b; Arahal and Ventosa 2006).
From the chemotaxonomic point of view, the fatty acid profile is available for most species described within this family whereas the analysis of respiratory lipoquinones or polar lipids has been addressed for only some of them. There are a set of features shared for the vast majority of the species belonging to the Halomonadaceae. The major respiratory lipoquinone is ubiquinone 9 (Q9), although ubiquinone 8 (Q8) and ubiquinone 10 (Q10) are also present in several species. With regards to polar lipid composition all the members possess phosphatidylethanolamine and phosphatidylglycerol (Franzmann and Tindall 1990), except for the species Aidingimonas halophila, which contains diphosphatidylglycerol instead of phosphatidylglycerol (Wang et al. 2009). The fatty acid profile notably varies depending on the growing media and conditions, but in general, the main fatty acids are C16:0, C18:1ω7c, C16:1ω7c, C19:0 cyclo ω8c, C17:0 cyclo, and C12:0 3-OH.
Table 17.3 shows the phenotypic and chemotaxonomic features, as well as the DNA G+C content comparison among the genera of the family Halomonadaceae. The type genus of the family is Halomonas (Vreeland et al. 1980).
Halomonas Vreeland et al. (1980), emend. Dobson and Franzmann (1996)
Ha.lo.mo’nas. Gr. n. hals, halos salt of the sea; Gr. n. monas a unit, monad; M.L. fem. n. Halomonas salt(-tolerant) monad.
Gram-negative, straight or curved, rod-shaped cells, generally 0.6–0.8 × 1.6–1.9 μm, except the species H. halodenitrificans that presents coccoid cells. Some species may produce poly-β-hydroxyalcanoates and/or exopolysaccharides. Endospores are not formed. Motile by means of peritrichous, lateral or polar flagella or nonmotile. Colonies are white to yellow, turning light brown with age. Slight to moderate halophiles, that are able to grow in NaCl concentrations ranging from 0.1 % to 32.5 % (w/v). Possess a mainly respiratory type of metabolism with oxygen as the terminal electron acceptor, but some species are also capable of anaerobic growth in the presence of nitrate. Some species have been reported to grow under anaerobic conditions in the absence of nitrate if supplied with glucose (but not other carbohydrates or amino acids). Some species reduce nitrate to nitrite; nitrogen gas is not formed. Catalase positive and most of them are also oxidase positive. Chemoorganotrophic. Carbohydrates, organic acids, polyols, and amino acids, can be used as sole carbon and energy sources or as sole carbon, nitrogen and energy sources (Vreeland et al. 1980; Dobson and Franzmann 1996; Vreeland 2005).
The major respiratory lipoquinone is ubiquinone 9 (Vreeland 2005), with the exception of H. alkaliphila that mainly possess ubiquinone 8 and ubiquinone 6 (Romano et al. 2006). The major fatty acids are C16:1 ω7c, C17:0 cyclo, C16:0, C18:1 ω7c, and C19:0 cyclo ω8c (Vreeland 2005). The predominant polar lipids are phosphatidylethanolamine and phosphatidylglycerol.
DNA G+C content ranges between 52.0 and 74.3 mol% (Martínez-Cánovas et al. 2004b; Arahal and Ventosa 2006), demonstrating the enormous heterogeneity within this genus.
Halomonas is the type genus of the family Halomonadaceae. The species Halomonas elongata is the type species of the genus.
Aidingimonas Wang et al. (2009)
Ai.ding.i.mo’nas. N.L. n. Aiding a lake located in Xinjiang province of north-west China; L. fem. n. monas, monad a unit, a monad; N.L. fem. n. Aidingimonas a monad from Aiding Lake.
Cells are Gram-negative, facultatively anaerobic, non-endospore-forming, short rods. Non-motile, without flagella. Moderately halophilic. Positive for catalase activity. Negative for oxidase activity and nitrate reduction. Ubiquinone 9 is present. Major fatty acids are C19:0 cyclo ω8c and C16 : 0. The polar lipid pattern consists of diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylinositol, phosphatidylinositol mannosides, two unknown phospholipids, two unknown phosphoglycolipids and one unknown glycolipid. The DNA G+C content is 57.2–57.5 mol% (Wang et al. 2009).
The only species within this genus is Aidingimonas halophila, which was isolated from a salt lake in Xinjiang province, north-west China. Cell size ranges between 0.1–0.3 × 0.7–1.5 μm. It forms colourless to yellow brown colonies, flat and opaque with slightly irregular edges. Growth occurs at 10–45 °C, at pH 5.0–10.0 and in 1–25 % (w/v) NaCl, with optimal growth at 37 °C, pH 7.0–8.0 and 5–10 % NaCl. Does not contain poly-β-hydroxybutyrate granules or produce exopolysaccharide. Growth occurs under anoxic conditions in the presence of nitrate ion as electron acceptor. The Voges–Proskauer test is variable. Indole and H2S are not produced. Milk peptonization and coagulation and the methyl red test are negative. Gelatin, aesculin, casein, starch, and Tweens 40, 60 and 80 are not hydrolysed, but positive for hydrolysis of Tween 20 and urea. ONPG, phenylalanine deaminase and lysine and ornithine decarboxylase tests are negative, but positive for arginine dihydrolase. Citrate and other substrates can be utilized as a sole carbon or nitrogen and energy sources. Acid is produced from different carbohydrates and organic acids. The type strain is YIM 90637T, with a DNA G+C content of 57.5 mol% (Wang et al. 2009).
Carnimonas Garriga et al. (1998)
Car.ni’mo.nas. L. gen. n. carnis of meat; Gr. n. monas a unit, monad. Carnimonas a monad of meat.
Straight or slightly curved rods, 0.5–0.6 × 1.0–1.7 μm, occurring singly or in pairs. Gram-negative. Does not form endospores. Nonmotile. Oxidase and catalase positive. Aerobic, having a strictly respiratory type of metabolism with oxygen as the terminal electron acceptor. Slightly or moderate halophile. No growth occurs in the presence of more than 8 % (w/v) NaCl. Optimum temperature for growth is 28–30 °C. No growth occurs at 5 °C or 37 °C. Chemoorganotrophic. Acid, but no gas, is produced from d-glucose, d-xylose, melibiose, maltose and sucrose. β-galactosidase (ONPG) activity occurs. Forms dark spots on the surface of raw, cured meat products. The main respiratory quinone is ubiquinone-9. Main components in the polar lipid composition are diphosphatidylglycerol, phosphatidylglycerol, and phosphatidylethanolamine. Major fatty acids are C16:0, C16:1, C18:1, and C19:0 cyclo. The G+C content of the DNA is 56.0 mol% (Garriga et al. 1998; 2005).
The type and the single species within this genus is Carnimonas nigrificans. In the species description a total of nine strains, CTCBS1T to CTCBS9 were reported to be isolated from cured meat products. Colonies are non-pigmented, white, convex, shiny and circular. Aesculin and starch are hydrolysed. Gelatin, casein and DNA are not hydrolysed. Voges-Proskauer negative. Arginine dihydrolase, urease, lecithinase and phenylalanine deaminase negative. Indole is not produced. Nitrate is not reduced (Garriga et al. 1998; 2005).
Chromohalobacter Ventosa et al. (1989) emend. Arahal et al. (2001a)
Chro.mo.ha’lo.bac’ter. Gr. n. chroma color; Gr. n. halos the sea, salt; M.L. n. bacter rod; M.L. masc. n.Chromohalobacter colored salt rod.
Gram-negative, straight or sometimes slightly curved, rods (0.4–1.2 × 0.8–6.1 μm). Motile by polar or peritrichous flagella. Cells occur singly, in pairs, and in short chains. Colonies are cream to brown-yellow pigmented, with the exception of C. nigrandesensis that shows black pigmentation. Endospores are not formed. Moderately halophilic. Salt is required for growth. The optimum salt concentration for growth is between 8 % and 10 %. May grow at salt concentrations up to 30 %. The broader ranges of temperature and pH observed for growth are 0–45 °C (optimal 30–37 °C) and pH 5.0–10.0 (optimal pH 7.5), respectively. Aerobic. Chemoorganotrophic. Catalase positive. Oxidase negative, with the exception of C. beijerinckii (Peçonek et al. 2006) and C. sarecensis (Quillaguamán et al. 2004a). Some strains reduce nitrates, but H2S and urease are not produced, with the exception of C. nigrandesensis (Prado et al. 2006) and C. salexigens (Arahal et al. 2001b). Phenylalanine deaminase test is negative. Starch, Tween 80, aesculin, DNA, and tyrosine are not hydrolyzed. The species C. japonicus is the only able to hydrolyse gelatin (Sánchez-Porro et al. 2007). Acid is produced aerobically from d-glucose and other carbohydrates. Carbohydrates, amino acids, and some polyols can serve as sole carbon or nitrogen sources (Arahal et al. 2001a; Ventosa 2005).
Predominant fatty acids are C16:0, C19:0 cyclo ω8c, and C18:1 ω7c; C17:0 cyclo, and C12:0 3-OH are present in smaller amounts. Diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine and two unknown phospholipids are major to moderate compounds in the polar lipid profile. The quinone system consists of the major compound Q-9 (>95 %) and small amounts of Q-8 (Peçonek et al. 2006; Sánchez-Porro et al. 2007). The DNA G+C base composition ranges from 56.1 to 66.0 mol%.
The type species of the genus is Chromohalobacter marismortui, previously named as “Chromobacterium marismortui”, which was originally isolated from the Dead Sea (Ventosa et al. 1989).
Cobetia Arahal et al. (2002b) emend. Romanenko et al. (2013)
Co.be’ti.a. N.L. fem. n. Cobetia named after A. B. Cobet, who originally described the type species as “Arthrobacter marinus”.
Gram-negative, straight, rod-shaped cells that are 1.6–4.0 × 0.8–1.2 μm, and occur singly and in pairs. Strains are non-motile or motile by means of a single polar flagellum and/or two to seven lateral flagella. Some strains can produce fimbria-like structures and capsules. Colonies are round, bright, smooth and cream pigmented. Poly-β-hydroxyalkanoate is accumulated. Oxidase negative. Aerobic; unable to grow anaerobically in the presence of nitrate or arginine. Sodium ions are not essential for growth. Most strains can grow without addition of NaCl to the medium, but optimal growth occurs in the presence of 5–6 % (w/v) salts and they can considered as slightly halophiles. Good growth is also obtained up to 20 % (w/v) but not at higher salinities. Hydrolyses tyrosine, but not gelatine, starch or Tween 80. Hydrolysis of casein, DNA and aesculin is strain-dependent (negative reaction for most strains). Negative for chitin hydrolysis and Simmons’ citrate test. ONPG positive. Phosphatase positive. Negative for phenylalanine deaminase, methyl red, indole production, and nitrate reduction. H2S is not produced. Acid is produced from glucose and other carbohydrates (Arahal et al. 2002a; Romanenko et al. 2013).
Polar lipids include phosphatidylethanolamine, phosphatidylglycerol, unknown phospholipids, unknown lipids, an unknown aminolipid and phosphatidic acid. The major fatty acids are C16:1 ω7c, C16:0, C12:0 3-OH, C18:1 ω7c and C17:0 cyclo. The DNA G+C content varies between 61.4 and to 64.2 mol% (Kim et al. 2010b; Romanenko et al. 2013).
The species Cobetia marina is the type of the genus and strain DSM 4741T is the type strain of this species. Strain DSM 5160, formerly the type strain of the species Halomonas halodurans, has been proposed as member of C. marina based on phylogenetic analysis and DNA-DNA hybridization and chemotaxonomic characteristics (Romanenko et al. 2013).
Halotalea Ntougias et al. (2007)
Ha.lo.ta.le’a. Gr. n. hals halos salt; L. fem. n. talea a staff, rod; N.L. fem. n. Halotalea rod-shaped cells living in saline conditions.
Cells are Gram-negative, rods, motile by peritrichous flagella and forming small, non-pigmented pale yellow colonies. Endospores are not formed. Strictly aerobic. Halotolerant and alkalitolerant. Sugar-tolerant. Oxidase- and catalase-positive. Chemo-organotrophic. Ubiquinone-9 is present in the respiratory chain. The major fatty acids are C18:1 ω7c, C16:0, C19:0 cyclo ω8c, C12:0 3-OH and C16:1 ω7c/iso-C15:0 2-OH. The DNA G+C content is 64.4 mol% (Ntougias et al. 2007).
The type species is Halotalea alkalilenta, which tolerates up to 15 % (w/v) NaCl, with an optimum salt concentration of 0–3 % (w/v) NaCl. Tolerates up to 45 % and 60 % w/v (+)-d-glucose and maltose, respectively. Grows at pH 5–11, with optimum at pH 7. The temperature range for growth is 5–45 °C, with an optimum temperature of 32–37 °C. Negative for arginine dihydrolase, lysine decarboxylase, ornithine decarboxylase, phenylalanine deaminase, nitrate reduction, H2S production from l-cysteine, indole production, methyl red and Voges-Proskauer. Acid is produced from (+)-d-glucose and other carbohydrates. Hydrolyses Tween 20, but does not hydrolyse casein, DNA, gelatin, starch, Tween 80 or urea. The type strain is AW-7T, isolated from olive mill waste (alkaline alpeorujo) obtained from the premises of the Toplou Monastery in the region of Sitia, Crete, Greece (Ntougias et al. 2007).
Kushneria Sánchez-Porro et al. (2009)
Kush.ne’ri.a. N.L. fem. n. Kushneria from the name Kushner, honouring Dr Donn J. Kushner, a Canadian microbiologist who carried out pioneering studies on halophilic micro-organisms.
