Abstract
Peripheral T-cell lymphoma (PTCL) is an uncommon group of lymphoma covering a diverse spectrum of entities. Little was known regarding the molecular and genomic landscapes of these diseases until recently but the knowledge is still quite spotty with many rarer types of PTCL remain largely unexplored. In this chapter, the recent findings from gene expression profiling (GEP) studies, including profiling data on microRNA, where available, will be presented with emphasis on the implication on molecular diagnosis, prognostication, and the identification of new entities (PTCL-GATA3 and PTCL-TBX21) in the PTCL-NOS group. Recent studies using next-generation sequencing have unraveled the mutational landscape in a number of PTCL entities leading to a marked improvement in the understanding of their pathogenesis and biology. While many mutations are shared among PTCL entities, the frequency varies and certain mutations are quite unique to a specific entity. For example, TET2 is often mutated but this is particularly frequent (70-80%) in angioimmunoblastic T-cell lymphoma (AITL) and IDH2 R172 mutations appear to be unique for AITL. In general, chromatin modifiers and molecular components in the CD28/T-cell receptor signaling pathways are frequently mutated. The major findings will be summarized in this chapter correlating with GEP data and clinical features where appropriate. The mutational landscape of cutaneous T-cell lymphoma, specifically on mycosis fungoides and Sezary syndrome, will also be discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Peripheral T-cell lymphoma
- Gene expression profiling
- Mutational landscape
- Pathogenesis
- Biology
- Mycosis fungoides
- Sezary syndrome
2.1 Introduction
Peripheral T-cell lymphoma (PTCL) constitutes ~10–15% of all non-Hodgkin lymphomas (NHLs) in the western world [1–3] but may represent a higher proportion (~20–30%) of NHL in Asia and South America [1, 4, 5, 6]. The current World Health Organization (WHO) classification includes at least 27 subtypes of PTCL, whose incidence varies significantly among different geographic locations and different ethnic groups [7–9]. The differences could be due to genetic predisposition, a combination of genetic and environmental factors, or predominantly environmental factors. Despite a plateau observed in the incidence of B-cell lymphoma in the USA, an increase in the incidence of PTCL has been noted in recent studies [9, 10]. Whether this is due to improved diagnosis or due to epidemiological changes is not known. In general, NHL incidence increases with age and is higher in males than females, and the same pattern is also true in PTCL, with a few exceptions. Compared to B-cell lymphomas, T-cell malignancies more often relapse after chemotherapy and have shorter overall survival (OS) [11–13] (Fig. 2.1). The introduction of a number of novel therapies has done little to improve the long-term survival, as recent meta-analysis found no improvement in clinical outcome of AITL or PTCL patients in the last two decades [14–16].
Signaling through the T-cell receptor and co-stimulatory pathways: Upon binding of the TCR to pMHC, LCK, and FYN phosphorylate CD3 ITAMs, followed by recruitment and activation of ZAP70, which in turn phosphorylates LAT. A signalosome condenses on LAT clusters through binding to phosphotyrosines by SH2-containing proteins, which in turn are phosphorylated. Binding to proline-rich motifs through SH3 domains contributes to these recruitment steps. This leads to activation of PLCG1 and the subsequent generation of IP3 and DAG and activation of downstream pathways. PI3K is recruited and activated through several mechanisms, including CD28 activation, as shown. The resulting PIP3 contributes to the membrane recruitment and activation of AKT. Only a small fraction of the involved signaling molecules are shown. Proteins with frequent gain-of-function mutations or dominant-negative mutations (RHOA) in (PTCL are shown in red
The current updated edition of the WHO classification (2016 update) recognizes 7 more entities than the 2008 edition and 14 more than the 2001 edition, reflecting, in part, an improvement in the understanding of PTCL biology in the past 15 years [2, 3]. Although the major diagnostic criteria are the pathological and immunophenotypical characteristics, recent genetic studies have substantially improved the accuracy of classification, and additional molecular entities have been recognized as a result of genomic studies in T-cell and NK-cell neoplasms. The major PTCL entities presenting as nodal disease include angioimmunoblastic T-cell lymphoma (AITL), anaplastic large cell lymphoma (ALCL), and a large group of cases that cannot be specifically classified with current methods and are categorized as PTCL, not otherwise specified (PTCL-NOS) [7, 2, 3]. Adult T-cell leukemia/lymphoma (ATLL) and extranodal NK/T-cell lymphoma of nasal type (ENKTCL) [17] are primarily extranodal tumors that are common in specific geographical regions but rare elsewhere. Our subsequent discussions will be focused mainly on the above entities, as much of the recent progress has been in these diseases. There are other less frequent PTCL entities that are mostly extranodal tumors [17], and we will describe relevant findings where available. We will also describe and discuss the recent studies on cutaneous T-cell lymphoma (CTCL), focusing on mycosis fungoides (MF) and Sézary syndrome.
2.2 T-Cell Development and Activation: An Overview
Mature T-cells can broadly be separated into two categories based on the T-cell receptor (TCR) types expressed; [18] the α-chain always pairs with β chain to form a αβ receptor in αβ T-cells, which comprise approximately 98% of mature T-cells. The remaining 2% of T-cells express the γ and δ chains of the TCR and are abundant in mucosal surfaces. These frequencies are also reflected in the incidence of the T-cell lymphoma entities, as PTCLs expressing αβ TCR are far more frequent than γδ-PTCLs.
A brief overview of human normal T-lymphocyte differentiation and activation is presented in this subsection because it is crucial for the understanding of the heterogeneity associated with PTCL and because molecular abnormalities in PTCL frequently target genes encoding proteins with important roles in normal T-cell activation. The hematopoietic stem cell (HSC) in the bone marrow differentiates to a common lymphoid progenitor (CLP) from which the entire lymphoid lineage is derived. The CLP emigrates from the bone marrow to the thymus, where it undergoes further maturation [19] through the sequential generation of functional β and α TCR chains through recombination events. As the thymocyte progresses through well-defined developmental stages, a functional TCRβ-chain and a pre-α chain (non-recombined) are synthesized in the initial stages, to form a pre-TCR with CD3 subunits γ, δ, ε, and ζ, and subsequently a mature αβ-TCR, with the replacement of the pre-α chain with a mature α chain [18]. At the end of this developmental stage, the T-cells are single positive, expressing either CD4 or CD8, which allows them to interact with MHC II or MHC I, respectively. In the thymus, T-cells undergo both positive and negative selection based on the strength of the interaction of the TCR with self-peptides bound to MHC (pMHC), so that only cells whose TCRs have a modest affinity for self pMHC survive.
Mature naïve αβ CD4+ T-cells circulate throughout the body and enter secondary lymphoid organs––spleen, lymph nodes, and oral and gastrointestinal mucosa—to provide immunosurveillance. These quiescent T-cells express a mature TCR complex (αβ-chains, and CD3γ, δ, two ε and two ζ chains) that interacts with pMHC on antigen-presenting cells (APCs). When a T-cell finds an APC presenting a pMHC with which it can interact strongly, it is activated by recruiting a large array of signaling molecules to the T-cell-APC junction, the immunological synapse. This signaling is enhanced by ligation of co-stimulatory molecules, of which the most important in naïve T-cells is CD28. Strikingly, genes encoding many proteins important in TCR and CD28 signaling are mutated or affected by copy number abnormalities in PTCL, and, in this brief overview, we will put emphasis on these proteins (Fig. 2.2).
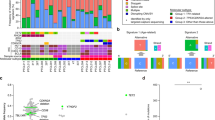
a Unique gene expression signatures were identified for major Peripheral T-cell Lymphoma (PTCL) entities. Anaplastic large cell lymphoma (ALCL) and Extranodal natural killer/T-cell lymphoma (ENKTL), groups are further differentiated into ALK-positive ALCL and ALK-negative ALCL, and NK and γδ T-cell subgroups, respectively. b Overall survival (OS) in the different molecular PTCL subgroups. (C and D) GATA3 and TBX21 subgroups within PTCL-NOS with (C) biological and (D) overall survival differences. OS analysis of molecularly defined GATA3 and TBX21 subgroups showed a significant difference in clinical outcome (P = 0.01)
Upon ligation of the TCR with a cognate pMHC, the MHC II also interacts with CD4, which is constitutively associated with the tyrosine kinase LCK. LCK phosphorylates the immunoreceptor tyrosine-based activation motifs (ITAMs) on the CD3ζ chains (and to a lesser extent those on the other CD3 subunits), which allows binding by the kinase ζ-associated protein of 70 kilodaltons (ZAP-70), which is then phosphorylated and activated by LCK. Active ZAP-70 then phosphorylates linker of activated T-cells (LAT) on multiple tyrosines, allowing the recruitment of a large signaling complex that includes VAV1, ITK, SOS1 (an activator of the RAS-MEK-ERK pathway), and phospholipase Cγ1 (PLCγ1), among others. The tyrosine kinase ITK phosphorylates and activates PLCγ1, which then cleaves phosphatidylinositol (4, 5) bisphosphate to generate two important second messengers, inosine trisphosphate (IP3) and diacylglycerol (DAG).
IP3 binds to receptors on the endoplasmic reticulum (ER), leading to an initial phase of calcium release, which, after calcium depletion in the ER, is followed by further calcium flux due to activation of plasma membrane calcium release-activated channels (CRACs). Intracellular calcium activates several pathways; in T-cells, a particularly important effect is activation of the nuclear factor of activated T-cells (NFAT) family of transcription factors, whose targets include many important cytokines such as interleukin-2 (IL-2). DAG activates several important proteins. These include several isoforms of protein kinase C (PKC) and RASGRP1 and −2, which are responsible for an initial phase of RAS activation, which is sustained and amplified by SOS1 [20].
LCK also interacts with and phosphorylates FYN [21], the other Src family tyrosine kinase highly expressed in T-cells. FYN has important positive and negative effects on T-cell signaling. FYN-deficient mouse T-cells have severe defects in signaling induced by anti-CD3 antibodies [22], a condition in which LCK is poorly recruited to the TCR. In contrast, under normal antigenic stimulation, abnormalities in FYN-deficient T-cells are subtle. In addition to contributing to ITAM phosphorylation, the positive effects of FYN include enhancement of NF-κB activation [23] and cell adhesion to B-cells [24] through its interaction with SAP (SH3D1A) bound to SLAM family members [25], whereas it is also important in a negative feedback loop that inactivates both FYN and LCK [26].
Despite decades of experimentation, the pathways through which CD28 ligation leads to costimulation are still incompletely understood. LCK phosphorylates CD28, leading to recruitment of PI3 K, VAV1, and several other molecules, including PKCθ [27]. PI3 K is similarly recruited and activated by ligation of ICOS. In addition to stimulation of the AKT–mTOR pathway, the PIP3 generated by PI3 K also contributes to membrane recruitment of several important proteins, including ITK, which activates PLCγ1, and PDK1, which activates not only AKT but also PKCθ, which in turn phosphorylates PDK1, increasing its stability [28]. PKCθ phosphorylates membrane-associated CARD11, which then assembles a complex with BCL10 and MALT1 (the CBM complex) [29], which, through several intermediate steps, activates nuclear factor κB (NF-κB). PKC-θ is also important in activating AP-1 [30] (heterodimers of the JUN and FOS family of transcription factors), which is essential for activating transcription of IL-2 and other genes required for T-cell proliferation.
2.3 TCR Rearrangement in the Diagnosis of PTCL
Assessment of T-cell clonality is a critical test in the diagnosis of PTCL. Each tumor arises from alterations in the progeny of a T-cell with uniquely rearranged TCR loci. Each rearrangement has distinctive TCR junctional regions, which can serve as a tumor-specific marker [31]. Although clonality is strongly supportive of the diagnosis of neoplasia, it is not sufficient to establish the diagnosis, which requires the integration of all relevant clinical, morphological, and immunophenotypic data. Clonal rearrangements of the TCR β and/or γ loci are detectable in > 90% of T-cell lymphomas [32, 33], even though some T-cell neoplasms lack TCR expression, a common feature in ALCL. The newly defined mutations and structural abnormalities identified in PTCL (described below), can provide additional markers for the diagnosis.
2.4 Pathobiology of PTCL with Molecular Genomic Approaches
Global gene expression profiling using RNA-seq or microarray analysis has been extensively applied to lymphomas to define their oncogenic signatures as well as other characteristic features common to a group of tumors or unique to each tumor [34, 35]. More recently, additional global studies including miRNA profiling [36–38], mutation analysis, copy number alterations, and epigenetics analysis have been published [39–44]. We will discuss major findings from GEP studies of the more common PTCL entities and some rarer ones with sufficient data [11–13]. We will also discuss complementary studies that may validate or provide insight into the biological basis of these functional profiles as well as their likely impact on the diagnosis and management of PTCL patients.
2.5 Peripheral T-Cell Lymphoma, Not Otherwise Specified (PTCL-NOS)
PTCL-NOS represent the most common group of PTCL (~30%). By definition, it does not represent an entity, but a heterogeneous group of mainly nodal cases that cannot be specifically assigned to any known WHO classified entities[2, 3, 17]. Therefore, the morphologic and immunophenotypic spectrum of PTCL-NOS is broad. Some morphological variants such as lymphoepithelioid (Lennert) lymphoma have been described within PTCL-NOS, but whether these variants define unique biological entities is not clear. The diagnosis is challenging with a high frequency of discordant diagnosis among pathologists [45–47].
2.6 Gene Expression Profiling Revealed Biological Subgroups in PTCL-NOS
Major challenges in the initial GEP studies of PTCL-NOS were due to the limited number of cases studied and lack of a standardized platform [48–50]. Although several distinct clusters were identified in earlier studies, their biological significance was not clear and largely reflected characteristics of the tumor microenvironment [48–50]. These studies also reported that NF-κB activation was associated with a favorable clinical outcome and a high proliferation gene signature with a poor outcome [51, 52] TH1/TH2-associated chemokine expression defined by immunohistochemistry was also found to be correlated with overall survival [36]. Additional studies showed an association of PTCL-NOS cases with activated CD4+ or CD8+ T-cells [50] and identified molecular classifiers in FFPE cases [53]. There was little consensus in these studies, which may be due to either platform differences, small sample size, or differences in the patient populations; nevertheless, overall they indicated the presence of multiple molecular subgroups within nodal PTCL-NOS [54].
Using GEP, we were able to define diagnostic molecular signatures for the major subgroups of PTCL, but about a third of the PTCL cases in this study remained uncharacterized, although a small subset of these cases showed features of αβ cytotoxic T-cells with poor clinical outcome [35, 11, 12, 13] (Fig. 2.3a, b). To further study PTCL-NOS, an international collaborative effort involving two major consortiums (LLMPP and I-PTCL) and a meta-analysis of two previously published series totaling 350 PTCL cases led to the identification of at least two novel biological subgroups. One subgroup (33% of PTCL-NOS) was characterized by high expression of GATA3 and its target genes (e.g., CCR4, IL18RA, CXCR7, IK), and the other subgroup by high expression of TBX21 and EOMES (49% of PTCL-NOS) and their known target genes (e.g., CXCR3, IL2RB, CCL3, IFNγ) (Fig. 2.3c, d). The “high GATA3” subgroup was associated with poor clinical outcome and the lack of a prominent microenvironmental signature. It was associated with high MYC and proliferation signatures, whereas the NF-κB pathway was enriched in the TBX21 subgroup, which may likely explain the more favorable outcome associated with the NF-κB pathway activation reported in a previous study [52]. Our study also suggested that the TBX21 subgroup may contain a subset with high cytotoxic signature, which is associated with a poorer clinical outcome compared to the rest of the TBX21 subgroup. GATA3 and TBX21 are master regulators of T-cell differentiation (Th2 and Th1/cytotoxic T-cells, respectively) and may suggest the cell of origin in these subsets [11]. However, the data need to be interpreted cautiously due to the plasticity of the T-cell molded by the cytokine milieu [55]. The adverse outcome of the GATA3 subgroup is further supported by an independent study using immunohistochemistry for GATA3 expression [56]. Our study also identified a diagnostic signature that distinguishes ALK-negative ALCL from PTCL-NOS which is useful particularly in cases with high CD30 expression [11].
a A unique gene expression signature for ALCL was identified, and then ALK-positive ALCL cases were separated from ALK-negative ALCL (reclassified PTCL-NOS cases are indicated at the top.) (Adapted from Iqbal et al. Blood 2014). b Unsupervised hierarchical clustering classifies ALCL into distinct groups which show significantly different overall survival (OS). (Adapted from Abate et al. Leukemia 2014) c OS of patients with ALCL, stratified by rearrangements of ALK, DUSP22, and TP63 and triple-negative cases (−/−/− ; lacking all 3 rearrangements) (Adapted from Parilla-Castellar et al. Blood 2014). d Recurrent mutations in the JAK-STAT3 pathway with JAK1 and STAT3 being the most common mutated genes
2.7 Genetic Studies Demonstrated Distinct Entities Within PTCL-NOS
We performed a high-resolution copy number analysis on the PTCL-GATA3 and PTCL-TBX21 subgroups, which revealed that these two subgroups showed significant differences in the profile of copy number aberrations (CNAs), and expectedly higher genomic complexity is associated with the PTCL-GATA3 subgroup. This preliminary analysis provided significant genetic evidence that these two subgroups represent distinct diseases that utilize different oncogenic pathways for tumorigenesis [57]. Efforts are being made to bring this information to clinical practice by translating the diagnostic signatures to platforms more amenable to routine clinical practice.
Scanty data are available so far regarding the mutational landscape of PTCL-NOS. Some mutations that are frequent in AITL (see below) are also detected in PTCL-NOS but at lower frequencies. A study using a limited capture panel of known mutated genes [58] reported frequent mutations in epigenetic regulators including MLL2, KDM6A, MLL, TET2, and DNMT3A. Cases showing alterations in histone methyltransferase genes (MLL, MLL2, or KDM6A) were associated with poor clinical outcome (P = 0.0198). Several recent studies have identified VAV1 fusion transcripts that have the common theme of deleting the terminal SH2 domain, thus eliminating an autoregulatory domain and possibly causing constitutive VAV1 activation [59, 60]. A small deletion affecting intron 25 extending to exon 26 of VAV1 is detected in number of PTCL cases resulting in the abnormal splicing of exon 26, again leading to a protein that is constitutively active. Thus, VAV1 activation could be a significant pathogenetic mechanism in this group of lymphomas.
2.8 Limited MicroRNA Profiling Studies in PTCL-NOS
One study [61] on microRNA profiling of 23 PTCL-NOS cases reported miRNAs that are differentially expressed compared to activated CD4+ and CD8+ T-lymphocytes. This study found that miR-132-3p may be an important modulator of the PTCL-NOS transcriptome, but this was a limited study, and a much more comprehensive international effort would be needed to characterize the miRNA transcriptome and understand its biological meaning.
2.9 Angioimmunoblastic T-Cell Lymphoma (AITL)
AITL represents the most common well-defined PTCL entity and accounts for about 30% of PTCL cases with unique clinical characteristics and pathological features[2, 3, 17]. Morphologically, it is characterized by a diffuse or T-zone expansion of CD4+ T-cells accompanied by a proliferation of arborizing blood vessels with features of high endothelial venules. By immunohistochemistry, the majority of the neoplastic cells express two or more T-follicular helper (TFH) markers, such as PD-1, BCL6, CXCL13, CD10, ICOS, and CXCR5. Epstein–Barr virus (EBV)-infected B-cells are frequently present in the tumor microenvironment, and scattered patches of follicular dendritic cells are commonly found. There has been little clinical improvement in the past two decades, despite the introduction of novel therapies [14].
In the 2016 updated WHO classification, the subgroup of PTCL-NOS that expresses at least two TFH markers, but without the typical morphology of AITL, is designated as nodal PTCL with TFH phenotype [2, 17]. Moreover, a recent study suggested that this group shares not only phenotypic features with AITL, but also similar clinical features, genetic abnormalities, and molecular signatures, and based on these unifying features are likely part of the AITL spectrum [62]. It has been proposed to unify the entities of nodal PTCL with TFH phenotype, follicular T-cell lymphoma, and AITL under the umbrella term of “nodal T-cell lymphomas with T-follicular helper phenotype” [2].
Cell of origin of AITL: We and others have now convincingly demonstrated that the follicular helper T-cell (TFH) is the cell of origin of AITL [11, 12, 48, 63]. TFH cells, located within germinal centers (GC) of lymph nodes, are essential for the development of high-affinity antibodies by driving cognate GC B-cells to proliferate and differentiate into plasma cells [64]. The GCs in AITL are often involuted or largely effaced; paradoxically, tumor cells do not show GC localization except in the rare follicular PTCL. Normal TFH cells express a chemokine receptor (CXCR5) that drives their migration into B-cell follicles, bringing them into close proximity with B-cells. High affinity TCR epitopes are preferentially observed in TFH populations compared to other CD4+ T-cell subsets [65]. BCL6 has been identified as a master regulator of TFH differentiation in murine models and has been shown to negatively regulate differentiation to other CD4+ T-cell subsets [66]. It appears to also have a critical role in human TFH cells, but other transcription factors, such as MAF, are also important. Even though BCL6 and MAF are crucial for human TFH development, BCL6 is infrequently detected in AITL tumor cells [67, 68], but MAF [69, 70] expression is noted in a substantial number of AITLs. The frequent lack of BCL6 expression is in contrast to the consistent expression of BCL6 in GC B-cells [71].
Initial GEP studies showed that the majority of AITLs clustered together and identified a molecular classifier for AITL, which was dominated by the tumor microenvironment, with the presence of B-cells and a complex cytokine profile in the tumor milieu. The molecular classifier also identified at least 18% of pathologically defined PTCL-NOS cases as AITL [37] (Fig. 2.3a, b). These studies also identified oncogenic pathways including the NF-κB pathway, IL-6 signaling, and the TGFβ pathway [48, 50, 72, 73] enriched in AITL. Recently, GEP on a much larger series refined the gene expression signatures for AITL [11] and reclassified 14% of PTCL-NOS cases as AITL, similar to the previous observation [37]. The validity of this reclassification was further supported by the analysis of IDH2 mutations (R172K/S/T/G/M), which are frequent (~33%) in molecularly defined AITL, but not in other PTCL entities [41]. In contrast, in a subset of pathologically defined AITL cases that were not classified by GEP as AITL, IDH2 mutation was infrequent (7%; 1/14) [11].
Genetic studies in AITL: Few molecular genetics studies have been reported in AITL [74], although a few cytogenetic studies [75–77] have shown recurrent trisomy-5, often co-occurring with trisomy 21. We recently performed a high-resolution aCGH study, which showed that AITL had the least abnormal genome (7%) compared to other major PTCL entities, with CN gains more frequent than CN losses. Chromosome 5 gain (15 of 38; 40%) and chromosome 21 gain (8 of 38; 21%) were the most frequent abnormalities and significantly co-occur (p = 0.003, Fisher’s exact test). Losses were infrequent in AITL, and more focal than the pattern of the entire chromosome/arm abnormalities observed in gains [57]. Several small focal deletions showed enrichment in genes resulting in dysregulation of the PI3 K–AKT–mTOR pathway. Whether constitutive activation of this pathway is critical for maintenance of a malignant phenotype in TFH cells will be crucial to determine in the future studies.
Mutation spectrum of AITL: TET2 inactivating missense or nonsense mutations or insertions/deletions (indels) have been found variably in up to 85% of AITL cases suggesting high selective pressure in AITLs for loss of TET2 function [78–80]. Mutations in other family members, TET1 and TET3, however, are not observed, despite their homology, similar functions, and the likely ability for TET2 to compensate for TET1 in knockout mice [81], even though Tet1/Tet2 double knockout mice display some defects in TH17 differentiation [82]. Loss-of-function TET2 mutations are expected to result in DNA hypermethylation (5mC) and a decrease in 5-hydroxymethylcytosine (5hmC) in regions of the chromosome. 5hmC may not be just a byproduct of oxidation of 5mC but may have physiological functions [83, 84]. Thus, it is important to determine the profiles of aberrant 5mC and 5hmC in AITL to gain further insight to the consequence of TET2 inactivation. TET2 protein may also recruit other proteins such as HDAC1/2 and hence may perform functions unrelated to DNA demethylation which may also be affected by the loss of the TET protein.
IDH1 and IDH2 encode cytosolic and mitochondrial forms of isocitrate dehydrogenase, respectively, that catalyze the interconversion of isocitrate and α-ketoglutarate (αKG). The mutations in IDH1 and IDH2 found in cancer confer a novel enzymatic activity with conversion of α-ketoglutarate to the (R)-enantiomer of 2-hydroxyglutarate ((R)-2-HG) [85, 86] with depletion of αKG and generation of a large quantity of 2-HG, which acts as an oncometabolite by competitively inhibiting α-KG-dependent dioxygenases such as histone lysine demethylases, prolyl hydroxylases, collagen prolyl-4-hydroxylases, several DNA repair and RNA modifying enzymes, and the TET family of 5-methylcytosine (5mC) hydroxylases [87]. IDH1R132H, IDH2R140Q, and IDH2R172K knock-in (KI) mouse models showed an increase in lineage-restricted hematopoietic progenitors, a decrease in BM cellularity, and extramedullary hematopoiesis, but no effect on HSC self-renewal or expansion [88–90]. As R-2-HG has a wide array of effects beyond the altering of DNA 5mC and 5hmC patterns through inhibition of TET family members [91], the functional role of these effects warrants further investigation.
There is a remarkable similarity in the mutations of epigenome modifiers (e.g., TET2, IDH2, and DNMT3A) between AITL and myeloid neoplasms [92–94]. There are, however, also striking differences in the pattern of mutations between the two diseases: (i) TET2 and IDH1 or IDH2 mutations are mutually exclusive in acute myeloid leukemia (AML) and chronic myelomonocytic leukemia (CMML) [95]; however, IDH2 mutations often co-occur with TET2 in AITL [96, 97]. (ii) IDH1 mutations are not observed in AITL [96]. Of the two recurrent IDH2 mutants (IDH2R140 and IDH2R172) in myeloid malignancies [85, 86, 91, 95], only IDH2R172 is observed in AITL [98], suggesting that the more efficient production of the oncometabolite 2-HG [85, 86, 91] associated with IDH2R172 may be crucial for T-cell transformation. (iii) There is insignificant or no overlap of IDH2R172-related gene signatures in CMML and AITL [98]. (iv) Mutations in polycomb group (PcG) genes (e.g., EZH2) are rare in AITLs, unlike myeloid proliferative disorders.
The identification of an oncometabolite allows a new consideration in diagnosis, by a noninvasive serum test rather than by histology as recently shown in AML [99]. Recent studies have shown that specific inhibition of IDH2 mutants can reverse the abnormal production of 2-HG and has shown efficacy in preclinical and phase-1 clinical studies of AML. Interestingly, a recent case report showed the efficacy of a hypomethylating agent, 5-azacytidine, in an AITL patient with TET2 mutation [100]. Epigenetic therapies, as single agent therapy, have shown early promise with objective durable response in at least 15–30% of AITL [101]. It would be of interest to see if a combination of epigenetic and PI3 K–mTOR inhibitors could prove to be synergistic.
RHOA has a specific G17 V mutation in up to 70% of AITLs, which exclusively occurs in the background of TET2 mutations with or without IDH2 mutations [92–94]. RHOA, a small GTPase, has long been known to be necessary for T-cell development, activation, and polarization [102, 103]. However, mice with transgenic T-cell-specific expression of a bacterial toxin that inhibits RHOA function develop aggressive thymic lymphomas [104]. A recent study has shown that RhoAG17V knock-in mice resulted in the spontaneous accumulation of Tfh cells in the absence of immunization, and expression of mutant RHOAG17V in hematopoietic progenitors from Tet2 knockout mice resulted in the specific development of AITL-like disease demonstrating an instrumental role for the RHOAG17V mutation in AITL transformation [105]. Unlike AITL, RHOAG17V mutations in ATLL do not co-occur with Tet2 mutation, suggesting a cooperative role of RHOAG17V and Tet2 in skewing differentiation [92]. The RHOAG17V mutant protein cannot bind GDP or GTP but binds strongly to guanine nucleotide exchange factors (GEFs), and the mutant acts as a dominant negative to block wild-type function, presumably by sequestering RHOA GEFs. RHOA mutant may also abnormally recruit VAV1 to the cell membrane where it may be activated. RHOA is involved in numerous intracellular processes, many through activation of two kinases, ROCK1 and −2. An important target of the ROCK kinases is PTEN [106], which is activated and, in turn, down-modulates PI3 K signaling and AKT activity. Consistent with this hypothesis, RHOAG17V mutation results in increased AKT activity [94].
2.10 Anaplastic Large Cell Lymphoma (ALCL)
This entity, initially called Ki-1 lymphoma, is characterized by clusters of large strongly CD30 (Ki-1)-positive anaplastic cells often involving lymph nodes. These lymphomas are predominately of T-cell lineage, but some cases with null cell phenotype and genotype have been described. Two distinct sub-entities differentiated by the expression of anaplastic lymphoma kinase (ALK) protein are recognized, which, except for the expression of ALK protein, have similar morphological and immunophenotypic features. ALK-positive ALCL has a disease-defining genetic translocation involving ALK [107] and most commonly NPM, generating a fusion gene encoding a chimeric protein with constitutive tyrosine kinase activity [108–110] Many additional fusion partners have since been identified, and a recent study identified a novel translocation resulting in the fusion of the TRAF1 and ALK genes, and aggressive clinical course. Genomic studies identified TP53 and PRDM1 loss and the activation of NF-κB signaling in TRAF1-ALK-ALCL. ALK-positive ALCL is more common in children and young adults, accounting for 10–20% of childhood NHL and 3% of adult NHL. The 5-year OS of ALK-positive ALCL (70–86%) is superior to that of ALK-negative ALCL (30–49%). ALK-negative ALCL occurs in older patients, and, when ALCL patients are stratified according to age and/or stage, ALK-positive, and ALK-negative individuals have more similar prognosis. In the current 2016 WHO classification, ALK-negative ALCL is considered as a true entity distinct from PTCL-NOS [2], and other rarer ALCL entities including breast implant-associated ALCL and primary cutaneous ALCL are included in the new version.
GEP studies: Several genome-wide GEP studies definitively demonstrated that ALK-ALCL is a distinct molecular entity, and this lymphoma shows enrichment of a number of signatures including IRF4 and MYC signatures (also see below). Both ALK+ and ALK-negative ALCL demonstrate low expression of genes associated with TCR signaling and high expression of CD30 (TNFRSF8), BATF3 and TMOD1 [111].
In our initial GEP study [12] we identified an ALK + ALCL classifier including TH17 cell-associated molecules (IL-17A, IL-17F, and ROR-γ) and a small group of immunoregulatory cytokines/receptors regulating the STAT3 pathway [112]. These TH17 cell-associated molecules (IL-17F,−A, ROR-γ) were shown to be regulated by miR-135b [113]. In a subsequent larger study, we defined a signature that included all systemic ALCL; the new signature was able to identify 97% of pathologically defined ALCL cases irrespective of ALK status. The signature reclassified 11% (17 of 150) of PTCL-NOS as ALCL, which were all ALK-negative ALCL cases [111] (Fig. 2.4a). Moreover, distinct signatures distinguishing ALK-positive ALCL from ALK-negative ALCL were identified including enrichment of MYC and IRF4 target gene signatures as well proliferation and mTOR gene signatures in ALK-negative ALCL. Although ALK status is prognostic in ALCL, in a recent study, NF-κB signatures could also identify prognostic subgroups in ALCL irrespective of the ALK status [114] (Fig.2.4b).
a NKCLs have a gene expression signature similar to γδ-PTCLs, but distinct from cytotoxic (αβ)-PTCL and Hepatosplenic T-cell Lymphoma (HSTCL) (Adapted from Iqbal et al. Leukemia 2013). b STAT5B mutations are common in γδ T-cell lymphoma (Adapted from Kucuk et al. Nature Communications, 2015). c Whole-exome sequencing (WES) in 25 cases of NKCL and subsequent custom seq in a larger number of cases identified mutations in genes of several categories that include: an RNA helicase, DDX3X (I), tumor suppressors (II), JAK-STAT pathway (III), epigenetic modifiers (IV), other RNA helicases (VI) and other genes (V)
Genetic studies in ALCL: Recent genomic studies have shown recurrent chromosomal rearrangements of genetic loci encoding DUSP22 (30%) and TP63 (8%) in ALK-negative ALCLs, but not in ALK-positive ALCLs and demonstrated that ALK-negative ALCLs have more complex genomes than ALK-positive cases (Fig.2.4c). DUSP22 and TP63 rearrangements were mutually exclusive, and DUSP22 rearranged cases appeared to have very favorable prognosis while TP63 rearranged cases had a very poor outcome. Similarly, TP53 and PRDM1 losses are more common in ALK-negative than in ALK-positive ALCL and may be associated with a more aggressive course [115]. A recent comprehensive genomic study identified frequent activating, occasionally co-occurring mutations of JAK1 and STAT3 genes in ALK-negative ALCLs (38%), resulting in the constitutive activation of the JAK/STAT3 pathway [116]. There are more than 20 sporadic fusion transcripts with fusions involving NFKB2 or NCOR2 being the most common (Fig.2.4d). Some of the common mutations have been shown to activate STAT3. STAT3 activation is also central to ALK-positive ALCL where ALK kinase mediates the activation [117].
MiRNA studies in ALCL: Two studies that aimed to characterize the role of miRNA as downstream effectors of the ALK oncogenic pathway found members of the miR-17-92 clusters to be highly expressed in ALK-positive ALCL whereas miR-155 was expressed at higher levels in ALK-negative ALCL [38, 118]. Expression of the miRNA-17-92 cluster partially rescues STAT3 knockdown by sustaining proliferation and survival of ALK-positive cells. Liu et al. corroborated the high expression of the miR-17-92 cluster in primary ALK-positive ALCL and found a signature of 7 additional miRNAs that could help to distinguish ALK-positive from ALK-negative ALCL cases and an 11-miRNA signature that is useful in differentiating ALK-negative ALCL from other PTCLs. Another study showed that miR-155 was expressed more than tenfold higher in ALK-negative than ALK-positive ALCL, and miR-101 was downregulated in all ALCL model systems [119], but its forced expression attenuated cell proliferation only in ALK-positive but not in ALK-negative cell lines, suggesting different modes of ALK-dependent regulation of its target proteins.
2.11 Adult T-Cell Leukemia/Lymphoma (ATLL)
Adult T-cell leukemia/lymphoma (ATLL) is a rare mature T-cell lymphoma that is etiologically linked to infection with the human T-cell lymphotropic virus type 1 (HTLV-1). HTLV-1 infection is endemic in southwestern Japan, the Caribbean, Central and South America, the Middle East, and tropical Africa. Morphologically, ATLL is characterized by a wide spectrum of morphological appearances including variable patterns of infiltration and cell morphology. Usually, the neoplastic cells show marked nuclear pleomorphism with polylobation giving rise to the characteristic “flower cell” appearance. Clinically, the disease can be divided into the less aggressive smoldering and chronic types and the aggressive acute and lymphomatous types. In Japan, the lifetime risk of ATLL among carriers is 6–7% for males and 2–3% for females, and malignancy typically develops many decades after infection. In addition to gag, pol, and env proteins expressed by other retroviruses, HTLV-1 encodes several other proteins, including Tax, which transcriptionally activates viral transcription and also activates several host signaling pathways, including NF-κB and AP-1 pathways, leading to T-cell proliferation. Tax has transforming activity for T-cells and can induce T-cell lymphomas in transgenic mice [120]. Despite its importance in initial immortalization of infected cells, its expression is almost completely lost in nearly all ATLL cases. The only viral protein that is always persistently expressed is a basic leucine zipper transcription factor, HBZ, encoded by a negative-strand transcript. Its transgenic expression can induce T-cell lymphomas resembling ATLL in mice. It also drives proliferation, inhibits apoptosis, and stimulates transcription of hTERT and several miRNAs such as miR-17 and miR-21 that act to disrupt genomic integrity [121]. It inhibits the acetyltransferase activity of two proteins: KAT7 (HBO1) and CREBBP (CBP) which in addition to modifying histones, also activates p53 by acetylation [122].
GEP studies: ATLL shows extensive abnormalities in gene expression, DNA methylation, and histone modifications and numerous mutations and copy number changes affecting key signaling pathways in T-cells. Gene expression studies have recently showed markedly increased expression of tumor suppressor in lung cancer 1 (TSLC1), CAV1, and prostaglandin D2 synthase [123]. Another study of uncultured lymphocytes from ATLL patients showed increased expression of genes linked to the cell cycle (CDC2, cyclin B), hypercalcemia (RANKL, PTHLH), tyrosine kinase signaling (SYK, LYN) pathways, and anti-apoptosis (BIRC5) [124] and identified BIRC5 as a rational clinical target in the treatment of ATLL. We also identified a unique signature for ATLL with many of the transcripts associated with the Tax viral oncoprotein [72] despite its frequent downregulation at this stage (see also below) or involved in TCR signaling but not an elaborate cytokine profile like AITL. Interestingly, CCR4 expression has been shown to be significantly associated with ATLL [11, 12], and recently, anti-CCR4 (mogamulizumab) was approved for the treatment of ATLL, PTCL-NOS, and CTCL [125, 126]. GSEA revealed the enrichment of TCR signaling genes, target genes of the transcription factor retinoic acid receptor-γ and mature CD4+ T-cell signature but not Treg-related genes.
Genetic studies in ATLL: A recent large integrated study [127] involving 426 patients included mutations (2 whole genome sequencing (WGS), 35 whole-exomes sequencing (WES), 42 both; targeted sequencing, (370), copy number and structural abnormalities (426), DNA methylome (109), and transcriptome (57) analysis. An average of 7.1 mutations per Mb/sample (SNVs, insertions, or deletions) and ~60 structural variants per case were found by WGS. Breakpoints tended to affect tumor suppressor genes and oncogenes relevant to ATLL but more frequently affected known fragile sites with multiple deletions of the NRXN3 locus (14q31.1 fragile site) in 60% of cases. Significant focal CNAs included 26 gains, often high-level amplifications, and 50 losses, many homozygous. A total of 50 genes were found to be significantly mutated (q < 0.1), 13 of them in > 10% of cases. Notably, 13 of these 50 genes lie within the significant CNAs. Many of the significantly altered genes encode proteins in the Tax interactome (p = 8 × 10−18), suggesting that these alterations compensate for the loss of Tax expression during disease evolution.
Driver abnormalities are highly enriched in components of the TCR pathway, the most significantly affected (p = 1.4 × 10−12) and the NF-κB pathway. Known or suspected gain-of-function mutations or focal amplifications predominated in these pathways, the most common being in PLCG1 (36%) and PRKCB (33%), which are the two most commonly mutated genes overall. CARD11 was also frequently mutated (24%) or amplified (12%), and four cases (8% of WGS cases) had focal small internal deletion within the protein’s autoinhibitory domain. Hotspot mutations of CARD11 or PRKCB enhance NF-κB activation, with even further activation when they are co-expressed, a possible explanation of the significant co-occurrence of the two mutations in ATLL. Other changes include hotspot mutations in VAV1 (18%), some of which are known to be activating by altering autoinhibitory domains and in FYN (4%), many in the C-terminal autoinhibitory region. IRF4, a transcription factor constitutively active in ATLL and a target of NF-κB, shows copy number gains in 25% of cases and is mutated (14%) at hotspots within the DNA-binding domain that are predicted to confer a gain of function. Signaling from CD28 is expected to be very frequently enhanced by several mechanisms, including focal gains (71%), which are often high-level amplifications (23%); mutations in hotspots that could increase activity; and tandem duplications leading to expression of CD28 sequences from the CTLA4 or ICOS promoter and occasionally substituting the extracellular domain with that of ICOS or CTLA4; the latter binds CD80 and CD86 more strongly than CD28, resulting in stronger signaling from the retained CD28 intracellular domain.
G-protein-coupled receptors (GPCRs) involved in T-cell trafficking are also commonly mutated. Mutations in CCR4 (29%) and CCR7 (11%) almost always result in truncation of the intracellular C-terminal tail, which prevents internalization upon stimulation with ligand and thereby enhances chemotaxis. CCR4 is typically well expressed in ATLL [128, 129] and likely contributes to the frequent skin involvement in this disease [130].
Other common mutations promote evasion of immune surveillance. Over half (54%) have alterations (copy number losses or inactivating mutations) affecting MHC class I (B2 M and HLA-A and –B), cell adhesion (CD58), cell death (FAS), and immune checkpoint (PDCD1 [PD-1]). Several transcription factors with important roles in T-cells are frequently affected. Strikingly, IKZF2 (Helios) has internal deletions or inversions in 40% of cases analyzed by WGS that generate abnormal short isoforms, previously reported in ATLL [131] that can act in a dominant-negative fashion and can induce T-cell lymphoma in mice [132]. GATA3, a master regulator of TH2 cells, is also frequently deleted (14%) or mutated (15%), with many mutations clearly conferring loss of function. Other transcription factors commonly targeted include CEBFA, ETV6, BCL11B, and ZEB1. STAT3 is mutated in 21% of cases, and activating mutations in NOTCH1 are present in 15% of cases. As in other cancers and PTCL subtypes, TP53 is commonly affected (18% mutation and 23% deletion), as is CDKN2A (2% mutation and 29% deletion). Alterations affecting DNA methylation (TET2, DNMT3A, and IDH2) occur, but at far lower frequencies than in AITL. Despite the low frequency of mutations in TET2 and IDH2 (8% and 1%), ATLL shows prominent CpG island (CGI) hypermethylation, and about 40% of cases showed extensive CGI hypermethylation (CpG island methylator phenotype; CIMP). The CIMP was associated with much poorer overall survival (p = 0.002) [127]. Among the 180 significantly hypermethylated and downregulated genes were HLA-A, −B, −C, and –E. Together with deletions and mutations, about 90% of ATLL cases had loss of MHC class I expression. Earlier studies had shown that DNA methylation, including at CDKN2A, was associated with disease progression, with CIMP most common in the lymphomatous type [133, 134].
Another striking global epigenetic abnormality in ATLL is enhanced Polycomb-dependent repression by trimethylation of histone3 lysine-27 (H3K27me3), affecting half of the genes in ATLL [135]. This is found even at early stages, and, in fact, lentiviral transduction of normal T-cells with Tax leads to enhanced H3K27me3 with 83% overlap between the induced peaks and the peaks found in ATLL in comparison to normal T-cells. EZH2 and other components of the Polycomb repressor complex 2 (PRC2) are considerably upregulated in ATLL, and pharmacological inhibition of EZH2 markedly induced apoptosis in ATLL cell lines [135]. Among the many downregulated genes with enhanced H3K27me3 were important tumor suppressor genes such as BCL2L11 (BIM) and CDKN1A and other genes encoding proteins whose expression is characteristically lost in ATLL such as CD7. Notably, KDM6B, encoding one of the two proteins that reverse H3K27me3 modification, is itself downregulated in association with H3K27me3. This modification may help to lock in the global changes initiated by Tax, resulting in their persistence when Tax expression is lost during disease evolution [135].
2.12 Cutaneous T-Cell Lymphoma (CTCL)
CTCL is heterogeneous, and in this discussion, we will focus on mycosis fungoides (MF) and Sézary syndrome (SS), where recent studies have provided significantly more molecular data. MF is far more common than SS and primarily comprises CD4+ skin-homing memory T-cells, typically with a TH2-like phenotype. The disease can vary from limited cutaneous plaques and patches to skin tumors. MF with limited skin involvement has a generally favorable prognosis, but with skin tumors and high stage disease, median survival is approximately four years. SS, with generalized erythema, lymphadenopathy, and high blood tumor burden, has even worse survival. SS may develop as a progression from MF but usually arises de novo. Most molecular analyses of CTCL have been directed at advanced stages of MF and included SS.
A chronic inflammatory infiltrate appears to have an important pathogenetic role in CTCL, with malignant cells being stimulated through paracrine loops involving malignant cells, inflammatory cells, keratinocytes, and other normal constituents of the skin [136]. There is suggestive but not yet conclusive evidence that TCR stimulation by superantigens [137], in particular, from Staphylococcus aureus, is important in MF. MFs adverse effects on skin barrier function [138, 139] likely contribute to infections that result in exposure to superantigens. Autocrine and paracrine loops involving cytokines appear to be very important in CTCL, but their relative importance appears to vary with the stage of the disease. Even at early stages of MF, autocrine or paracrine IL-15 appears to be important in malignant cell proliferation and survival [140, 141] and may be responsible for constitutive STAT5 activation [142]. ZEB1 binding to sites in the promoter negatively regulates IL-15 transcription. Methylation of the region of the promoter containing these sites [143] may enhance IL-15 expression, as would the frequent deletion or mutation of ZEB1 found in CTCL (see below). Transgenic expression of IL-15 in T-cells in mice leads to a spontaneous CTCL resembling the human disease [143]. In SS, JAK1, JAK2, and STAT3 are consistently constitutively active [144], perhaps due to autocrine IL-21 secretion [145]. There is evidence that several other cytokines function as autocrine or paracrine growth factors in CTCL. These include TSLP [146, 147], IL-13 [148], IL-16 [147], and IL-32 [149, 150] Perhaps due to aberrant STAT5 activation, lymphotoxin-α (LTA) is expressed in CTCL [151] and acts as an autocrine factor through its receptor, TNF receptor 2, encoded by TNFRSF1B, which, as noted below, has gain-of-function mutations and focal copy gains in CTCL.
Gene expression studies in CTCL: Numerous studies have compared the gene expression profiles (GEPs) of various CTCL subtypes to those of normal T-cells, normal skin, and inflammatory skin conditions. SS cells commonly express several genes not normally expressed in T-cells, including TWIST1, PLS3 (plastin 3), and two NK-cell markers, NCR1 (NKp46) and KIR3DL2 [152]. TWIST1 is also expressed in many cases of MF, especially in advanced stages. Aberrant DNA hypomethylation appears to play a role in inappropriate TWIST1 and PLS3 expression. [153] A recent study using RNA-seq compared SS to normal CD4+ T-cells and found a total of 345 transcripts to be significantly upregulated in SS. Gene set enrichment analysis showed the upregulated genes to be enriched in only a few canonical pathways, including cell cycle control, MYC transcriptional activation, immune system regulation, and chemokine signaling. Several genes with critical roles in TCR, cytokine, and chemokine signaling are strongly upregulated (≥ 5x). These include CD3G, RAC2, PRKCQ (PKCθ), and HRAS (TCR signaling); chemokine receptors CCR4 and CCR8; IL6R; and IL2RG. IL2RG encodes the IL-2 receptor common gamma chain, which is a subunit of six cytokine receptors including receptors for IL-15 and IL-21, which are important autocrine factors in CTCL, as described above. Among upregulated transcripts of transcription factors is TOX (~19-fold increase), which has been suggested as a specific biomarker for MF and SS. There is evidence for a pathogenic role for TOX in CTCL by downregulating cell cycle inhibitory proteins CDKN1B and –C [154], as well as RUNX [155], which may act as a tumor suppressor gene in CTCL [156].
Genetic abnormalities: Many early studies identified individual mutated genes and studied copy number abnormalities at low resolution or in a limited number of patients, and demonstrated that CNAs found in CTCL are shared with other PTCLs, especially adult T-cell leukemia/lymphoma (ATLL), which often also shows homing to the skin. In recent years, several studies have performed WES and high-resolution copy number analysis on large numbers of patients [157–160]. The frequencies of the various abnormalities varied among the studies, sometimes considerably, probably due to the considerable heterogeneity of the disease; nevertheless, most of their results were very similar. About three-quarters of single nucleotide variants (SNVs) in most SS cases are C-to-T transitions, which mainly fall into two mutations signatures—one from spontaneous deamination at CpG dinucleotides, associated with age, and a second associated with ultraviolet-B light exposure. The presence of the latter in SS suggests that the disease may originate in skin-resident memory T (TRM) cells. In addition to SNVs, SS cases show large numbers of structural abnormalities. One study [159] of six SS cases by WGS found an average of 168 abnormal junctions per case due to structural alterations. Although no recurring abnormalities were found, 42 potential fusion genes were identified, several of which appeared likely to have a pathogenetic impact. Many of the genes most commonly found deleted or deleteriously mutated in SS are also commonly lost in many other tumors. Loss of 17p occurs in ~40% of cases, and, including small deletions at 17p13.1, TP53 is deleted in 50% or more of SS cases. In addition, mutations are also common (24% in one report). Deletions at 9p21.3 that include CDKN2A occur in ~50% of cases and are often homozygous; mutations also occasionally occur. Loss of CDKN2A and TP53 are expected to confer resistance to apoptosis and cellular senescence and enhance proliferation and genomic instability. Other deletions that appear to be driven by the loss of cell cycle regulators include deletion at 13q14.2 (RB1) and 12p13.1 (CDKN1B). An apoptosis regulator that is commonly deleted or mutated is FAS (10q23.3, which also includes PTEN). Loss of FAS is expected to interfere with activation-induced cell death (AICD), an important homeostatic mechanism in T-cells.
Genes involved in chromatin remodeling are frequently affected. A focused deletion on 1p36.1 includes ARID1A, which also shows a mutation in ~10% of cases. Together, ~40–60% of cases have either deletion or mutation. ARID1A is a targeting subunit of SWI/SNF chromatin-remodeling complexes, which can move nucleosomes along DNA. ARID1A is deleted or mutated in many other tumor types. In a colon cancer model, loss of ARID1A led to lost or decreased H3K27 acetylation and loss of other SWI/SNF components at most enhancers, often with associated decreased expression of the associated genes [161]. Intermediate effects were seen with heterozygous loss. ARID1A also has a role in DNA repair, and its loss sensitizes cells to DNA damage [162], and in deleted cells, inhibitors of the DNA damage checkpoint kinase ATR are synthetically lethal [163]. ARID5B is also frequently (~40%) included in deletions within 10q and ARID3A in deletions of distal 19p [159]. Other SWI/SNF components are also frequently affected by deletions. In particular, SMARCC1 at 3p21.31 was included in deletions of < 4 Mb in 21% of tumors. Deletions or mutations affecting ARID1A, ARID5B, or SMARCC1 occurred in 61% of cases.
Other genes mutated in CTCL also have roles in enhancer function. For example, mutations in MLL2, MLL3, and MLL4 (KDM2D, −C, and B) occur in SS although the frequencies vary considerably among reports; for example, MLL3-mutant SS cases vary between 1 of 25 (4%) [158] to 39 of 66 (59%) [159]. The MLL family catalyzes mono-, di-, and trimethylation of histone H3 lysine 4 and seems to be particularly important in monomethylation at enhancers [164, 165], which is associated with enhancer activity. Other chromatin regulators mutated in SS include BRD9, CHD3, and CREBBP. Alterations affecting DNA methylation are also common in CTCL. Focal deletions at 2p23.3, sometimes homozygous, likely target de novo DNA methyltransferase DNMT3A in ~20% of SS [158]; another report found 37.5% of cases with DNMT3A deletions in CTCL [157]. Deleterious mutations also occur. Mutations and deletions also occur in TET family genes, including TET2 [158].
Another commonly deleted gene with a broad effect on transcription is NCOR1 on 17p, with deletions in almost half of SS cases. Although 17p also includes TP53, NCOR1 mutations, including nonsense and frameshift mutations, were found in 15% of SS cases, often together with deletion of the other allele, suggesting a pathogenetic role [159]. NCOR1 interacts with a large number of transcription factors and recruits histone deacetylases (HDACs) to repress transcription. Several transcription factors have either activating or inactivating alterations. In addition to TP53, one of the most commonly deleted genes is ZEB1, on 10p, which was reported to be deleted in 36% of SS cases and to have mutations in ~10%, half of them frameshifts. ZEB1 is a candidate TSG in ATLL, and ZEB1 KO mice develop CD4 + T-cell lymphoma [166]. ZEB1 is a transcription factor that usually represses, but can activate transcription, and binds to some E-box sites in competition with E2A (TCF3). Among the genes repressed by ZEB1 are BCL6 [167], IL2 [168], and IL15. [143] Gain-of-function mutations in STAT3 and STAT5B and JAK1 and JAK3 are found occasionally in SS, altogether occurring in ~10% of cases. In addition, STAT3, −5A, and −5B are on chromosome 17q, which is very frequently gained in CTCL. Deletions in 10q frequently include NFKB2, which encodes the NF-κB inhibitory subunit p100. In the noncanonical pathway, the C-terminus of p100 is proteolytically removed, yielding a p52 subunit, which, as a heterodimer with RELB, activates a subset of NF-κB target genes. In one study, 10% of CTCLs had 10q deletions causing C-terminal truncation of NFKB2 [157], mimicking natural activation through the noncanonical pathway; such truncations in CTCL had been reported 20 years before [169, 170]. Several other mechanisms contribute to NF-κB activation in CTCL. TCR-induced NF-κB activation involves recruitment and phosphorylation of a CARD11-BCL10-MALT1 complex by PKCθ. Activating mutations in CARD11 are found in CTCL (6% in one study [159]). Focal gains of PRKCQ, encoding PKCθ, are also frequent. Focal deletions, sometimes homozygous, of the negative feedback regulator TNFAIP3 are common [29, 157]. A recurrent point mutation in TNFRSF1B, encoding TNFR2, was identified in 5% of CTCL and SS cases [171]. This mutation was shown to result in enhanced activation of the noncanonical NF-κB pathway. Focal copy gains of the gene also occur.
Other alterations that are expected to have broader effects on signaling pathways include point mutations in CD28 and occasional partial tandem duplications that result in CTLA4-CD28 fusion, resulting in enhanced co-stimulatory signaling. Mutations in PLCG1, encoding phospholipase C γ1, have been reported at various frequencies, often ~10%. Some of the recurring abnormalities have been shown to confer a gain of function. PLCγ1 generates two critically important second messengers, inosine trisphosphate (IP3), which induces calcium release from the ER, and diacylglycerol (DAG), which activates several pathways in T-cells, including the RAS-ERK and NF-κB pathways (Fig. 2.1). Notably, the RAS-ERK pathway can also be activated by occasion gain-of-function mutations in NRAS and KRAS and MAPK1 (ERK2) in CTCL.
miRNA expression in CTCL: Several studies of miRNA expression in CTCL tumor cells from various stages of the disease have been reported [172–174]. Different stages appear to show different patterns of miRNA expression. In SS, the most strikingly upregulated miRNAs are miR-214 and miR-199a-5p and −3p (~50, 40 and tenfold), generated from the same primary RNA, which is upregulated due to aberrant TWIST1 expression. The miRNA cluster has been shown to promote cell survival in CTCL cell lines [175]. MiR-155 is upregulated even in early-stage MF, and miR-21 is upregulated in more advanced stages [143]. Both miRNAs have clear oncogenic functions.
2.13 ENK/T-Cell Lymphomas
NK/T-cell lymphoma comprises 1–2% of all NHL and 10% of PTCL [7]. The vast majority of cases present as an extranodal lymphoma characterized by vascular damage and destruction, prominent necrosis, cytotoxic phenotype, and the consistent presence of the EBV genome in tumor cells. This lymphoma has a higher incidence in Asia [7] and Central and South America and there is an increased incidence in the US due to changes in the demographic of the population [176, 177]. Except for localized nasal or paranasal disease, the prognosis is unfavorable, and most patients succumb to their disease.
Although ENKTCL is strongly associated with EBV, the role of the virus in the etiology of the disease is unclear. In addition, rare cases and NK-cell lines without EBV have been reported [178–182]. The tumor cells typically show a viral latency II pattern with expression of LMP1, LMP2, EBERs, and EBNA1. The expression of LMP1 may be significant as it is the main transforming protein in B-lymphoblastoid cell lines [183] and can activate both the canonical and alternative NF-kB pathways. EBV has been reported to produce some microRNAs from two primary transcripts from the BART and the BHRF1 loci [184, 185]. Only the BART locus associated miRNAs are expressed with type I or II latency, and there is evidence that at least some of these miRNAs promote transformation in B-lymphocytes [186]. A recent report [187] suggests that miR-BART20-5p and miR-BART8 may repress the IFN-STAT1 pathway with decreased expression of TP53 in ENKTCL. The expression profile and implications of these miRNAs in ENKTCL pathogenesis warrant further investigation [188].
GEP studies on ENKTCL identified unique signatures with the majority contributed by the neoplastic cells, and a classifier has been derived (Fig. 2.5a). Interestingly, this classifier also identified a set of γδ-PTCLs both in the ENKTCL and in cases initially classified as PTCL-NOS. These γδ-PTCLs expressed transcripts associated with the T-cell receptor (TCR)/CD3 complex consistent with their T-cell lineage. They were very similar to NK-cell tumors by GEP but were distinct from cytotoxic (αβ)-PTCL indicating derivation from an ontogenically and functionally distinct subset of γδ T-cells (Fig. 2.5). The platelet-derived growth factor receptor (PDGFR) signaling pathway has been reported to be activated in ENKTCL, and the expression of phosphorylated PDGFRα has been confirmed by immunohistochemistry. If this is a clinically important pathway, it is possible that imatinib could be an effective treatment. GEP studies also revealed activation of the NOTCH pathway, and tumor cell lines with NOTCH pathway activation were highly sensitive to γ-secretase inhibitors (Fig. 2.6) [189, 190].Aurora kinase A (AURKA) is frequently upregulated in ENKTCL, and NK-cell lines are highly sensitive to AURKA inhibitors. In addition to being an important regulator of mitosis, AURKA also downregulates the TP53 pathway, with an increase in survivin (BIRC5) expression [13]. The frequent angiocentric and angiodestructive feature of the tumor is expected to produce hypoxia and thereby activation of hypoxia-induced factors (HIFs). Indeed, activation of the HIF1α pathway is observed in GEP studies and may promote tumorigenesis and angiogenesis. These findings provide a rationale for further study of inhibitors to all these pathways as novel therapeutic agents in NK-cell malignancies.
Mutated genes in Hepatosplenic T-cell Lymphoma patients (a). Percentage of cases and types of mutations affected per gene (b). Every box represents the mutation status of a patient for a particular gene and black dots indicate more than one mutation in that gene, with boxes split to show the different types of mutations affecting that gene
Mutations in EATL patients (a) and percentage of cases and types of mutations affected per gene (b). Every box represents the mutation status of a patient for a particular gene and black dots indicate more than one mutation in that gene, with boxes split to show different types of mutations affecting that gene
Genetic studies in ENKTCL: Cytogenetic information on ENKTCL is limited, but some studies on chromosomal copy number changes and loss of heterozygosity (LOH) have been reported [189–194]. A number of frequent gains and losses including gains of 1q21-q44 and losses of 6q21 and 17p11-p13 were detected. The minimal common region of del 6q21 includes PRDM1, ATG5, AIM1, and HACE1. Additional studies have found inactivating mutations affecting PRDM1 in cell lines and infrequently in patient samples. In contrast, DNA methylation that inhibits the transcription of this gene is frequently detected. In vitro functional studies also indicate that PRDM1 is a tumor suppressor gene in ENKTCL [195, 196]. There is also evidence implicating HACE [197]; an E3 ubiquitin ligase reported to be a tumor suppressor gene in other tumors [198]. The 17p deletion includes the TP53 gene which has also been found to be frequently mutated in ENKTCL [199]. It is interesting that EBNA1 has been found to decrease TP53 expression, probably through the interruption of the interaction of USP7 (HAUSP) with TP53. These observations, together with the possibility of decreased TP53 expression due to miR-BART20-5p and 8, indicate that the TP53 pathway is likely to be impaired in most ENKTCL lymphomas through diverse mechanisms [200–202].
Several important genes have also been found to be frequently methylated such as TP73 and SHP1 [203, 204]. TP73 is a member of the TP53 family that may regulate a similar set of target genes as TP53, although it may also have functions other than that of a transactivator. [205, 206]. SHP1 is one of the major negative regulators of NK-cell activation [207] through dephosphorylation of phosphotyrosines in activated receptor/adaptor. In addition, SHP1 also regulates STAT3 activation in several systems although the exact mechanism has not been elucidated [208–212]. Thus, SHP1 loss may contribute, similar to the loss of SOSC6 and other negative regulators, to the aberrant activation of the JAK/STAT3 pathway (see below). By genome-wide methylation analysis, we found frequent prominent global hypermethylation in ENKTCL and detected additional potentially important methylated loci, including some that have been previously reported to be methylated or inactivated by mutation in other tumors or are known tumor suppressor genes. These include BIM, a crucial pro-apoptotic BH3-only protein; SOCS6, an inhibitor of the JAK/STAT pathway; DDX3X; and DAPK1 [43]. One interesting finding is the frequent methylation of ASNS, encoding asparagine synthetase. Its inactivation may explain the sensitivity of the tumor cells to asparaginase treatment [213].
FAS have been reported to be mutated or deleted in ENKTCL [214, 215] which may help to overcome the fratricidal function of NK-cells. Also, survivin (BIRC5), an inhibitor of apoptosis, has been reported to be overexpressed in ENKTCL, probably through TP53 inactivation. These changes, together with BIM methylation and upregulation of BCLXL and BCL2 through STAT3 or STAT5B activation, represent some of the mechanisms for ENKTCL to resist apoptosis.
The STAT3 pathway is frequently activated in ENKTCL. This may be achieved through autocrine/paracrine activation of STAT3 via cytokine receptors [216]. Inactivation of negative regulators may play a role as noted above; however, STAT3 mutations affecting the SH2 domain are one of the most frequent mutations (Fig. 2.5b) [213]. These mutations may prolong and enhance the activation of STAT3, which may be particularly relevant when cytokine concentrations are suboptimal [213]. These STAT3 mutants may also promote tumor development through the production of tumor-promoting inflammation and an immunosuppressive tumor microenvironment as a result of the production of cytokines such as VEGF and IL-10 [217–219]. There were two reports of frequent JAK3 mutations in ENKTCL [220, 221]. However, other studies failed to confirm such a high frequency [213, 222]. As STAT3 is activated by JAK kinases, JAK inhibitors are expected to suppress tumor proliferation/survival, and this is indeed observed both for wild-type and mutant cell lines [213].
A recent mutation analysis using whole exome sequencing and subsequently more focused sequencing has described the mutational landscape of ENKTCL [223] (Fig. 2.5c). Interestingly, the most frequent mutations involved DDX3X, an RNA helicase that has been found to be recurrently methylated as well [43]. The mutants have reduced RNA helicase activity, but the precise roles of DDX3X in the pathogenesis of ENKTCL need further investigation. This study also confirmed the frequent TP53 and STAT3 mutations and reported mutations affecting epigenetic modifiers. Two other studies have reported frequent inactivating BCOR mutations [224, 225].
Hepatosplenic T-cell lymphoma (HSTL): HSTL is a rare entity mostly derived from γδ T-cells, and the disease is usually fatal. It occurs predominantly in young male adults and may arise in the setting of chronic immune suppression such as in solid organ transplant recipients or patients treated with azathioprine and infliximab for Crohn’s disease. Morphologically, HSTL typically presents with intermediate-sized T-cells that preferentially infiltrate the liver, spleen, and bone marrow in a sinusoidal pattern. CD4 and CD8 are frequently absent in the gamma–delta expressing form of the tumor. GEP studies on HSTL demonstrated that irrespective of TCR cell lineage, these lymphomas share unique gene signatures and in general form a distinct hierarchical cluster, but with closer association with ENKTCL than other αβPTCLs (Fig.2.3) [13]. The differentially expressed transcripts reflected the characteristic anatomical (liver and spleen) distribution of the neoplastic cells, with high expression of metabolic and B-cell-related genes, mainly from stromal components. Overall, there were differences in the expression of cytotoxicity genes compared with NKCL and also other non-HS γδ PTCLs [226]. No significant differential expression of KIR or KLR family members was observed between HSTCL and other γδ-PTCL in this study. [226] Genomic profiles confirm frequent recurrent isochromosome 7q and trisomy 8 in HSTL [227]. Despite the rarity of this disease, a comprehensive genomic study of HSTL identified frequent mutations in chromatin-modifying genes (SETD2, INO80, and ARID1B) affecting 62% of cases, and also in STAT5B (31%), STAT3 (9%), and PIK3CD (9%). This study also demonstrated that SETD2 acts as a tumor suppressor gene in HSTCL [44] (Fig. 2.5).
Intestinal T-cell lymphoma: Enteropathy-associated T-cell lymphoma (EATL) defines a primary intestinal tumor derived from intestinal intraepithelial lymphocytes (IEL), with a variable abundance of large lymphoid cells. It accounts for approximately 5% of PTCL. The neoplastic cells express the mucosal homing receptor CD103 characteristic of IEL and are usually CD3+ CD5− CD4− CD8− αβ T-cells with an activated cytotoxic profile and frequent co-expression of CD30. This tumor is associated with gluten sensitivity and enteropathy (celiac disease) and is, therefore, prevalent in people of Northern European descent. Type II EATL, characterized by a monomorphic proliferation of medium-sized cells, is more prevalent in Asia, and is not associated with celiac disease. The neoplastic cells are usually CD3+ CD5− CD7+ CD4−, CD8+ CD56+, and more commonly CD20+. The majority of the cases are of γδ̣ origin and about a third of the cases have mutation of STAT5B similar to HSTCL [213]. Type II EATL has been reclassified as monomorphic epitheliotropic intestinal T-cell lymphoma (MEITL). A recent genetic study identified SETD2 as the most frequently (32%) mutated gene in EATL (32% of cases) and the JAK-STAT pathway as the most frequently mutated pathway, with frequent mutations in STAT5B, JAK1, JAK3, STAT3, and SOCS1. EATL and MEITL shared mutations in KRAS, TP53, and TERT and had overlapping genetic alterations indicating shared mechanisms underlying their pathogenesis [228] (Fig. 2.6).
2.14 The Expanding Scope of Genomics for Therapeutic Opportunities in PTCLs
Recent genomics studies have significantly improved our understanding of the pathogenesis and biology of the major PTCL entities and even some rare ones such as HSTCL and EATL [35]. More and more agents that specifically target biologic pathways are now becoming available for clinical studies. It is possible to rationally design clinical trials in which treatment cohorts are stratified according to molecular profiling defined criteria. This approach will test the feasibility of personalized treatment based on agents specifically targeting genetic abnormalities or oncogenic pathways found in the tumor.
GEP studies can help elucidate oncogenic pathways in different types of cancers [229, 230] which may suggest novel therapeutic targets. An example of this comes from the profiling studies of DLBCL, in which the ABC and PMBL subgroups of DLBCL showed an increased expression of target genes of the transcription factor NF-κB [229, 231, 232]. Davis et al. [229] demonstrated constitutive activation of NF-κB in ABC-derived cell lines, which underwent apoptosis when the NF-κB pathway was inhibited. Several NF-κB inhibitors are currently available and could be tested in clinical trials. Bortezomib (a proteasome inhibitor that also has NF-κB inhibitory activity) has demonstrated some early promise in the treatment of PTCL [233]. Another known drug for this pathway, CEP-18770, has a strong antiangiogenic activity and potently represses RANKL-induced osteoclastogenesis [234]. We have recently demonstrated differential NF-κB activation in T-cell lymphomas, particularly in AITL [72], and this class of drugs may be particularly effective in AITL. Since the microenvironment and angiogenesis are important particularly in AITL, lenalidomide, an angiogenesis inhibitor and immune modulator may be a potential agent for future trials. Clinical grade epigenetic modifiers are being tested in several diseases and could potentially prove synergistic with other drugs in AITL.
Another possible target is SYK, a receptor-associated tyrosine kinase which is expressed in 94% of all PTCL [235]. In vitro studies have demonstrated the efficacy of a small molecule SYK inhibitor (R402) in cell lines, which induced apoptosis and growth inhibition of the cells [236]. PDGFRA is a member of the vascular endothelial growth factor (VEGF) receptor family, and a recent study indicates a high level of PDGFR subunit expression in PTCL-NOS cases, including downstream activation of STAT1, STAT3, and STAT5 transcription factors in primary tumor samples [237]. Notably, PDGFR interacts with the CD3/TCR and CD28 signaling through SH2-binding proteins such as GRB2 and their downstream targets (reviewed in [238]).
The JAK/STAT3 or −5B pathways are often aberrantly activated in PTCL and ENKTCL and are a potential target for therapy. JAK1/2 and JAK3 inhibitors are already in the clinics for myeloid disorders and may be tested for their efficacy in diseases with JAK/STAT activation. Some of the mutations may aberrantly activate the TCR signaling pathway such as the frequent gain-of-function mutations in PLCγ1, which is important for calcium flux and RAS pathway activation. Mutations or other changes affecting VAV1, CD28, PI3 K/AKT/mTOR, and occasionally the RAS pathway are also observed and potentially targetable. We are entering an exciting era with tremendous potential to improve the outcome of patients with T- and NK-cell lymphomas employing single agents or combinations of targeted agents. Further studies are needed to improve our understanding of these rather uncommon tumors so that even more effective and precise targets can be identified for future clinical trials. As these diseases are uncommon or even rare, national or international collaborative efforts are critical to advance our objectives.
References
Rudiger T, Weisenburger DD, Anderson JR et al (2002) Peripheral T-cell lymphoma (excluding anaplastic large-cell lymphoma): results from the Non-Hodgkin’s Lymphoma Classification Project. Ann Oncol 13(1):140–149
Swerdlow SH, Campo E, Pileri SA et al (2016) The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 127(20):2375–2390
Swerdlow SH, Campo E, Harris NL et al (2008) WHO classification of Tumours of Haematopoietic and Lymphoid Tissues, 4th edn
Bellei M, Chiattone CS, Luminari S et al (2012) T-cell lymphomas in South america and europe. Rev Bras Hematol Hemoter. 34(1):42–47
Arora N, Manipadam MT, Nair S (2013) Frequency and distribution of lymphoma types in a tertiary care hospital in South India: analysis of 5115 cases using the World Health Organization 2008 classification and comparison with world literature. Leuk Lymphoma 54(5):1004–1011
Perry AM, Diebold J, Nathwani BN et al (2016) Non-Hodgkin lymphoma in the Far East: review of 730 cases from the international non-Hodgkin lymphoma classification project. Ann Hematol 95(2):245–251
International peripheral T-cell and natural killer/t-cell lymphoma study (2008) pathology findings and clinical outcomes. J Clin Oncol 26(25):4124–4130
Perry AM, Diebold J, Nathwani BN et al (2016) Non-Hodgkin lymphoma in the developing world: review of 4539 cases from the International Non-Hodgkin Lymphoma Classification Project. Haematologica 101(10):1244–1250
Adams SV, Newcomb PA, Shustov AR (2016) Racial Patterns of Peripheral T-Cell Lymphoma Incidence and Survival in the United States. J Clin Oncol 34(9):963–971
Wang. SS, Vose. JM (2103) Epidemiology and Prognosis of T-Cell Lymphoma. Springer Science, New York
Iqbal J, Wright G, Wang C et al (2014) Gene expression signatures delineate biological and prognostic subgroups in peripheral T-cell lymphoma. Blood 123(19):2915–2923
Iqbal J, Weisenburger DD, Greiner TC et al (2010) Molecular signatures to improve diagnosis in peripheral T-cell lymphoma and prognostication in angioimmunoblastic T-cell lymphoma. Blood 115(5):1026–1036
Iqbal J, Weisenburger DD, Chowdhury A et al (2011) Natural killer cell lymphoma shares strikingly similar molecular features with a group of non-hepatosplenic gammadelta T-cell lymphoma and is highly sensitive to a novel aurora kinase A inhibitor in vitro. Leukemia 25(2):348–358
Xu B, Liu P (2014) No survival improvement for patients with angioimmunoblastic T-cell lymphoma over the past two decades: a population-based study of 1207 cases. PLoS ONE 9(3):e92585
Croziera JA,, Shera T, Yangb D et al (2015) Persistent disparities among patients with T-cell Non-Hodgkin Lymphomas and B-cell Diffuse Large Cell Lymphomas over 40 years: a seer database review. Clin Lymphoma Myeloma Leukemia
Briski R, Feldman AL, Bailey NG et al (2014) The role of front-line anthracycline-containing chemotherapy regimens in peripheral T-cell lymphomas. Blood Cancer J. 4:e214
Swerdlow SH, Elias Campo, Harris NL et al (2017) WHO classification of Tumours of the Haematopoitic and Lymphoid Tissues. Lyon, IARC Press, France
Germain RN (2002) T-cell development and the CD4-CD8 lineage decision. Nat Rev Immunol 2(5):309–322
Martin CH, Aifantis I, Scimone ML et al (2003) Efficient thymic immigration of B220+ lymphoid-restricted bone marrow cells with T precursor potential. Nat Immunol 4(9):866–873
Poltorak M, Meinert I, Stone JC, Schraven B, Simeoni L (2014) Sos1 regulates sustained TCR-mediated Erk activation. Eur J Immunol 44(5):1535–1540
Filipp D, Zhang J, Leung BL et al (2003) Regulation of Fyn through translocation of activated Lck into lipid rafts. J Exp Med 197(9):1221–1227
Sugie K, Jeon MS, Grey HM (2004) Activation of naive CD4 T cells by anti-CD3 reveals an important role for Fyn in Lck-mediated signaling. Proc Natl Acad Sci U S A. 101(41):14859–14864
Cannons JL, Yu LJ, Hill B et al (2004) SAP regulates T(H)2 differentiation and PKC-theta-mediated activation of NF-kappaB1. Immunity 21(5):693–706
Cannons JL, Qi H, Lu KT et al (2010) Optimal germinal center responses require a multistage T cell: B cell adhesion process involving integrins, SLAM-associated protein, and CD84. Immunity 32(2):253–265
Latour S, Roncagalli R, Chen R et al (2003) Binding of SAP SH2 domain to FynT SH3 domain reveals a novel mechanism of receptor signalling in immune regulation. Nat Cell Biol 5(2):149–154
Yasuda K, Nagafuku M, