Abstract
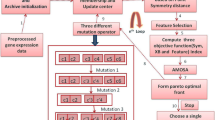
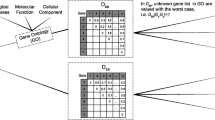
Microarray experiments generate a large amount of data which is used to discover the genetic background of diseases and to know the characteristics of genes. Clustering the tissue samples according to their co-expressed behavior and characteristics is an important tool for partitioning the dataset. Finding the clusters of a given dataset is a difficult task. This task of clustering is even more difficult when we try to find the rank of each gene, which is known as Gene Ranking, according to their abilities to distinguish different classes of samples. In the literature, many algorithms are available for sample clustering and gene ranking or selection, separately. A few algorithms are also available for simultaneous clustering and feature selection. In this article, we have proposed a new approach for clustering the samples and ranking the genes, simultaneously. A novel encoding technique for the chromosomes is proposed for this purpose and the work is accompleshed using a multi-objective evolutionary technique. Results have been demonstrated for both artificial and real-life gene expression data sets.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
References
Knowles, J.D., Corne, D.W.: Pareto archived evolution strategy: A new baseline algorithm for Pareto multiobjective optimization. In: Congress on Evolutionary Computation (CEC 1999), vol. 1, pp. 98–105. IEEE Press, Piscataway (1999)
Ben-Dor, A., et al.: Clustering gene expression patterns. Journal of Computational Biology 6, 281–297 (1999)
Duda, R.O., Hart, P.E.: Pattern Classification and Scene Analysis. John Wiley and Sons, Inc., New York (1973)
Guyon, I., Elissee, A.: An introduction to variable and feature selection. Journal of Machine Learning Research 3, 1157–1182 (2003)
Deb, K., Pratap, A., Agrawal, S., Meyarivan, T.: A fast and elitist multiobjective genetic algorithm: NSGA-II. IEEE Transaction on Evolutionary Computation 6, 182–197 (2002)
Theodoridis, S., Koutroubas, K.: Pattern Recognition. Academic Press, London (1999)
Halkidi, M., Vazirgiannis, M., Batistakis, Y.: Quality scheme assessment in the clustering process. In: Zighed, D.A., Komorowski, J., Żytkow, J.M. (eds.) PKDD 2000. LNCS (LNAI), vol. 1910, pp. 265–276. Springer, Heidelberg (2000)
Rezaee, R., Lelieveldt, B.P.F., Reiber, J.H.C.: A New Cluster Validity Index for the Fuzzy c-Mean. Pattern Recognition Letters 19, 237–246 (1998)
Sharma, S.C.: Applied Multivariate Techniques. John Wiley and Sons, Chichester (1996)
Xie, X., Beni, G.: A validity measure for fuzzy clustering. IEEE Transaction on P.A.M.I. 13(4), 841–846 (1991)
Davies, D., Bouldin, D.: A cluster separation measure. IEEE Transaction on P.A.M.I 1(2), 224–227 (1979)
Dunn, J.C.: A Fuzzy Relative of the ISODATA Process and Its Use in Detecting Compact Well-Separated Clusters. Journal of Cybernetics 3(3), 32–57 (1973)
Maulik, U., Bandyopadhyay, S.: Performance Evaluation of Some Clustering Algorithms and Validity Indices. IEEE Transactions on Pattern Analysis and Machine Intelligence 24(12) (December 2002)
Mukhopadhyay, A., Maulik, U., Bandyopadhyay, S.: Multi-objective Genetic Algorithm based Fuzzy Clustering of Categorical Attributes. IEEE Transactions on Evolutionary Computation 13(5), 991–1005 (2009)
Bandyopadhyay, S., Saha, S., Maulik, U., Deb, K.: A Simulated Annealing-Based Multiobjective Optimization Algorithm: AMOSA. IEEE Transactions on Evolutionary Computation 12(3), 269–283 (2008)
Zhang, C., Lu, X., Zhang, X.: Significance of gene ranking for classification of microarray samples. IEEE/ACM Transaction on Computational Biology and Bioinformatics 3(3), 312–320 (2006)
Garey, M.R., Johnson, D.S.: Computers and Intractability: A Guide to the Theory of NP-Completeness. W.H. Freeman, New York (1979)
Jong, K., Mary, J., Cornuéjols, A., Marchiori, E., Sebag, M.: Ensemble feature ranking. In: Boulicaut, J.-F., Esposito, F., Giannotti, F., Pedreschi, D. (eds.) PKDD 2004. LNCS (LNAI), vol. 3202, pp. 267–278. Springer, Heidelberg (2004)
Streib, F.E., Dehmer, M., Liu, J., Muhlhuser, M.: A systems approach to gene ranking from dna microarray data of cervical cancer. In: World Academy of Science, Engineering and Technology, vol. 8 (October 2005)
Sharan, R.: Click and expander: a system for clustering and visualizing gene expression data. Bioinformatics 19, 1787–1799 (2003)
Burges, C., Shaked, T., Renshaw, E., Lazier, A., Deeds, M., Hamilton, N., Hullender, G.: Learning to rank using gradient descent. In: ICML (2005)
Freund, Y., Schapire, R.E.: Experiments with a new boosting algorithm. In: Proceedings of the Thirteenth International Conference on Machine Learning, pp. 148–156 (1996)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Mondal, K.C., Mukhopadhyay, A., Maulik, U., Bandhyapadhyay, S., Pasquier, N. (2011). MOSCFRA: A Multi-objective Genetic Approach for Simultaneous Clustering and Gene Ranking. In: Rizzo, R., Lisboa, P.J.G. (eds) Computational Intelligence Methods for Bioinformatics and Biostatistics. CIBB 2010. Lecture Notes in Computer Science(), vol 6685. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-21946-7_14
Download citation
DOI: https://doi.org/10.1007/978-3-642-21946-7_14
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-21945-0
Online ISBN: 978-3-642-21946-7
eBook Packages: Computer ScienceComputer Science (R0)