Cells are Gram-negative, motile rods (0.5–2.0 × 1.7–5.0 μm). Endospores are not formed. Strictly aerobic. Colonies are yellow-orange to cream. Moderately halophilic; Na+ is required for growth. The optimal NaCl range supporting the growth is 0.5–12 % (w/v). Mesophilic (able to grow from 4 to 42° C, optimum at 25–37° C). Growth occurs at pH 4.5–10.0 (optimally at pH 7.0–8.0). Chemo-organotrophic. Catalase-positive and oxidase-negative. Aesculin and gelatin are hydrolysed, with the exception of Kushneria sinocarnis that does not possess gelatinase activity (Zou and Wang 2010). Casein, starch, Tween 80, DNA, and tyrosine are not hydrolysed. Indole and H2S production, and Voges-Proskauer test are negative. Phosphatase is produced, but urease, arginine dihydrolase, and lysine decarboxylase are not. Except K. indalinina (Cabrera et al. 2007) and K. sinocarnis (Zou and Wang 2010), the other species do not reduce nitrate to nitrite. The major respiratory quinone is Q9. Major fatty acids are C16:0, C18:1 ω7c, C19:0 cyclo ω8c, C12:0 3-OH, and C17:0 cyclo. Polar lipids are phosphatidylglycerol, diphosphatidylglycerol, phosphatidylethanolamine and unidentified phospholipids and glycolipids. The DNA G+C content is 59.0–61.7 mol% (Yoon et al. 2001; Cabrera et al. 2007; Soto-Ramírez et al. 2007; Sánchez-Porro et al. 2009; Zou and Wang 2010).
The type species of the genus is Kushneria aurantia, with the type strain A10T isolated from the leaf surface of Avicennia germinans (black mangrove).
Modicisalibacter Ben Ali Gam et al. (2007)
Mo’di.ci.sa’li.bac’ter. L. adj. modicus moderate, limited; L. n. sal, salis salt; N.L. masc. n. bacter a rod; N.L. masc. n. Modicisalibacter a moderately halophilic rod.
Cells are Gram-negative, non-endospore-forming, motile rods. Moderately halophilic. Strictly aerobic and require Na+ for growth. Mesophilic, growing well at 15–45 °C, oxidase-negative and reduce nitrate. Predominant fatty acids are C16:0, C18:1 ω7c, C16:1 ω7c, C19:0 cyclo ω8c and C17:0. Contents of C19:0 cyclo ω8c and C17:0 differ significantly from those of other members of the Halomonadaceae. The DNA G+C content is 53.7 mol% (Ben Ali Gam et al. 2007).
The genus contains a single species, namely Modicisalibacter tunisiensis, whose cells are approximately 1.0–4.0 μm long × 0.6–1.0 μm wide. Colonies on marine agar are circular, smooth, convex and 2–3 mm in diameter after 48 h of incubation at 37 °C. Cells grow at 4–45 °C, with optimum at 37 °C. The pH range for growth is 5–10, with an optimum at pH 7.2. Growth occurs in the range 0.1–25 % (w/v) NaCl and optimally at 10 % (w/v) NaCl. Catalase reaction is positive. ONPG hydrolysis is negative. Citrate is not utilized. Urease and arginine dihydrolase are not produced. Gelatin, alginate and aesculin are not hydrolysed. H2S and indole are not produced. d-glucose, d-fructose, tryptone, peptone and Casamino acids are utilized. The type strain is LIT2T, which was isolated from a sample of oilfield-water injection collected in the Sidi Litayem area near Sfax, Tunisia (Ben Ali Gam et al. 2007).
Salinicola Anan’ina et al. (2008)
Sa.li.ni.co’.la. L. fem. pl. n. salinae salterns, salt-works; N.L. suff. -cola, derived from incola, inhabitant; N.L. masc. n. Salinicola, inhabitant of salterns.
Cells are Gram-negative, non-endospore-forming rods (0.5–1.3 × 1.0–3.2 μm). Motile by means of a single lateral/polar flagellum or by peritrichous flagella (Huo et al. 2013). Colonies are circular, smooth, convex and creamy yellow-coloured. Aerobic and chemoorganotrophic. Moderately halophilic that grows in the range of 0–30 % (w/v) NaCl with the optimum of 0.5–20 % (w/v) NaCl, at pH 4.5–10.0 (optimum pH 5.0–8.0). Mesophilic, growing in the range 4–45 °C (optimal growth temperature is 25–37 °C). Exopolysaccharides are not produced, but the species Salinicola salarius and S. socius produce poly-β-hydroxyalkanoate (de la Haba et al. 2010b). Respiration on fumarate, nitrate and nitrite is negative. Shows positive reaction in the oxidation/fermentation of d-glucose. Catalase reaction is positive, oxidase reaction is negative, except for S. salarius (Kim et al. 2007). Tween 20 is hydrolysed. Tyrosine and aesculin are not hydrolysed. β-Galactosidase is not produced. Phosphatase activity is present. For all species, except S. peritrichatus, methyl red test is positive and Voges-Proskauer is negative (de la Haba et al. 2010b; Huo et al. 2013). Indole is not produced. Lysine- and ornithine-decarboxylases and phenylalanine deaminase are negative. Gluconate is not oxidized. Selenite is not reduced. The predominant respiratory lipoquinone is ubiquinone with nine isoprene units (Q9). The major fatty acids are C16:1 ω7c, C16:0, C18:1 ω7c, C19:0 cyclo ω8c, and C12:0 3-OH. The DNA G+C content ranges between 58.8 and 63.6 mol% (Anan’ina et al. 2007; Kim et al. 2007; Aguilera et al. 2007; de la Haba et al. 2010b; Huo et al. 2013).
The type species of the genus is Salinicola socius, isolated from the microbial community obtained from the soil of salt mines (Berezniki, Perm region of Russia), which grows on naphthalene as a sole carbon and energy source. The other three species comprising the genera are S. halophilus (formerly Chromohalobacter salarius), S. peritrichatus, and S. salarius (basonym of Halomonas salaria). The species name S. zeshunii (Cao et al. 2013) has been effectively, but not validly published.
Zymobacter Okamoto et al. (1995)
Zy.mo.bac’ter. Gr. n. zyme leaven, ferment; M.L. n. bacter masc. equivalent of Gr. neut. n. bakterion rod; M. L. masc. n. Zymobacter the fermenting rod.
Non-endospore forming rod-shaped cells with rounded ends, 1.3–2.4 × 0.7–0.9 μm; usually single. Motile by as many as 20 peritrichous flagella that are non-sheathed. Gram-negative. Facultatively anaerobic. Chemoorganotrophic. Grow on and ferment 1 mol of glucose or hexose moiety of maltose to produce approximately 2 mol each of ethanol and CO2, with a trace amount of acids. Ferments hexoses, α-linked di- and tri-saccharides and sugar alcohols. Growth initiates at pH values of 4.7–8.1. Catalase-positive and oxidase-negative. The major cellular fatty acids are C16:0, C19:0 cyclo, C18:1 ω9c, and C12:0 3-OH. Quinone system is ubiquinone-9. The DNA G+C base composition ranges from 55.4 to 56.2 mol% (Okamoto et al. 1993, 2005).
Currently, the genus include a single species, Zymobacter palmae, with the type strain T109T isolated from palm sap. Colonies are round, entire, smooth, opaque, and milky white. Colonies of similar size develop under aerobic and anaerobic growth conditions. Growth is better in static cultures than in shaken cultures. Requires nicotinic acid for growth. Growth does not occur in the absence of a sugar or sugar alcohol. Growth occurs at 15–37 °C (optimum, 30 °C) and pH 4.7–8.1 (optimum, pH 6.0). The organisms are neither halophilic nor halotolerant, but they are considerably tolerant to ethanol and produce ethanol mainly from maltose (5.8 % ethanol after 6 days of fermentation), but also from glucose, fructose, sucrose, melibiose, raffinose, sorbitol, mannitol, and –depending on the strain– mannose and galactose. Methyl red, Voges-Proskauer and α-glucosidase are positive. The following tests are negative: indole production, utilization of citrate, nitrate reduction, chromogenicity, gelatin liquefaction, hydrolysis of starch, phenylalanine deaminase, arginine dihydrolase, lysine decarboxylase, ornithine decarboxylase and β-galactosidase (Okamoto et al. 1993, 2005).
Isolation and Maintenance Procedures
Isolation
Halomonads may be isolated following standard microbiological techniques on complex defined media with a suitable salinity and incubated at room temperature for 1–7 days.
Since not all species show the same requirements (or tolerance) of salinity (Table 17.4 ), differences are expected to occur depending on the final salt content of the media. With the only exception of Zymobacter palmae, all species are able to grow at salinities in the range of 5–10 % but not necessarily at their optimum over the whole range. Media for the selective isolation of moderately halophilic bacteria can be prepared with higher salt contents (for instance 20 %) to prevent the growth of non-adapted competitors. The selectivity of such media can be increased by lowering the concentration of Mg2+ to prevent the growth of extremely halophilic archaea (Ventosa et al. 1982). However, not all the species within the family Halomonadaceae are able to grow at such high salinities.
The growth of species of Aidingimonas, Chromohalobacter, some Halomonas (such as H. alkaliantarctica, H. alkaliphila, H. almeriensis, H. anticariensis, H. aquamarina, H. beimenensis, H. campaniensis, H. cerina, H. cibimaris, H. cupida, H. daqiaonensis, H. daqingensis, H. denitrificans, H. eurihalina, H. fontilapidosi, H. gomseomensis, H. gudaonensis, H. halmophila, H. halodenitrificans, H. halophila, H. ilicicola, H. janggokensis, H. jeotgali, H. korlensis, H. lutea, H. maura, H. olivaria, H. organivorans, H. pantelleriensis, H. sabkhae, H. shengliensis, H. sinaiensis, H. smyrnensis, H. stenophila, H. subglaciescola, H. taeanensis, H. variabilis, H. ventosae, H. vilamensis, and H. xinjiangensis), and the species Kushneria aurantia, K. indalinina, K. sinocarnis, Modicisalibacter tunisiensis, Salinicola halophilus, and S. salarius will be favored in media with a salt content of 7.5–10 %. Some of them will yield reasonable growth at 15 % salts or even at 20 %. At these concentrations, growth of extremely halophilic Archaea can occur, but they are easily distinguished by their red pigmentation. Most of the above-mentioned species have a minimum requirement for NaCl of 0.5–5 %.
Other species (Cobetia amphilecti, Cob. crustatorum, Cob. litoralis, Cob. marina, Cob. pacifica, Halomonas alimentaria, H. boliviensis, H. campisalis, H. elongata, H. flava, H. hamiltonii, H. hydrothermalis, H. koreensis, H. kribbensis, H. mongoliensis, H. muralis, H. nitroreducens, H. qijiaojingensis, H. ramblicola, H. rifensis, H. saccharevitans, H. salina, H. stevensii, H. xianhensis, H. zincidurans, K. avicenniae, K. marisflavi, and Salinicola socius) are also moderately halophilic, but their optimum salinity for growth is lower (around 5 %). Most of these species show no requirement of salts or require only as little as 0.5–1 %.
The slightly halophilic species (Halomonas andesensis, H. arcis, H. axialensis, H. caseinilytica, H. desiderata, H. halocynthiae, H. johnsoniae, H. kenyensis, H. magadiensis, H. meridiana, H. nanhaiensis, H. neptunia, H. pacifica, H. salifodinae, H. subterranea, H. sulfidaeris, H. titanicae, H. venusta, H. zhanjiangensis, Halotalea alkalilenta and S. peritrichatus) grow optimally at around 3 % NaCl. However, these species are halotolerant of up to 15–20 % NaCl (25 % in the case of H. neptunia and H. titanicae and 10 % in the case of H. kenyensis).
Carnimonas nigrificans shows only a limited level of halotolerance. For its isolation, two selective media have been proposed (Garriga et al. 1998), cetrimide agar and MacConkey agar, although it may grow in other media such as tryptone soy agar. Although the organism is not pigmented, it produces black spots on the surface of cured meat products (the isolation source). This coloration seems to be the consequence of nonenzymatic reactions of substances derived from the meat surface.
On the contrary, Zymobacter palmae is neither halophilic nor halotolerant, but other special features of this organism can be used for its isolation. Thus, Okamoto et al. (1993) first selected ethanol-tolerant bacteria using media with 5 % (v/v) ethanol and then tested isolates for production of ethanol from maltose. Zymobacter palmae was found to produce ethanol from the fermentation of a variety of sugars: hexoses, α-linked di- and tri-saccharides, and sugar alcohols (fructose, galactose, glucose, mannose, maltose, melibiose, saccharose, raffinose, mannitol and sorbitol).
Regarding pH, most species are neutrophilic and therefore the pH of media can be adjusted to around 7.2–7.5. Exceptions to this rule are H. alkaliantarctica, H. alkaliphila, H. campaniensis, H. campisalis, H. daqingensis, H. desiderata, H. kenyensis, H. korlensis, H. magadiensis, H. mongoliensis, and H. pantelleriensis that grow optimally at pH 9.0 or 9.5 or Cobetia crustatorum, Salinicola peritrichatus, and Zymobacter palmae, with optimal growth at pH 5 or 6 (Table 17.4 ).
Finally, although all halomonads are mesophilic, some differences in their behavior towards temperature can be used for selective isolation. Aidingimonas halophila, Chromohalobacter japonicus, C. marismortui, C. salexigens, Cobetia amphilecti, C. litoralis, C. marina, C. pacifica, Halomonas alkaliphila, H. beimenensis, H. campaniensis, H. desiderata, H. elongata, H. flava, H. fontilapidosi, H. halophila, H. ilicicola, H. kenyensis, H. lutea, H. magadiensis, H. meridiana, H. mongoliensis, H. organivorans, H. qijiaojingensis, H. ramblicola, H. rifensis, H. sabkhae, H. smyrnensis, H. titanicae, H. xinjiangensis, Halotalea alkalilenta, Kushneria aurantia, K. sinocarnis, Modicisalibacter tunisiensis, Salinicola peritrichatus, and S. socius grew well at 37 °C while the rest grew only suboptimally (or not at all). Temperatures around 30 °C fit all known species so far. In the lower limit most species show no apparent growth below 4–15 °C (20 °C in the case of H. anticariensis, H. flava, H. magadiensis, H. qijiaojingensis, Kushneria aurantia, and Z. palmae; 25 °C in the case of H. ilicicola, H. rifensis, and H. sinaiensis; 30 °C in the case of H. sabkhae), but Chromohalobacter sarecensis, H. boliviensis, H. subglaciescola are capable of growing at 0 °C, and H. axialensis, H. neptunia, H. sulfidaeris at −1 °C. With respect to the upper limit, the species H. alkaliphila, H. beimenensis, H. campisalis, H. daqingensis, H. denitrificans, H. mongoliensis, H. olivaria, H. sabkhae, H. sinaiensis, H. ventosae, and H. xinjiangensis are able to grow up to 50 °C, although the most thermotolerant species is H. kenyensis, growing up to 55 °C (Table 17.4 ).
Maintenance and Preservation
A variety of media have been described for the routine maintenance of halomonads strains in the laboratory. In many cases the same media served also for the isolation of the strains, whereas in others different formulations were employed. Some of these formulations, for instance the Artificial Organic Lake (AOL) medium of Franzmann et al. (1987), are intended to mimic the chemical composition of the environment from where the organisms are isolated. Once that isolation has been achieved many authors find it advantageous, especially if preparation of the isolation medium is too laborious, to employ a more general medium that permits the growth of both fresh isolates and reference strains. Thus, one of the most commonly employed media is MH, for moderately halophilic bacteria (Ventosa et al. 1982).
MH medium (Ventosa et al. 1982) | |
Yeast extract | 10 g |
Proteose peptone | 5 g |
Glucose | 1 g |
NaCl | 81 g |
MgCl2 · 6H2O | 7 g |
MgSO4 · 7H2O | 9.6 g |
CaCl2 · 2H2O | 0.36 g |
KCl | 2 g |
NaHCO3 | 0.06 g |
NaBr | 0.026 g |
Distilled water q.s. | 1 L |
Adjust pH to 7.2 with 1 M KOH or NaOH. Add agar (20 g liter−1) for preparation of solid media. Adjust total saline content (commonly 10 %) to any other desired value by lowering or raising proportionally the amounts of salts.
In 2002 Mata et al. employed MH medium at 7.5 % salt content to maintain 104 Halomonas strains (including 21 type strains). For the vast majority of the strains the pH was adjusted to 7.2–7.5, except for two of the strains, H. magadiensis NCIMB 13595T and H. campisalis ATCC 700597T, for which the pH was adjusted to 9. All the Halomonas species described from 2002 up to now can be mantained in the same medium at pH 7.2–7.5, with the exception of H. alkaliantarctica, H. alkaliphila, H. campaniensis, H. daqingensis, H. desiderata, H. kenyensis, H. korlensis, H. mongoliensis, and H. pantelleriensis that grow optimally at pH 9.0 or 9.5.
For the species of the family Halomonadaceae with a lower salt requirement, commercial media such as Marine Agar (MA) can be satisfactorily employed (Yoon et al. 2001; Heyrman et al. 2002; Romanenko et al. 2002, 2013; Kim et al. 2010b). Alternatively, other commercial media such as trypticase soy agar (TSA) can also be employed by adding NaCl (or a mixture of salts) or even no salts (for the non-halophilic species) (Garriga et al. 1998). With regards to the species Zymobacter palmae (non-halophile) the MY medium is recommended (Okamoto et al. 1993), which consisted of 1 % Bacto yeast extract (Difco), 2 % maltose, 0.2 % KH2PO4 and 0.5 % NaCl, pH 6.0. The species Cobetia crustatorum and Salinicola peritrichatus are also slightly acidophiles (with optimal growth at pH 5 or 6), but they can be mantained in MH medium at 7.5 % salts.
Cultures on agar slants can be sealed and stored at 4–10 °C. Although viability may last for much longer periods it is recommended that they be transferred regularly every 1–3 months.
For long-term preservation, lyophilization is advised. Prior to the vacuum drying, actively growing cells can be suspended on protecting fluids such as 5 % inositol solution, and then the vials can be frozen by immersion into liquid nitrogen.
Cryopreservation at −80 °C or under liquid nitrogen is also possible. To enhance survival of the cells they have to be suspended with a cryoprotectant such as 20 % glycerol solution. Such prepared vials can also be stored at −20 °C for middle-term preservation. However, since the quality of the culture may diminish faster (especially after frequent freeze-thawing) it is recommended that new stocks be prepared regularly.
Ecology
Habitat
According to the original description of the family Halomonadaceae (Franzmann et al. 1988), its members typically occurr in “temperate and Antarctic saline lakes, solar salt facilities, saline soils and marine environments”. This is still true for the majority of the current species (Table 17.5 ). Indeed the only exceptions to the above definition are Carnimonas nigrificans, Chromohalobacter beijerinckii, Chr. canadensis, Chr. japonicus, Cobetia crustatorum, Halomonas alimentaria, H. cibimaris, H. daqingensis, H. desiderata, H. halodenitrificans, H. hamiltonii, H. jeotgali, H. johnsoniae, H. muralis, H. olivaria, H. stevensii, Halotalea alkalilenta, Kushneria sinocarnis, Modicisalibacter tunisiensis, and Zymobacter palmae. Of these 20 organisms, only Zymobacter palmae is neither halophilic nor halotolerant (however, it is very tolerant to ethanol –up to 6 %).
So, the halomonads can be found in any saline environment, regardless of its geographical location. This includes oceans and seas (even at considerable depths), saline soils, salty foods, naturally occurring saline lakes, solar pans, etc. Since some of its members are also alkaliphilic, they are found in soda lakes and alkaline soils. Additionally, three species have been isolated from blood patients and from dialysis machines of a renal care centre.
On the genus level, not surprisingly, Halomonas is found to be the most ubiquitous genus with the largest number of species, and these are very heterogeneous.
As for the interactions of halomonads with other microorganims, Ivanova et al. (2002) characterized a heterotrophic microbial enrichment community established during the degradation of brown algae Fucus evanescens, and consisting of two species, Pseudoalteromonas sp. and C. marina. While the first was highly metabolically active (14 hydrolytic activities could be detected) and likely plays the main role in the initial stages of algal degradation, the second, C. marina, produced only caseinase and DNase but was resistant to the bacteriolytic activity of the former and utilized the degradation products of polysaccharides.
In a study about the temporal stability and biodiversity of two complex antilisterial cheese-ripening microbial consortia (Maoz et al. 2003), out of 400 isolates, three were identified as H. venusta, two as H. variabilis, and two as Halomonas sp. by Fourier-transform infrared spectroscopy and 16S ribosomal RNA sequence analysis.
A recent study based on the metagenomic analysis of two hypersaline saltern ponds (19 % and 37 % NaCl) from Santa Pola (Spain) has demonstrated that the genera Halomonas and Chromohalobacter, which are commonly obtained in pure culture from similar salinity samples, are almost not represented in those ponds according to the metagenomic reads obtained (Ghai et al. 2011). This means that members of Halomonadaceae are less abundant in hypersaline environment than it was previously thought when using culture-dependent techniques.
The diversity and distribution of Halomonas populations in the hypersaline habitat Rambla Salada, situated in south-east Spain, have been studied using different molecular techniques (Oueriaghli et al. 2014). Denaturing gradient gel electrophoresis (DGGE) using specific primers for the 16S rRNA gene of Halomonas followed by a multivariate analysis of the results indicated that richness and evenness of the Halomonas populations were mainly influenced by the season, being the summer (the season with the highest salinity) the one with the highest value of diversity. Furthermore, canonical correspondence analysis (CCA) demonstrated that both salinity and pH significantly affected the structure of the Halomonas community. Halomonas almeriensis and two denitrifiers, H. ilicicola and H. ventosae were the predominant species. CARD-FISH showed that the percentage of Halomonas cells with respect to the total number of microorganisms in that habitat ranged from 4.4 % to 5.7 %. Finally, no significant differences between the types of samples studied, from either watery sediments or soil samples, were found (Oueriaghli et al. 2014). Classical cultivation methods have been also employed very recently to analyze the diversity of the halophilic bacterial community from Rambla Salada, being Halomonas the most abundant genus, representing 41.2 % of the 364 isolated strains (Luque, R., Béjar, V., Quesada, E., and Llamas, I., unpublished).
Pathogenicity, Clinical Relevance
Members of the Halomonadaceae were thought to be not pathogenic. There is one case report on the isolation of H. venusta from a human infection in a wound that originated from a fish bite (Von Graevenitz et al. 2000); however the identification of the organism alone does not prove its pathogenicity.
In 2007, Berger et al. (2007) reported an outbreak of “Halomonas phocaeensis” bacteraemia in a neonatal intensive care unit in Tunisia, attributed to contamination from a water bath used to warm fresh frozen plasma.
In a study performed seeking bacterial DNA signatures in unexplained deaths and critical illnesses, polymerase chain reaction of 16S RNA gene from culture-negative materials identified an unspeciated Halomonas organism in one patient’s blood (Nikkari et al. 2002). The phylogenetic analysis of the sequence data determined that this organism was part of the H. variabilis–H. boliviensis–H. neptunia cluster, but in an apparently distinct position (Stevens et al. 2009). Additionally, Stevens et al. (2009) isolated a total of 14 strains recognized as human pathogens causing infection and contamination in a dyalisis center. Exhaustive taxonomic characterization of these isolates led to the description of three new Halomonas species, H. stevensii, H. hamiltonii and H. johnsoniae (Kim et al. 2010a).
More recently, a patient developed a bacteremia caused by Halomonas johnsoniae (previously reported only as dialysis unit environmental contaminants) (Stevens et al. 2013). The medical community is alerted to the pathogenic potential of the genus in humans, but also in algae and animals (Kim et al. 2013).
Applications
The species of the Halomonadaceae can be used for several biotechnological purposes and, as in the case of other extremophilic microorganisms, many different applications have been suggested. The biotechnological potential and applications of moderately and halotolerant microorganisms have been reviewed in detail (Ventosa et al. 1998; Margesin and Schinner 2001; Mellado and Ventosa 2003; Quillaguamán et al. 2010; Oren 2010). Some of the most promising applications of members of Halomonadaceae include the production of compatible solutes and polyhydroxyalkanoates as well as extracellular compounds such as exopolysaccharides and enzymes, and their use in environmental bioremediation processes.
Compatible solutes are known for their stabilizing and protective effect on enzymes, nucleic acids, cell structures or whole cells subjected to low water activities, temperature stress, and other adverse conditions (Lippert and Galinski 1992; Galinski 1993; Knapp et al. 1999) and therefore could be useful for industrial and clinical purposes. A “bacterial milking” process to obtain ectoine from Halomonas elongata has been developed and patented (Sauer and Galinski 1998) and later industrially exploited by Bitop (Witten, Germany). The process is based on subjecting the bacteria repeatedly to osmotic shocks. An osmotic down-shock permits the excretion of the intracellular ectoine to the surrounding medium while subsequent exposure of the cells to a hyperosmotic shock quickly restores the original level of ectoine (Sauer and Galinski 1998). Very recently, an ectD (ectoine hydroxylase) deficient H. elongata mutant has been proved to produce ectoine from a variety of sugars derived from lignocellulosic biomass and thus has tremendous potential as a host for producing useful compounds from biomass resources (Tanimura et al. 2013). Concerning other halomonads, a process comprising two-step fed-batch cultivation has been investigated for the production of ectoine and hydroxyectoine using Halomonas boliviensis DSM 15516T (Guzmán et al. 2009; Van-Thuoc et al. 2010a, b). The first cultivation was performed under optimal conditions for cell growth and resulted in a high cell mass concentration. During the second cultivation at higher salt concentration, accumulation of ectoines increased while cell mass decreased. Maximum productivity of total ectoines reached was 10 g l−1 d−1 at 18.5 % NaCl, which is among the highest reported so far. The accumulated ectoines were released by subjecting the cells to hypoosmotic shock and the cells were further recycled for the production process (Van-Thuoc et al. 2010a). A similar method in two stages has been employed for efficient ectoine production using Halomonas salina DSM 5928T (Zhang et al. 2009; Lang et al. 2011). An ectoine absorption defective H. salina DSM 5928T mutant (lacking the ectoine-specific transporter TeaABC), which compromised the negative feedback regulation of ectoine synthesis, was constructed to improve the efficiency of ectoine production up to 9.93 g l−1 d−1 (Xu and Zhang 2012). Apart from Halomonas species, only the species Chromohalobacter salexigens has been optimized for ectoine and hydroxyectoine production (Fallet et al. 2010; Rodríguez-Moya et al. 2013). Ectoine and its derivative hydroxyectoine are used in the cosmetic industry because of their moisturizing properties. The potential use of the Chromohalobacter salexigens and Halomonas elongata ect genes (responsible for the synthesis of ectoine) to obtain agriculturally important transgenic organisms tolerant to osmotic stress has already been proposed (Vargas et al. 2004). Nakayama et al. (2000) obtained transgenic cultured tobacco cells that accumulated the compatible solute ectoine from H. elongata, exhibiting a normal growth pattern under hyperosmotic conditions.
Poly-β-hydroxyalkanoate (PHA) is a polymer accumulated by many prokaryotes, that can be used for the production of biodegradable plastics (“biological polyesters”) with properties resembling that of polypropylene. Several bacterial species of the family Halomonadaceae accumulate PHAs: Cobetia amphilecti, Cob. crustatorum, Cob. litoralis, Cob. marina, Cob. pacifica, Chromohalobacter salexigens, Chr. sarecensis, Halomonas almeriensis, H. andesensis, H. anticariensis, H. alkaliphila, H. aquamarina, H. boliviensis, H. campaniensis, H. campisalis, H. caseinilytica, H. cerina, H. cibimaris, H. cupida, H. daqiaonensis, H. daqingensis, H. desiderata, H. elongata, H. eurihalina, H. fontilapidosi, H. halmophila, H. halodenitrificans, H. halophila, H. hamiltonii, H. jeotgali, H. johnsoniae, H. magadiensis, H. maura, H. meridiana, H. nitroreducens, H. olivaria, H. pacifica, H. pantelleriensis, H. ramblicola, H. rifensis, H. salina, H. sinaiensis, H. stevensii, H. subglaciescola, H. variabilis, H. ventosae, H. venusta, H. zhanjiangensis, Kushneria marisflavi, Salinicola salarius, and S. socius. From those, the strain Halomonas boliviensis LC1T reached PHA yields and volumetric productivities close to the highest reported so far, accumulating the compound to up to 88 % of its dry weight (Quillaguamán et al. 2006, 2007). Furthermore, H. boliviensis and other Halomonas species are able to co-produce PHA and osmolytes, i.e., ectoines and hydroxyectoine, in one process (Quillaguamán et al. 2010).
Strains from several species of Halomonas are of interest as producers of exopolysaccharides (EPS). These include H. alkaliantarctica, H. alkaliphila, H. almeriensis, H. anticariensis, H. caseinilytica, H. cerina, H. daqiaonensis, H. daqingensis, H. eurihalina, H. fontilapidosi, H. halophila, H. maura, H. nitroreducens, H. olivaria, H. ramblicola, H. rifensis, H. sabkhae, H. salina, H. sinaiensis, H. smyrnensis, H. stenophila, and H. ventosae. EPS production has also been described in the species Cobetia crustatorum. The nutritional and environmental factors influencing the production of EPS have been thoroughly investigated (Quesada et al. 1993; Béjar et al. 1996; Béjar et al. 1998; Bouchotroch et al. 1999; Bouchotroch et al. 2000; Martinez-Checa et al. 2002; 2007; Arias et al. 2003; Mata et al. 2006; Llamas et al. 2012). Yield can reach 1–3 g of EPS per liter of medium after five days of cultivation. Glucose and sucrose are the most efficient carbon sources (Béjar et al. 1996) although many other nutrients can be used including end products such as molasses from sugar beet (Quesada et al. 2004). Production of the EPS is not inhibited by the presence of crude oil. Moreover, the composition of the biopolymer is different under such conditions showing a highly efficient emulsifying activity towards crude oil (Calvo et al. 2002). Interestingly, it has been observed that strains of H. maura and H. eurihalina produce more EPS at salt concentrations below 7.5 % (w/v), which are suboptimal for growth (Quesada et al. 1993; Bouchotroch et al. 2000). As for the chemical composition of the EPSs produced from Halomonas strains, they are anionic polymers composed mainly of carbohydrates and a minor fraction of proteins, uronic acids and acetyls. Sulfate substituents have also been detected (in some strains exceeding 20 % of the dry weight), and together with the uronic acids, make the EPS anionic. Depending on their composition EPSs have different functional properties and thus different applications, which include immunomodulation, biodetoxification of heavy-metal polluted environments, crude oil emulsification, viscosity enhancement in foods, protection against oxidative stress, among others (Quesada et al. 2004; Raveendran et al. 2013). Recently, it has been demonstrated that the EPS produced by H. stenophila strain B100 selectively induces apoptosis in human T leukaemia cells (Ruiz-Ruiz et al. 2011), suggesting that the search for new antineoplastic drugs should include the screening of other bacterial EPSs, particularly those isolated from halophiles. The properties of some of these EPSs such as mauran (from H. maura), H28 and V2-7 (from H. eurihalina) and those from strains Al12T and Al16 (H. ventosae), from strains FP35T and FP36 (H. anticariensis) and from strain M8T (H. almeriensis) have been studied in detail highlighting in each case their biotechnological potentials (Pérez-Fernandez et al. 2000; Martinez-Checa et al. 2002, 2007; Arias et al. 2003; Mata et al. 2006; Llamas et al. 2012). Besides, genes involved in the biosynthesis of the exopolysaccharide mauran produced by H. maura have been identified, which form part of a gene cluster epsABCDJ (Arco et al. 2005). Furthermore, production of EPS in H. anticariensis was found to be influenced by a two-component regulatory system, GacS/GacA (Tahrioui et al. 2013b).
Another interesting field is the production of extracellular enzymes (i.e., amylases, proteases and nucleases) by moderately halophilic representatives of the family. Although few studies have been carried out, this is a promising subject that has been reviewed (Mellado et al. 2004). Sanchez-Porro et al. (2003) studied the diversity of moderately halophilic bacteria able to produce several extracellular hydrolytic enzymes (amylases, DNases, lipases, proteases and pullulanases) by screening different hypersaline locations in South Spain. In contrast to the scarce hydrolytic activity shown by culture collection strains, environmental isolates produce a variety of extracellular enzymes that could be of potential biotechnological interest. Identification at the genus level revealed that 25 strains and 2 out of 122 isolates belonged to Halomonas and Chromohalobacter, respectively. Another study with the aims to isolate and characterize the cultivable community of hydrolase producers inhabiting heavy-metal-contaminated soils in extreme conditions from the Atacama Desert showed that only 1 out of 25 isolates was closely related to the genus Halomonas, in particular to H. organivorans, possessing protease, lipase, amylase, DNase, and pullulanase activities (Moreno et al. 2012). Concerning the characterization of extracellular enzymes, an α-amylase produced by Halomonas meridiana has been studied at the biochemical and molecular level (Coronado et al. 2000a, b). The amylase showed optimal activity at 10 % NaCl, 37 °C and pH 7.0 (being relatively stable under alkaline conditions). The enzyme showed activity at high salt concentrations (up to 30 %). The main products resulting from the hydrolysis of starch were maltose and maltotriose. The gene encoding this α-amylase (amyH) has been cloned and expressed in the heterologous hosts H. elongata and the non-halophilic bacterium E. coli (Coronado et al. 2000b). It encodes a 457-residue protein with a molecular mass of 50 kDa. Besides, an extracellular α-amylase gene from the hyperthermophilic archaeon Pyrococcus woesei has been cloned and expressed in H. elongata, under the control of a native H. elongata promoter (Frillingos et al. 2000). More recently, a novel endoglucanase (Cel8H) from Halomonas sp. strain S66-4 has been biochemically characterized, cloned, expressed in E. coli and purified. The purified recombinant enzyme had an optimal activity of 4.9 U/mg at pH 5 and 45° C toward the substrate carboxymethylcellulose. It exhibited extraordinary properties which differed from endoglucanases reported previously at the point of high salt tolerance above 5 M, simultaneously with high stability at pH 4–12 and 40–60° C (Huang et al. 2010). Moreover, an α-glucosidase (HaG) with a estimated molecular mass of 58 kDa was isolated from Halomonas sp. strain H11 (Ojima et al. 2012a). HaG showed high hydrolytic activities toward maltose, sucrose, and p-nitrophenyl-α-d-glucoside but to almost no other disaccharides or malto-oligosaccharides higher than trisaccharides. HaG showed optimum activity to maltose at 30 °C and pH 6.5. Monovalent cations, such as K+, Rb+, Cs+, and NH4 + increased the enzymatic activity to 2- to 9-fold of the original activity. This enzyme was tested to be useful for efficient synthesis of α-d-glucosylglycerol and α-glucosylated 6-gingerol (Ojima et al. 2012a, b).
Halomonads and other moderately halophilic bacteria can be also used for the degradation of toxic compounds both at high and intermediate salt concentrations, permitting the restoration of saline industrial residues and contaminated saline environments (Mellado and Ventosa 2003). However, few studies have been carried out. Some examples of halomonads with a potential for biodegradation and biotransformation of contaminants (such as aminomethane sulfonate, p-aminosalicylate, organophosphonates, benzoate, 3-chlorobenzoate, cinnamate, p-coumarate, crude-oil, 2,4-dichlorophenoxyacetate, ferulate, formaldehyde, p-hydroxybenzoate, phenol, phenylacetate, phenylpropionate, salicylate, selenate and uranium compounds) are shown in Table 17.6 . Maltseva et al. (1996) isolated halomonads able to use chloroaromatic compounds as the sole source of carbon and energy and studied in detail the degradation of the herbicide 2,4-dichlorophenoxyacetic acid (2,4-D). Kleinsteuber et al. (2001) reported the use of the alkaliphilic, moderately halophilic Halomonas sp. strain EF43 for the expression of the 2,4-d-degradative pathway by conjugation of the broad host range plasmid pJP4. This strain was able to degrade 2,4-d- and 3-chlorobenzoate under alkaline conditions in the presence of an additional carbon source. In 2004, García et al. (2004) described a novel species, Halomonas organivorans, which was able to use benzoic acid, p-hydroxybenzoic acid, cinnamic acid, salicylic acid, phenylacetic acid, phenylpropionic acid, phenol, p-coumaric acid, ferulic acid and p-aminosalicylic acid. Later, a screening of moderately halophilic bacteria able to use aromatic compounds permitted the isolation of a large number of members of the genus Halomonas able to degrade different compounds (García et al. 2005). Recently, cat and ben genes, involved in phenol and benzoate degradation, respectively, from H. organivorans, have been cloned, characterized and analyzed (Moreno et al. 2011), providing an ideal model system to investigate the potential use of this group of extremophiles in the decontamination of saline environments. Other recent publications also show the ability of H. organivorans to degradate contaminants, such as crude-oil (Hassanshahian et al. 2012) and phenol (Bonfá et al. 2013). The tolerance patterns of several Halomonas species and Chromohalobacter marismortui to ten heavy metals have been studied, as well as the influence of salinity and composition of culture media (Nieto et al. 1989). These studies may be interesting for future use of metal-tolerant halophilic strains as biological detoxicants.
Few studies have been carried out on the role of halomonads in the fermentation processes of foods and other products. Cobetia crustatorum, Halomonas alimentaria, H. cibimaris, and H. jeotgali were isolated from the traditional Korean fermented seafood (Kim et al. 2010b; Yoon et al. 2002; Jeong et al. 2013; Kim et al. 2011). The hydrolytic activity of H. elongata in bacon curing brines has been investigated (Hinrichsen et al. 1994).
Fuel ethanol production has been proposed using the pyruvate decarboxylase (PDC) of Z. palmae as biocatalyst (Raj et al. 2002). This enzyme is thermostable and showed the highest specific activity and lowest Km for pyruvate of all four known bacterial PDCs.
References
Aguilera M, Cabrera A, Incerti C, Fuentes S, Russell NJ, Ramos-Cormenzana A, Monteoliva-Sánchez M (2007) Chromohalobacter salarius sp. nov., a moderately halophilic bacterium isolated from a solar saltern in Cabo de Gata, Almería, southern Spain. Int J Syst Evol Microbiol 57:1234–1242
Alva VA, Peyton BM (2003) Phenol and catechol biodegradation by the haloalkaliphile Halomonas campisalis: influence of pH and salinity. Environ Sci Technol 37:4397–4402
Amjres H, Béjar V, Quesada E, Abrini J, Llamas I (2011) Halomonas rifensis sp. nov., an exopolysaccharide-producing, halophilic bacterium isolated from a solar saltern. Int J Syst Evol Microbiol 61:2600–2605
Amouric A, Liebgott PP, Joseph M, Brochier-Armanet C, Lorquin J (2014) Halomonas olivaria sp. nov., a moderately halophilic bacterium isolated from olive-processing effluents. Int J Syst Evol Microbiol 64:46–54
Anan’ina LN, Plotnikova EG, Gavrish EY, Demakov VA, Evtushenko LI (2007) Salinicola socius gen. nov., sp. nov., a moderately halophilic bacterium from a naphthalene-utilizing microbial association. Microbiology 76:324–330 (English traslation of Mikrobiologiya 76:369–376)
Anan’ina LN, Plotnikova EG, Gavrish EY, Demakov VA, Evtushenko LI (2008) Salinicola socius gen. nov. In: List of new names and new combinations previously effectively, but not validly, published List No. 124. Int J Syst Evol Microbiol 58:2471–2472
Arahal DR, Ventosa A (2006) The family Halomonadaceae. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 6, 3rd edn, Proteobacteria: gamma subclass. Springer, New York, pp 811–835
Arahal DR, Garcia MT, Ludwig W, Schleifer K-H, Ventosa A (2001a) Transfer of Halomonas canadensis and Halomonas israelensis to the genus Chromohalobacter as Chromohalobacter canadensis comb. nov. and Chromohalobacter israelensis comb. nov. Int J Syst Evol Microbiol 51:1443–1448
Arahal DR, Garcia MT, Vargas C, Canovas D, Nieto JJ, Ventosa A (2001b) Chromohalobacter salexigens sp. nov., a moderately halophilic species that includes Halomonas elongata DSM 3043 and ATCC 33174. Int J Syst Evol Microbiol 51:1457–1462
Arahal DR, Castillo AM, Ludwig W, Schleifer K-H, Ventosa A (2002a) Proposal of Cobetia marina gen. nov., comb. nov., within the family Halomonadaceae, to include the species Halomonas marina. Syst Appl Microbiol 25:207–211
Arahal DR, Castillo AM, Ludwig W, Schleifer K-H, Ventosa A (2002b) Cobetia marina gen. nov., comb. nov. In: Validation of publication of new names and new combinations previously effectively published outside the IJSEM List No. 88. Int J Syst Evol Microbiol 52:1915–1946
Arahal DR, Ludwig W, Schleifer K-H, Ventosa A (2002c) Phylogeny of the family Halomonadaceae based on 23S and 16S rDNA sequence analyses. Int J Syst Evol Microbiol 52:241–249
Arahal DR, Vreeland RH, Litchfield CD, Mormile MR, Tindall BJ, Oren A, Bejar V, Quesada E, Ventosa A (2007) Recommended minimal standards for describing new taxa of the family Halomonadaceae. Int J Syst Evol Microbiol 57:2436–2446
Arco Y, Llamas I, Martínez-Checa F, Argandoña M, Quesada E, del Moral A (2005) epsABCJ genes are involved in the biosynthesis of the exopolysaccharide mauran produced by Halomonas maura. Microbiology 151:2841–2851
Arenas M, Bañón PI, Copa-Patiño JL, Sánchez-Porro C, Ventosa A, Soliveri J (2009) Halomonas ilicicola sp. nov., a moderately halophilic bacterium isolated from a saltern. Int J Syst Evol Microbiol 59:578–582
Argandoña M, Martínez-Checa F, Llamas I, Quesada E, del Moral A (2003) Megaplasmids in Gramnegative, moderately halophilic bacteria. FEMS Microbiol Lett 227:81–86
Arias S, del Moral A, Ferrer MR, Tallón R, Quesada E, Béjar V (2003) Mauran, an exopolysaccharide produced by the halophilic bacterium Halomonas maura, with a novel composition and interesting properties for biotechnology. Extremophiles 7:319–326
Azachi M, Henis Y, Oren A, Gurevich P, Sarig S (1995) Transformation of formaldehyde by a Halomonas sp. Can J Microbiol 41:548–553
Baumann L, Baumann P, Mandel M, Allen RD (1972) Taxonomy of aerobic marine eubacteria. J Bacteriol 110:402–429
Baumann L, Bowditch RD, Baumann P (1983) Description of Deleya gen. nov. created to accommodate the marine species Alcaligenes aestus, A. pacificus, A. cupidus, A. venustus, and Pseudomonas marina. Int J Syst Bacteriol 33:793–802
Béjar V, Calvo C, Moliz J, Diaz-Martinez F, Quesada E (1996) Effect of growth conditions on the rheological properties and chemical composition of Volcaniella eurihalina exopolysaccharide. Appl Biochem Biotechnol 59:77–86
Béjar V, Llamas I, Calvo C, Quesada E (1998) Characterization of exopolysaccharides produced by 19 halophilic strains of the species Halomonas eurihalina. J Biotechnol 61:135–141
Ben Ali Gam Z, Abdelkafi S, Casalot L, Tholozan JL, Oueslati R, Labat M (2007) Modicisalibacter tunisiensis gen. nov., sp. nov., an aerobic, moderately halophilic bacterium isolated from an oilfield-water injection sample, and emended description of the family Halomonadaceae Franzmann et al. 1989 emend Dobson and Franzmann 1996 emend. Ntougias et al. 2007. Int J Syst Evol Microbiol 57:2307–2313
Berendes F, Gottschalk G, Heine-Dobbernack E, Moore ERB, Tindall BJ (1996) Halomonas desiderata sp. nov., a new alkaliphilic, halotolerant and denitrifying bacterium isolated from a municipal sewage works. Syst Appl Microbiol 19:158–167
Berger P, Barguellil F, Raoult D, Drancourt M (2007) An outbreak of Halomonas phocaeensis sp. nov. bacteraemia in a neonatal intensive care unit. J Hosp Infect 67:79–85
Boltyanskaya YV, Kevbrin VV, Lysenko AM, Kolganova TV, Tourova TP, Osipov GA, Zhilina TN (2007) Halomonas mongoliensis sp. nov. and Halomonas kenyensis sp. nov., new haloalkaliphilic denitrifiers capable of N2O reduction, isolated from soda lakes. Microbiology 76:739–747 (English traslation of Mikrobiologiya 76:834–843)
Bonfá MR, Grossman MJ, Piubeli F, Mellado E, Durrant LR (2013) Phenol degradation by halophilic bacteria isolated from hypersaline environments. Biodegradation 24:699–709
Bouchotroch S, Quesada E, del Moral A, Béjar V (1999) Taxonomic study of exopolysaccharide-producing, moderately halophilic bacteria isolated from hypersaline environments in Morocco. Syst Appl Microbiol 22:412–419
Bouchotroch S, Quesada E, Izquierdo I, Rodriguez M, Béjar V (2000) Bacterial exopolysaccharides produced by newly discovered bacteria belonging to the genus Halomonas, isolated from hypersaline habitats in Morocco. J Ind Microbiol Biotechnol 24:374–378
Bouchotroch S, Quesada E, del Moral A, Llamas I, Béjar V (2001) Halomonas maura sp. nov., a novel moderately halophilic, exopolysaccharide-producing bacterium. Int J Syst Evol Microbiol 51:1625–1632
Cabrera A, Aguilera M, Fuentes S, Incerti C, Russell NJ, Ramos-Cormenzana A, Monteoliva-Sánchez M (2007) Halomonas indalinina sp. nov., a moderately halophilic bacterium isolated from a solar saltern in Cabo de Gata, Almería, southern Spain. Int J Syst Evol Microbiol 57:376–380
Cai L, Tan D, Aibaidula G, Donq XR, Chen JC, Tian WD, Chen GQ (2011) Comparative genomics study of polyhydroxyalkanoates (PHA) and ectoine relevant genes from Halomonas sp. TD01 revealed extensive horizontal gene transfer events and co-evolutionary relationships. Microb Cell Fact 10:88
Calvo C, Martínez-Checa F, Toledo FL, Porcel J, Quesada E (2002) Characteristics of bioemulsifiers synthesised in crude oil media by Halomonas eurihalina and their effectiveness in the isolation of bacteria able to grow in the presence of hydrocarbons. Appl Microbiol Biotechnol 60:347–351
Cánovas D, Vargas C, Calderón MI, Ventosa A, Nieto JJ (1998) Characterization of the genes for the biosynthesis of the compatible solute ectoine in the moderately halophilic bacterium Halomonas elongata DSM 3043. Syst Appl Microbiol 21:487–497
Cánovas D, Vargas C, Kneip S, Morón M-J, Ventosa A, Bremer E, Nieto JJ (2000) Genes for the synthesis of the osmoprotectant glycine betaine from choline in the moderately halophilic bacterium Halomonas elongata DSM 3043. Microbiology 146:455–463
Cao L, Yan Q, Ni H, Hu G, Hong Q, Li S (2013) Salinicola zeshunii sp. nov., a moderately halophilic bacterium isolated from soil of a chicken farm. Curr Microbiol 66:192–196
Chen Y-G, Zhang Y-Q, Huang H-Y, Klenk H-P, Tang S-K, Huang K, Chen Q-H, Cui S-L, Li W-J (2009) Halomonas zhanjiangensis sp. nov., a halophilic bacterium isolated from a sea urchin. Int J Syst Evol Microbiol 59:2888–2893
Chen C, Shi R, Liu BB, Zhang YJ, Sun HZ, Li CT, Tang SK, Zhang LL, Li WJ (2011) Halomonas qijiaojingensis sp. nov. and Halomonas flava sp. nov., two moderately halophilic bacteria isolated from a salt lake. Antonie Van Leeuwenhoek 100:365–373
Copeland A, O’Connor K, Lucas S, Lapidus A, Berry KW, Detter JC, Del Rio TG, Hammon N, Dalin E, Tice H, Pitluck S, Bruce D, Goodwin L, Han C, Tapia R, Saunders E, Schmutz J, Brettin T, Larimer F, Land M, Hauser L, Vargas C, Nieto JJ, Kyrpides NC, Ivanova N, Goker M, Klenk HP, Csonka LN, Woyke T (2011) Complete genome sequence of the halophilic and highly halotolerant Chromohalobacter salexigens type strain (1H11(T)). Stand Genomic Sci 5:379–388
Coronado MJ, Vargas C, Hofemeister J, Ventosa A, Nieto JJ (2000a) Production and biochemical characterization of an alpha-amylase from the moderate halophile Halomonas meridiana. FEMS Microbiol Lett 183:67–71
Coronado MJ, Vargas C, Mellado E, Tegos G, Drainas C, Nieto JJ, Ventosa A (2000b) The α-amylase gene amyH of the moderate halophile Halomonas meridiana: Cloning and molecular characterization. Microbiology 146:861–868
de la Haba RR, Arahal DR, Márquez MC, Ventosa A (2010a) Phylogenetic relationships within the family Halomonadaceae based on 23S and 16S rRNA comparative sequence analysis. Int J Syst Evol Microbiol 60:737–748
de la Haba RR, Sánchez-Porro C, Márquez MC, Ventosa A (2010b) Taxonomic study of the genus Salinicola: transfer of Halomonas salaria and Chromohalobacter salarius to the genus Salinicola as Salinicola salarius comb. nov. and Salinicola halophilus nom. nov., respectively. Int J Syst Evol Microbiol 60:963–971
de la Haba RR, Márquez MC, Papke RT, Ventosa A (2012) Multilocus sequence analysis of the family Halomonadaceae. Int J Syst Evol Microbiol 62:520–538
De Souza MP, Amini A, Dojka MA, Pickering IJ, Dawson SC, Pace NR, Terry N (2001) Identification and characterization of bacteria in a selenium-contaminated hypersaline evaporation pond. Appl Environ Microbiol 67:3785–3794
Dobson SJ, Franzmann PD (1996) Unification of the genera Deleya (Baumann et al. 1983), Halomonas (Vreeland et al. 1980), and Halovibrio (Fendrich 1988) and the species Paracoccus halodenitrificans (Robinson and Gibbons 1952) into a single genus, Halomonas, and placement of the genus Zymobacter in the family Halomonadaceae. Int J Syst Bacteriol 46:550–558
Dobson SJ, James SR, Franzmann PD, McMeekin TA (1990) Emended description of Halomonas halmophila (NCMB 1971T). Int J Syst Bacteriol 40:462–463
Dobson SJ, McMeekin TA, Franzmann PD (1993) Phylogenetic relationships between some members of the genera Deleya, Halomonas, and Halovibrio. Int J Syst Bacteriol 43:665–673
Duckworth AW, Grant WD, Jones BE, Meijer D, Márquez MC, Ventosa A (2000) Halomonas magadii sp. nov., a new member of the genus Halomonas, isolated from a soda lake of the East African rift valley. Extremophiles 4:53–60
Elazari-Volcani B (1940) Studies on the microflora of the dead sea. Doctoral thesis, Hebrew University, Jerusalem, 1–116 and i–xiii
Fallet C, Rohe P, Franco-Lara E (2010) Process optimization of the integrated synthesis and secretion of ectoine and hydroxyectoine under hyper/hypo-osmotic stress. Biotechnol Bioeng 107:124–133
Fernández-Castillo R, Vargas C, Nieto JJ, Ventosa A, Ruiz-Berraquero F (1992) Characterization of a plasmid from moderately halophilic eubacteria. J Gen Microbiol 138:1133–1137
Francis AJ, Dodge CJ, Gillow JB, Papenguth HW (2000) Biotransformation of uranium compounds in high ionic strength brine by a halophilic bacterium under denitrifying conditions. Environ Sci Technol 34:2311–2317
Franzmann PD, Tindall BJ (1990) A chemotaxonomic study of members of the family Halomonadaceae. Syst Appl Microbiol 13:142–147
Franzmann PD, Burton HR, McMeekin TA (1987) Halomonas subglaciescola, a new species of halotolerant bacteria isolated from Antarctica. Int J Syst Bacteriol 37:27–34
Franzmann PD, Wehmeyer U, Stackerbrandt E (1988) Halomonadaceae fam. nov., a new family of the class Proteobacteria to accommodate the genera Halomonas and Deleya. Syst Appl Microbiol 11:16–19
Franzmann PD, Wehmeyer U, Stackerbrandt E (1989) Halomonadaceae fam. nov. In: Validation of the publication of new names and new combinations previously effectively published outside the IJSB List No. 29. Int J Syst Biol 39:205–206
Frillingos S, Linden A, Niehaus F, Vargas C, Nieto JJ, Ventosa A, Antranikian G, Drainas C (2000) Cloning and expression of alpha-amylase from the hyperthermophilic archaeon Pyrococcus woesei in the moderately halophilic bacterium Halomonas elongata. J Appl Microbiol 88:495–503
Galinski EA (1993) Compatible solutes of halophilic eubacteria: molecular principles, water-solute interactions, stress protection. Experientia 49:487–495
García MT, Mellado E, Ostos JC, Ventosa A (2004) Halomonas organivorans sp. nov., a moderate halophile able to degrade aromatic compounds. Int J Syst Evol Microbiol 54:1723–1728
García MT, Ventosa A, Mellado E (2005) Catabolic versatility of aromatic compound-degrading halophilic bacteria. FEMS Microbiol Ecol 54:97–109
Garriga M, Ehrmann MA, Arnau J, Hugas M, Vogel RF (1998) Carnimonas nigrificans gen. nov., sp. nov., a bacterial causative agent for black spot formation on cured meat products. Int J Syst Bacteriol 48:677–686
Garriga M, Ehrmann MA, Arnau J, Hugas M, Vogel RF (2005) Genus II. Carnimonas. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, 2nd edn. Springer, New York, pp 313–315
Garrity GM, Bell JA, Lilburn TG (2005a) Order VIII. Oceanospirillales ord. nov. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, 2nd edn. Springer, New York, p 270
Garrity GM, Bell JA, Lilburn TG (2005b) Family IV. Halomonadaceae. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, 2nd edn. Springer, New York, p 300
Ghai R, Pašić L, Fernández AB, Martin-Cuadrado AB, Mizuno CM, McMahon KD, Papke RT, Stepanauskas R, Rodriguez-Brito B, Rohwer F, Sánchez-Porro C, Ventosa A, Rodríguez-Valera F (2011) New abundant microbial groups in aquatic hypersaline environments. Sci Rep 1:135
Göller K, Ofer A, Galinski EA (1998) Construction and characterization of an NaCl-sensitive mutant of Halomonas elongata impaired in ectoine biosynthesis. FEMS Microbiol Lett 161:293–300
González-Domenech CM, Béjar V, Martínez-Checa F, Quesada E (2008a) Halomonas nitroreducens sp. nov., a novel nitrate- and nitrite-reducing species. Int J Syst Evol Microbiol 58:872–876
González-Domenech CM, Martínez-Checa F, Quesada E, Béjar V (2008b) Halomonas cerina sp. nov., a moderately halophilic, denitrifying, exopolysaccharide-producing bacterium. Int J Syst Evol Microbiol 58:803–809
González-Domenech CM, Martínez-Checa F, Quesada E, Béjar V (2009) Halomonas fontilapidosi sp. nov., a moderately halophilic, denitrifying bacterium. Int J Syst Evol Microbiol 59:1290–1296
González-Domenech CM, Martínez-Checa F, Béjar V, Quesada E (2010) Denitrification as an important taxonomic marker within the genus Halomonas. Syst Appl Microbiol 33:85–93
Grammann K, Volke A, Kunte HJ (2002) New type of osmoregulated solute transporter identified in halophilic members of the Bacteria domain: TRAP transporter TeaABC mediates uptake of ectoine and hydroxyectoine in Halomonas elongata DSM 2581 T. J Bacteriol 184:3078–3085
Guan T-W, Xiao J, Zhao K, Luo X-X, Zhang X-P, Zhang L-L (2010) Halomonas xinjiangensis sp. nov., a halotolerant bacterium isolated from a salt lake. Int J Syst Evol Microbiol 60:349–352
Guzmán H, Van-Thuoc D, Martín J, Hatti-Kaul R, Quillaguamán J (2009) A process for the production of ectoine and poly(3-hydroxybutyrate) by Halomonas boliviensis. Appl Microbiol Biotechnol 84:1069–1077
Guzmán D, Quillaguamán J, Muñoz M, Hatti-Kaul R (2010) Halomonas andesensis sp. nov., a moderate halophile isolated from the saline lake Laguna Colorada in Bolivia. Int J Syst Evol Microbiol 60:749–753
Guzmán D, Balderrama-Subieta A, Cardona-Ortuño C, Guevara-Martínez M, Callisava-Quispe N, Quillaguamán J (2012) Evolutionary patterns of carbohydrate transport and metabolism in Halomonas boliviensis as derived from its genome sequence: influences on polyester production. Aquat Biosyst 8:9
Hassanshahian M, Emtiazi G, Cappello S (2012) Isolation and characterization of crude-oil-degrading bacteria from the Persian Gulf and the Caspian Sea. Mar Pollut Bull 64:7–12
Hayes VE, Ternan NG, McMullan G (2000) Organophosphonate metabolism by a moderately halophilic bacterial isolate. FEMS Microbiol Lett 186:171–175
Heyrman J, Balcaen A, De Vos P, Swings J (2002) Halomonas muralis sp. nov., isolated from microbial biofilms colonizing the walls and murals of the Saint-Catherine chapel (Castle Herberstein, Austria). Int J Syst Evol Microbiol 52:2049–2054
Hinrichsen LL, Montel MC, Talon R (1994) Proteolytic and lipolytic activities of Micrococcus roseus (65), Halomonas elongata (16) and Vibrio sp. (168) isolated from Danish bacon curing brines. Int J Food Microbiol 22:115–126
Hinteregger C, Streichsbier F (1997) Halomonas sp., a moderately halophilic strain, for biotreatment of saline phenolic waste-water. Biotechnol Lett 19:1099–1102
Huang X, Shao Z, Hong Y, Lin L, Li C, Huang F, Wang H, Liu Z (2010) Cel8H, a novel endoglucanase from the halophilic bacterium Halomonas sp. S66-4: molecular cloning, heterogonous expression, and biochemical characterization. J Microbiol 48:318–324
Huo Y-Y, Meng F-X, Xu L, Wang C-S, Xu X-W (2013) Salinicola peritrichatus sp. nov., isolated from deep-sea sediment. Antonie Van Leeuwenhoek 104:55–62
Ivanova EP, Bakunina IY, Sawabe T, Hayashi K, Alexeeva YV, Zhukova NV, Nicolau DV, Zvaygintseva TN, Mikhailov VV (2002) Two species of culturable bacteria associated with degradation of brown algae Fucus evanescens. Microb Ecol 43:242–249
James SR, Dobson J, Franzmann PD, McMeekin TA (1990) Halomonas meridiana, a new species of extremely halotolerant bacteria isolated from Antarctic saline lakes. Syst Appl Microbiol 13:270–277
Jeon CO, Lim J-M, Lee JR, Lee GS, Park D-J, Lee J-C, Oh H-W, Kim C-J (2007) Halomonas kribbensis sp. nov., a novel moderately halophilic bacterium isolated from a solar saltern in Korea. Int J Syst Evol Microbiol 57:2194–2198
Jeong SH, Lee JH, Jung JY, Lee SH, Park MS, Jeon CO (2013) Halomonas cibimaris sp. nov., isolated from jeotgal, a traditional Korean fermented seafood. Antonie Van Leeuwenhoek 103:503–512
Kawata Y, Kawasaki K, Shigeri Y (2012) Draft genome sequence of Halomonas sp. Strain KM-1, a moderately halophilic bacterium that produces the bioplastic poly(3-hydroxybutyrate). J Bacteriol 194:2738–2739
Kaye JZ, Márquez MC, Ventosa A, Barros JA (2004) Halomonas neptunia sp. nov., Halomonas sulfidaeris sp. nov., Halomonas axialensis sp. nov. and Halomonas hydrothermalis sp. nov.: halophilic bacteria isolated from deep-sea hydrothermal-vent environments. Int J Syst Evol Microbiol 55:499–511
Kharroub K, Jiménez-Pranteda ML, Aguilera M, Boulahrouf A, Ramos-Cormenzana A, Monteoliva-Sánchez M (2008) Halomonas sabkhae sp. nov., a moderately halophilic bacterium isolated from an Algerian sabkha. Int J Syst Evol Microbiol 58:40–44
Kim KK, Jin L, Yang HC, Lee S-T (2007) Halomonas gomseomensis sp. nov., Halomonas janggokensis sp. nov., Halomonas salaria sp. nov. and Halomonas denitrificans sp. nov., moderately halophilic bacteria isolated from saline water. Int J Syst Evol Microbiol 57:675–681
Kim KK, Lee KC, Oh HM, Lee JS (2010a) Halomonas stevensii sp. nov., Halomonas hamiltonii sp. nov. and Halomonas johnsoniae sp. nov., isolated from a renal care centre. Int J Syst Evol Microbiol 60:369–377
Kim MS, Roh SW, Bae JW (2010b) Cobetia crustatorum sp. nov., a novel slightly halophilic bacterium isolated from traditional fermented seafood in Korea. Int J Syst Evol Microbiol 60:620–626
Kim MS, Roh SW, Bae JW (2011) Halomonas jeotgali sp. nov., a new moderate halophilic bacterium isolated from a traditional fermented seafood. J Microbiol 48:404–410
Kim KK, Lee KC, Jeong H, Stevens DA, Lee JS (2012) Draft genome sequence of the human pathogen Halomonas stevensii S18214T. J Bacteriol 194:5143
Kim KK, Lee J-S, Stevens DA (2013) Microbiology and epidemiology of Halomonas species. Future Microbiol 8:1559–1573
Kleinsteuber S, Muller RH, Babel W (2001) Expression of the 2,4-D degradative pathway of pJP4 in an alkaliphilic, moderately halophilic soda lake isolate, Halomonas sp. EF43. Extremophiles 5:375–384
Knapp S, Landstein R, Galinski EA (1999) Extrinsic protein stabilization by the naturally occurring osmolytes β-hydroxyectoine and betaine. Extremophiles 3:191–198
Kraegeloh A, Amendt B, Kunte HJ (2005) Potassium transport in a halophilic member of the bacteria domain: identification and characterization of the K + uptake systems TrkH and TrkI from Halomonas elongata DSM 2581 T. J Bacteriol 187:1036–1043
Kraiwattanapong J, Ooi T, Kinoshita S (1999) Cloning and sequencing of a Deleya marina gene encoding for alginate lyase. Biotechnol Lett 21:169–174
Lang YJ, Bai L, Ren YN, Zhang LH, Nagata S (2011) Production of ectoine through a combined process that uses both growing and resting cells of Halomonas salina DSM 5928T. Extremophiles 15:303–310
Lapage SP, Sneath PHA, Lessel EF, Skerman VBD, Seeliger HPR, Clark WA (eds) (1992) International code of nomenclature of bacteria (1990 Revision). Bacteriological code. American Society for Microbiology, Washington, DC
le Phung T, Silver S, Trimble W, Gilbert JA (2012) Draft genome of Halomonas species strain GFAJ-1 (ATCC BAA-2256). J Bacteriol 194:1835–1836
Lee J-C, Jeon CO, Lim J-M, Lee S-M, Lee J-M, Song S-M, Park D-J, Li W-J, Kim C-J (2005) Halomonas taeanensis sp. nov., a novel moderately halophilic bacterium isolated from a solar saltern in Korea. Int J Syst Evol Microbiol 55:2027–2032
Li HB, Zhang LP, Chen SF (2008) Halomonas korlensis sp. nov., a moderately halophilic, denitrifying bacterium isolated from saline and alkaline soil. Int J Syst Evol Microbiol 58:2582–2588
Lim J-M, Yoon J-H, Lee J-C, Jeon CO, Park D-J, Sung C, Kim C-J (2004) Halomonas koreensis sp. nov., a novel moderately halophilic bacterium isolated from a solar saltern in Korea. Int J Syst Evol Microbiol 54:2037–2042
Lin Y, Fan H, Hao X, Johnstone L, Hu Y, Wei G, Alwathnani HA, Wang G, Rensing C (2012) Draft genome sequence of Halomonas sp. strain HAL1, a moderately halophilic arseniteoxidizing bacterium isolated from gold-mine soil. J Bacteriol 194:199–200
Lippert K, Galinski EA (1992) Enzyme stabilization by ectoine-type compatible solutes: protection against heating, freezing, and drying. Appl Microbiol Biotechnol 37:61–65
Llamas I, del Moral A, Béjar V, Girón MD, Salto R, Quesada E (1997) Plasmids from Halomonas eurihalina, a microorganism which produces an exopolysaccharide of biotechnological interest. FEMS Microbiol Lett 156:251–257
Llamas I, Sánchez MJ, Argandoña M, Béjar V, Quesada E, del Moral A (2002) Analysis of the genome of the moderate halophile Halomonas eurihalina. Curr Microbiol 45:233–239
Llamas I, Suárez A, Quesada E, Béjar V, del Moral A (2003) Identification and characterization of the carAB genes responsible for encoding carbamoylphosphate synthetase in Halomonas eurihalina. Extremophiles 7:205–211
Llamas I, del Moral A, Martínez-Checa F, Arco Y, Arias S, Quesada E (2006) Halomonas maura is a physiologically versatile bacterium of both ecological and biotechnological interest. Antonie Van Leeuwenhoek 89:395–403
Llamas I, Béjar V, Martínez-Checa F, Martínez-Cánovas MJ, Molina I, Quesada E (2011) Halomonas stenophila sp. nov., a halophilic bacterium that produces sulphate exopolysaccharides with biological activity. Int J Syst Evol Microbiol 61:2508–2514
Llamas I, Amjres H, Mata JA, Quesada E, Béjar V (2012) The potential biotechnological applications of the exopolysaccharide produced by the halophilic bacterium Halomonas almeriensis. Molecules 17:7103–7120
Long MR, Zhang DF, Yang XY, Zhang XM, Zhang YG, Zhang YM, Zhu H, Li WJ (2013) Halomonas nanhaiensis sp. nov., a halophilic bacterium isolated from a sediment sample from the South China Sea. Antonie Van Leeuwenhoek 103:997–1005
Luque R, Béjar V, Quesada E, Martínez-Checa F, Llamas I (2012) Halomonas ramblicola sp. nov., a moderately halophilic bacterium from Rambla Salada, a Mediterranean hypersaline rambla. Int J Syst Evol Microbiol 62:2903–2909
Maltseva O, McGowan C, Fulthorpe R, Oriel P (1996) Degradation of 2,4-dichlorophenoxyacetic acid by haloalkaliphilic bacteria. Microbiology 142:1115–1122
Maoz A, Mayr R, Scherer S (2003) Temporal stability and biodiversity of two complex antilisterial cheese-ripening microbial consortia. Appl Environ Microbiol 69:4012–4018
Margesin R, Schinner F (2001) Potential of halotolerant and halophilic microorganisms for biotechnology. Extremophiles 5:73–83
Martínez-Cánovas MJ, Béjar V, Martínez-Checa F, Quesada E (2004a) Halomonas anticariensis sp. nov., from Fuente de Piedra, a saline-wetland wildfowl reserve in Málaga, southern Spain. Int J Syst Evol Microbiol 54:1329–1332
Martínez-Cánovas MJ, Quesada E, Llamas I, Béjar V (2004b) Halomonas ventosae sp. nov., a moderately halophilic, denitrifying, exopolysaccharide-producing bacterium. Int J Syst Evol Microbiol 54:733–737
Martinez-Checa F, Toledo FL, Vilchez R, Quesada E, Calvo C (2002) Yield production, chemical composition, and functional properties of emulsifier H28 synthesized by Halomonas eurihalina strain H-28 in media containing various hydrocarbons. Appl Microbiol Biotechnol 58:358–363
Martínez-Checa F, Béjar V, Martínez-Cánovas MJ, Llamas I, Quesada E (2005) Halomonas almeriensis sp. nov., a moderately halophilic, exopolysaccharide-producing bacterium from Cabo de Gata, Almería, south-east Spain. Int J Syst Evol Microbiol 55:2007–2011
Martínez-Checa F, Toledo FL, El Mabrouki K, Quesada E, Calvo C (2007) Characteristics of bioemulsifier V2-7 synthesized in culture media added of hydrocarbons: chemical composition, emulsifying activity and rheological properties. Bioresour Technol 98:3130–3135
Mata JA, Martinez-Canovas J, Quesada E, Béjar V (2002) A detailed phenotypic characterisation of the type strains of Halomonas species. Syst Appl Microbiol 25:360–375
Mata JA, Béjar V, Llamas I, Arias S, Bressollier P, Tallon R, Urdaci MC, Quesada E (2006) Exopolysaccharides produced by the recently described halophilic bacteria Halomonas ventosae and Halomonas anticariensis. Res Microbiol 157:827–835
Mellado E, Ventosa A (2003) Biotechnological potential of moderately and extremely halophilic microorganisms. In: Barredo JL (ed) Microorganisms for health care, food and enzyme production. Research Signpost, Kerala, pp 233–265
Mellado E, Asturias JA, Nieto JJ, Timmis KN, Ventosa A (1995a) Characterization of the basic replicon of pCM1, a narrow-host-range plasmid from the moderate halophile Chromohalobacter marismortui. J Bacteriol 177:3443–3450
Mellado E, Moore ER, Nieto JJ, Ventosa A (1995b) Phylogenetic inferences and taxonomic consequences of 16S ribosomal DNA sequence comparison of Chromohalobacter marismortui, Volcaniella eurihalina, and Deleya salina and reclassification of V. eurihalina as Halomonas eurihalina comb. nov. Int J Syst Bacteriol 45:712–716
Mellado E, Garcia MT, Roldan E, Nieto JJ, Ventosa A (1998) Analysis of the genome of the gram-negative moderate halophiles Halomonas and Chromohalobacter by using pulsed-field gel electrophoresis. Extremophiles 2:435–438
Mellado E, Sanchez-Porro C, Martin S, Ventosa A (2004) Extracellular hydrolytic enzymes produced by moderately halophilic bacteria. In: Ventosa A (ed) Halophilic microorganisms. Springer, Berlin, pp 285–295
Menes RJ, Viera CE, Farías ME, Seufferheld MJ (2011) Halomonas vilamensis sp. nov., isolated from high-altitude Andean lakes. Int J Syst Evol Microbiol 61:1211–1217
Moreno ML, Sánchez-Porro C, Piubeli F, Frias L, García MT, Mellado E (2011) Cloning, characterization and analysis of cat and ben genes from the phenol degrading halophilic bacterium Halomonas organivorans. PLoS One 6:e21049
Moreno ML, Piubeli F, Bonfá MR, García MT, Durrant LR, Mellado E (2012) Analysis and characterization of cultivable extremophilic hydrolytic bacterial community in heavy-metal-contaminated soils from the Atacama Desert and their biotechnological potentials. J Appl Microbiol 113:550–559
Mormile MR, Romine MF, García MT, Ventosa A, Bailey TJ, Peyton BM (1999) Halomonas campisalis sp. nov., a denitrifying, moderately haloalkaliphilic bacterium. Syst Appl Microbiol 22:551–558
Muñoz JA, Pérez-Esteban B, Esteban M, de la Escalera S, Gómez MA, Martinez-Toledo MV, Gonzalez-López J (2001) Growth of moderately halophilic bacteria isolated from sea water using phenol as the sole carbon source. Folia Microbiol (Praha) 46:297–302
Nakayama H, Yoshida K, Ono H, Murooka Y, Shinmyo A (2000) Ectoine, the compatible solute of Halomonas elongata, confers hyperosmotic tolerance in cultured tobacco cells. Plant Physiol 122:1239–1247
Nieto JJ, Fernández-Castillo R, Márquez MC, Ventosa A, Quesada E, Ruiz-Berraquero F (1989) Survey of metal tolerance in moderately halophilic eubacteria. Appl Environ Microbiol 55:2385–2390
Nikkari S, Lopez FA, Lepp PW, Cieslak PR, Ladd-Wilson S, Passaro D, Danila R, Relman DA (2002) Broad-range bacterial detection and the analysis of unexplained death and critical illness. Emerg Infect Dis 8:188–194
Ntougias S, Zervakis GI, Fasseas C (2007) Halotalea alkalilenta gen. nov., sp. nov., a novel osmotolerant and alkalitolerant bacterium from alkaline olive mill wastes, and emended description of the family Halomonadaceae Franzmann et al. 1989, emend. Dobson and Franzmann 1996. Int J Syst Evol Microbiol 57:1975–1983
Ojima T, Saburi W, Yamamoto T, Kudo T (2012a) Characterization of Halomonas sp. strain H11 α-glucosidase activated by monovalent cations and its application for efficient synthesis of α-D-glucosylglycerol. Appl Environ Microbiol 78:1836–1845
Ojima T, Aizawa K, Saburi W, Yamamoto T (2012b) α-Glucosylated 6-gingerol: chemoenzymatic synthesis using α-glucosidase from Halomonas sp. H11, and its physical properties. Carbohydr Res 354:59–64
Okamoto T, Taguchi H, Nakamura K, Ikenaga H, Kuraishi H, Yamasato K (1993) Zymobacter palmae gen. nov., sp. nov., a new ethanol-fermenting peritrichous bacterium isolated from palm sap. Arch Microbiol 160:333–337
Okamoto T, Taguchi H, Nakamura K, Ikenaga H, Kuraishi H, Yamasato K (1995) Zymobacter palmae gen. nov., sp. nov. In: Validation of the publication of new names and new combinations previously effectively published outside the IJSB List No. 53. Int J Syst Bacteriol 45:418–419
Okamoto T, Maruyama A, Imura S, Takeyama H, Naganuma T (2004) Comparative phylogenetic analyses of Halomonas variabilis and related organisms based on 16S rRNA, gyrB and ectBC gene sequences. Syst Appl Microbiol 27:323–333
Okamoto T, Kuraishi H, Yamasato K (2005) Genus IV. Zymobacter. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, 2nd edn. Springer, New York, pp 319–323
Oren A (2010) Industrial and environmental applications of halophilic microorganisms. Environ Technol 31:825–834
Oren A, Ventosa A (2013) Subcommittee on the taxonomy of Halobacteriaceae and Subcommittee on the taxonomy of Halomonadaceae: minutes of the joint open meeting, 24 June 2013, Storrs, Connecticut, USA. Int J Syst Evol Microbiol 63:3540–3544
Oueriaghli N, González-Domenech CM, Martínez-Checa F, Muyzer G, Ventosa A, Quesada E, Béjar V (2014) Diversity and distribution of Halomonas in Rambla Salada, a hypersaline environment in the southeast of Spain. FEMS Microbiol Ecol 87:460–474
Parte AC (2014) List of prokaryotic names with standing in nomenclature. http://www.bacterio.net
Peçonek J, Gruber C, Gallego V, Ventosa A, Busse H-J, Kämpfer P, Radax C, Stan-Lotter H (2006) Reclassification of Pseudomonas beijerinckii Hof 1935 as Chromohalobacter beijerinckii comb. nov., and emended description of the species. Int J Syst Evol Microbiol 56:1953–1957
Pérez-Fernandez ME, Quesada E, Galvez J, Ruiz C (2000) Effect of exopolysaccharide V2-7, isolated from Halomonas eurihalina, on the proliferation in vitro of human peripheral blood lymphocytes. Immunopharmacol Immunotoxicol 22:131–141
Poli A, Esposito E, Orlando P, Lama L, Giordano A, de Appolonia F, Nicolaus B, Gambacorta A (2007) Halomonas alkaliantarctica sp. nov., isolated from saline lake Cape Russell in Antarctica, an alkalophilic moderately halophilic, exopolysaccharide-producing bacterium. Syst Appl Microbiol 30:31–38
Poli A, Nicolaus B, Denizci AA, Yavuzturk B, Kazan D (2013) Halomonas smyrnensis sp. nov., a moderately halophilic, exopolysaccharide-producing bacterium. Int J Syst Evol Microbiol 63:10–18
Prado B, Lizama C, Aguilera M, Ramos-Cormenzana A, Fuentes S, Campos V, Monteoliva-Sánchez M (2006) Chromohalobacter nigrandesensis sp. nov., a moderately halophilic, Gram-negative bacterium isolated from Lake Tebenquiche on the Atacama Saltern, Chile. Int J Syst Evol Microbiol 56:647–651
Qu L, Lai Q, Zhu F, Hong X, Zhang J, Shao Z, Sun X (2011) Halomonas daqiaonensis sp. nov., a moderately halophilic, denitrifying bacterium isolated from a littoral saltern. Int J Syst Evol Microbiol 61:1612–1616
Quesada E, Béjar V, Calvo C (1993) Exopolysaccharide production by Volcaniella eurihalina. Experientia 49:1037–1041
Quesada E, Béjar V, Ferrer MR, Calvo C, Llamas I, Martínez-Checa F, Arias S, Ruiz-García C, Páez R, Martínez-Cánovas MJ, del Moral A (2004) Moderately halophilic, exopolysaccharide-producing bacteria. In: Ventosa A (ed) Halophilic microorganisms. Springer, Berlin, pp 135–153
Quillaguamán J, Delgado O, Mattiasson B, Hatti-Kaul R (2004a) Chromohalobacter sarecensis sp. nov., a psychrotolerant moderate halophile isolated from the saline Andean region of Bolivia. Int J Syst Evol Microbiol 54:1921–1926
Quillaguamán J, Hatti-Kaul R, Mattiasson B, Alvarez MT, Delgado O (2004b) Halomonas boliviensis sp. nov., an alkalitolerant, moderate halophile isolated from soil around a Bolivian hypersaline lake. Int J Syst Evol Microbiol 54:721–725
Quillaguamán J, Delgado O, Mattiasson B, Hatti‐Kaul R (2006) Poly(β-hydroxybutyrate) production by a moderate halophile, Halomonas boliviensis LC1. Enzyme Microb Technol 38:148–154
Quillaguamán J, Muñoz M, Mattiasson B, Hatti‐Kaul R (2007) Optimizing conditions for poly(β-hydroxybutyrate) production by Halomonas boliviensis LC1 in batch culture with sucrose as carbon source. Appl Microbiol Biotechnol 74:981–986
Quillaguamán J, Guzmán H, Van-Thuoc D, Hatti-Kaul R (2010) Synthesis and production of polyhydroxyalkanoates by halophiles: current potential and future prospects. Appl Microbiol Biotechnol 85:1687–1696
Raj KC, Talarico LA, Ingram LO, Maupin-Furlow JA (2002) Cloning and characterization of the Zymobacter palmae pyruvate decarboxylase gene (pdc) and comparison to bacterial homologues. Appl Environ Microbiol 68:2869–2876
Raveendran S, Palaninathan V, Chauhan N, Sakamoto Y, Yoshida Y, Maekawa T, Mohanan PV, Kumar DS (2013) In vitro evaluation of antioxidant defense mechanism and hemocompatibility of mauran. Carbohydr Polym 98:108–115
Rodríguez-Moya J, Argandoña M, Iglesias-Guerra F, Nieto JJ, Vargas C (2013) Temperature- and salinity-decoupled overproduction of hydroxyectoine by Chromohalobacter salexigens. Appl Environ Microbiol 79:1018–1023
Romanenko LA, Schumann P, Rohde M, Mikhailov VV, Stackebrandt E (2002) Halomonas halocynthiae sp. nov., isolated from the marine ascidian Halocynthia aurantium. Int J Syst Evol Microbiol 52:1767–1772
Romanenko LA, Tanaka N, Svetashev VI, Falsen E (2013) Description of Cobetia amphilecti sp. nov., Cobetia litoralis sp. nov. and Cobetia pacifica sp. nov., classification of Halomonas halodurans as a later heterotrophic synonym of Cobetia marina and emended descriptions of the genus Cobetia and Cobetia marina. Int J Syst Evol Microbiol 63:288–297
Romano I, Nicolaus B, Lama L, Manca MC, Gambacorta A (1996) Characterization of a haloalkalophilic strictly aerobic bacterium, isolated from Pantelleria Island. Syst Appl Microbiol 19:326–333
Romano I, Giordano A, Lama L, Nicolaus B, Gambacorta A (2005) Halomonas campaniensis sp. nov., a haloalkaliphilic bacterium isolated from a mineral pool of Campania Region, Italy. Syst Appl Microbiol 28:610–618
Romano I, Lama L, Nicolaus B, Poli A, Gambacorta A, Giordano A (2006) Halomonas alkaliphila sp. nov., a novel halotolerant alkaliphilic bacterium isolated from a salt pool in Campania (Italy). J Gen Appl Microbiol 52:339–348
Romano I, Lama L, Orlando P, Nicolaus B, Giordano A, Gambacorta A (2007) Halomonas sinaiensis sp. nov., a novel halophilic bacterium isolated from a salt lake inside Ras Muhammad Park, Egypt. Extremophiles 11:789–796
Rosenberg A (1983) Pseudomonas halodurans sp nov, a halotolerant bacterium. Arch Microbiol 136:117–123
Ruiz-Ruiz C, Srivastava GK, Carranza D, Mata JA, Llamas I, Santamaría M, Quesada E, Molina IJ (2011) An exopolysaccharide produced by the novel halophilic bacterium Halomonas stenophila strain B100 selectively induces apoptosis in human T leukaemia cells. Appl Microbiol Biotechnol 89:345–355
Sakurai N, Sakurai T (1998) Genomic DNA cloning of the region encoding nitric oxide reductase in Paracoccus halodenitrificans and a structure model relevant to cytochrome oxidase. Biochem Biophys Res Commun 243:400–406
Sakurai N, Asada A, Mano S, Kataoka K, Sakurai T (2006) Tandem and single genes of three membrane-bound nitrate transporters in the nar gene cluster of the moderately halophilic denitrifier, Halomonas halodenitrificans. DNA Seq 17:363–369
Sanchez-Porro C, Martin S, Mellado E, Ventosa A (2003) Diversity of moderately halophilic bacteria producing extracellular hydrolytic enzymes. J Appl Microbiol 94:295–300
Sánchez-Porro C, Tokunaga H, Tokunaga M, Ventosa A (2007) Chromohalobacter japonicus sp. nov., a moderately halophilic bacterium isolated from a Japanese salty food. Int J Syst Evol Microbiol 57:2262–2266
Sánchez-Porro C, de la Haba RR, Soto-Ramírez N, Márquez MC, Montalvo-Rodríguez R, Ventosa A (2009) Description of Kushneria aurantia gen. nov., sp. nov., a novel member of the family Halomonadaceae, and a proposal for reclassification of Halomonas marisflavi as Kushneria marisflavi comb. nov., of Halomonas indalinina as Kushneria indalinina comb. nov. and of Halomonas avicenniae as Kushneria avicenniae comb. nov. Int J Syst Evol Microbiol 59:397–405
Sánchez-Porro C, Kaur B, Mann H, Ventosa A (2010) Halomonas titanicae sp. nov., a halophilic bacterium isolated from the RMS Titanic. Int J Syst Evol Microbiol 60:2768–2774
Sánchez-Porro C, de la Haba RR, Cruz-Hernández N, González JM, Reyes-Guirao C, Navarro-Sampedro L, Carballo M, Ventosa A (2013) Draft genome of the marine gammaproteobacterium Halomonas titanicae. Genome Announc 1:e0008313
Sauer T, Galinski EA (1998) Bacterial milking: a novel bioprocess for production of compatible solutes. Biotechnol Bioeng 57:306–313
Schwibbert K, Marin-Sanguino A, Bagyan I, Heidrich G, Lentzen G, Seitz H, Rampp M, Schuster SC, Klenk HP, Pfeiffer F, Oesterhelt D, Kunte HJ (2011) A blueprint of ectoine metabolism from the genome of the industrial producer Halomonas elongata DSM 2581T. Environ Microbiol 13:1973–1994
Sogutcu E, Emrence Z, Arikan M, Cakiris A, Abaci N, Öner ET, Ustek D, Arga KY (2012) Draft genome sequence of Halomonas smyrnensis AAD6T. J Bacteriol 194:5690–5691
Soto-Ramírez N, Sánchez-Porro C, Rosas S, González W, Quiñones M, Ventosa A, Montalvo-Rodríguez R (2007) Halomonas avicenniae sp. nov., isolated from the salty leaves of the black mangrove Avicennia germinans in Puerto Rico. Int J Syst Evol Microbiol 57:900–905
Sripo T, Phongdara A, Wanapu C, Caplan AB (2002) Screening and characterization of aldehyde dehydrogenase gene from Halomonas salina strain AS11. J Biotechnol 95:171–179
Stackebrandt E, Frederiksen W, Garrity GM, Grimont PAD, Kämpfer P, Maiden MCJ, Nesme X, Rosselló-Mora R, Swings J, Trüper HG, Vauterin L, Ward AC, Whitman WB (2002) Report of the ad hoc committee for the re-evaluation of the species definition in bacteriology. Int J Syst Evol Microbiol 52:1043–1047
Stevens DA, Hamilton JR, Johnson N, Kim KK, Lee JS (2009) Halomonas, a newly recognized human pathogen, causing infections and contamination in a dialysis center; 3 new species. Medicine (Baltimore) 88:244–249
Stevens DA, Kim KK, Johnson N, Lee JS, Hamilton JR (2013) Halomonas johnsoniae: review of a medically underappreciated genus of growing human importance. Am J Med Sci 345:335–338
Tahrioui A, Quesada E, Llamas I (2013a) Draft genome sequence of the moderately halophilic gammaproteobacterium Halomonas anticariensis FP35T. Genome Announc 1:e00497-13
Tahrioui A, Quesada E, Llamas I (2013b) Genetic and phenotypic analysis of the GacS/GacA system in the moderate halophile Halomonas anticariensis. Microbiology 159:462–474
Tanimura K, Nakayama H, Tanaka T, Kondo A (2013) Ectoine production from lignocellulosic biomass-derived sugars by engineered Halomonas elongata. Bioresour Technol 142:523–529
Ternan NG, McMullan G (2002) Utilisation of aminomethane sulfonate by Chromohalobacter marismortui VH1. FEMS Microbiol Lett 207:49–53
Tindall BJ, Rosselló-Móra R, Busse H-J, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266
Van-Thuoc D, Guzmán H, Quillaguamán J, Hatti-Kaul R (2010a) High productivity of ectoines by Halomonas boliviensis using a combined two-step fed-batch culture and milking process. J Biotechnol 147:46–51
Van-Thuoc D, Guzmán H, Thi-Hang M, Hatti-Kaul R (2010b) Ectoine production by Halomonas boliviensis: optimization using response surface methodology. Mar Biotechnol (NY) 12:586–593
Vargas C, Fernández-Castillo R, Cánovas D, Ventosa A, Nieto JJ (1995) Isolation of cryptic plasmids from moderately halophilic eubacteria of the genus Halomonas. Characterization of a small plasmid from H. elongata and its use for shuttle vector construction. Mol Gen Genet 246:411–418
Vargas C, Calderón MI, Capote N, Carrasco R, Garcia R, Morón MJ, Ventosa A, Nieto JJ (2004) Genetics of osmoadaptation by accumulation of compatible solutes in the moderate halophile Chromohalobacter salexigens: its potential in agriculture under osmotic stress conditions. In: Ventosa A (ed) Halophilic microorganisms. Springer, Berlin, pp 135–153
Ventosa A (2005) Genus III. Chromohalobacter. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, 2nd edn. Springer, New York, pp 316–319
Ventosa A, Quesada E, Rodriguez-Valera F, Ruiz-Berraquero F, Ramos-Cormenzana A (1982) Numerical taxonomy of moderately halophilic Gram-negative rods. J Gen Microbiol 128:1959–1969
Ventosa A, Gutiérrez MC, Garcia MT, Ruiz-Berraquero F (1989) Classification of “Chromobacterium marismortui” in a new genus, Chromohalobacter gen. nov., as Chromohalobacter marismortui comb. nov., nom. rev. Int J Syst Bacteriol 39:382–386
Ventosa A, Nieto JJ, Oren A (1998) Biology of moderately halophilic aerobic bacteria. Microbiol Mol Biol Rev 62:504–544
Von Graevenitz A, Bowman J, Del Notaro C, Ritzler M (2000) Human infection with Halomonas venusta following fish bite. J Clin Microbiol 38:3123–3124
Vreeland RH (2005) Genus I. Halomonas. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, 2nd edn. Springer, New York, pp 300–313
Vreeland RH, Ventosa A (2003) International Committee on Systematics of Prokaryotes Subcommittee on the Taxonomy of the Halomonadaceae: Minutes of the inaugural meetings, 31 July 2002, Paris, France. Int J Syst Evol Microbiol 53:921–922
Vreeland RH, Litchfield CD, Martin EL, Elliot E (1980) Halomonas elongata, a new genus and species of extremely salt-tolerant bacteria. Int J Syst Bacteriol 30:485–495
Wang Y-N, Cai H, Chi C-Q, Lu A-H, Lin X-G, Jiang Z-F, Wu X-L (2007a) Halomonas shengliensis sp. nov., a moderately halophilic, denitrifying, crude-oil-utilizing bacterium. Int J Syst Evol Microbiol 57:1222–1226
Wang Y-N, Cai H, Yu S-L, Wang Z-Y, Liu J, Wu X-L (2007b) Halomonas gudaonensis sp. nov., isolated from a saline soil contaminated by crude oil. Int J Syst Evol Microbiol 57:911–915
Wang Y, Tang S-K, Lou K, Mao P–H, Jin X, Jiang C-L, Xu L-H, Li W-J (2008a) Halomonas lutea sp. nov., a moderately halophilic bacterium isolated from a salt lake. Int J Syst Evol Microbiol 58:2065–2069
Wang Y, Wu Y-H, Wang C-S, Xu X-W, Oren A, Zhu X-F, Wu M (2008b) Halomonas salifodinae sp. nov., a halophilic bacterium isolated from a salt mine in China. Int J Syst Evol Microbiol 58:2855–2858
Wang Y, Tang S-K, Lou K, Lee J-C, Jeon CO, Xu L-H, Kim C-J, Li W-J (2009) Aidingimonas halophila gen. nov., sp. nov., moderately halophilic bacteria isolated from a salt lake. Int J Syst Evol Microbiol 59:3088–3094. doi:10.1099/ijs.0.036871-0. Epub 2012 Feb 3
Wang CY, Wu SJ, Ng CC, Tzeng WS, Shyu YT (2012) Halomonas beimenensis sp. nov., isolated from an abandoned saltern. Int J Syst Evol Microbiol 62:3013–3017
Wu G, Wu X-Q, Wang Y-N, Chi C-Q, Tang Y-Q, Kida K, Wu X-L, Luan Z-K (2008a) Halomonas daqingensis sp. nov., a moderately halophilic bacterium isolated from an oilfield soil. Int J Syst Evol Microbiol 58:2859–2865
Wu Y-H, Xu X-W, Huo Y-Y, Zhou P, Zhu X-F, Zhu H-B, Wu M (2008b) Halomonas caseinilytica sp. nov., a halophilic bacterium isolated from a saline lake on the Qinghai–Tibet Plateau, China. Int J Syst Evol Microbiol 58:1259–1262
Xu R, Zhang L (2012) Construction of ectoine absorption defective mutant for efficient ectoine production. Wei Sheng Wu Xue Bao 52:661–667
Xu X-W, Wu Y-H, Zhou Z, Wang C-S, Zhou Y-G, Zhang H-B, Wang Y, Wu M (2007) Halomonas saccharevitans sp. nov., Halomonas arcis sp. nov. and Halomonas subterranea sp. nov., halophilic bacteria isolated from hypersaline environments of China. Int J Syst Evol Microbiol 57:1619–1624
Xu L, Xu XW, Meng FX, Huo YY, Oren A, Yang JY, Wang CS (2013) Halomonas zincidurans sp. nov., a heavy-metal-tolerant bacterium isolated from the deep-sea environment. Int J Syst Evol Microbiol 63:4230–4236
Yarza P, Ludwig W, Euzèby J, Amann R, Schleifer K-H, Glöckner FO, Rosselló-Móra R (2010) Update of the All-Species Living Tree Project based on 16S and 23S-28S rRNA sequence analyses. Syst Appl Microbiol 33:291–299
Yoon J-H, Choi SH, Lee K-C, Kho YH, Kang KH, Park Y-H (2001) Halomonas marisflavae sp. nov., a halophilic bacterium isolated from the Yellow Sea in Korea. Int J Syst Evol Microbiol 51:1171–1177
Yoon JH, Lee KC, Kho YH, Kang KH, Kim CJ, Park YH (2002) Halomonas alimentaria sp. nov., isolated from jeotgal, a traditional Korean fermented seafood. Int J Syst Evol Microbiol 52:123–130
Zeigler DR (2003) Gene sequences useful for predicting relatedness of whole genomes in bacteria. Int J Syst Evol Microbiol 53:1893–1900
Zhang LH, Lang YJ, Nagata S (2009) Efficient production of ectoine using ectoine-excreting strain. Extremophiles 13:717–724
Zhao B, Wang H, Mao X, Li R, Zhang YJ, Tang S, Li WJ (2012) Halomonas xianhensis sp. nov., a moderately halophilic bacterium isolated from a saline soil contaminated with crude oil. Int J Syst Evol Microbiol 62:173–178
Zou Z, Wang G (2010) Kushneria sinocarnis sp. nov., a moderately halophilic bacterium isolated from a Chinese traditional cured-meat. Int J Syst Evol Microbiol 60:1881–1886
Acknowledgements
The work of authors was supported by grants from the Spanish Ministry of Science and Innovation (CGL2010-19303 and CGL2013-46941-P) and the Junta de Andalucía (P10-CVI-6226). FEDER funds also supported authors’ research.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this entry
Cite this entry
de la Haba, R.R., Arahal, D.R., Sánchez-Porro, C., Ventosa, A. (2014). The Family Halomonadaceae . In: Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F. (eds) The Prokaryotes. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-38922-1_235
Download citation
DOI: https://doi.org/10.1007/978-3-642-38922-1_235
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-38921-4
Online ISBN: 978-3-642-38922-1
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences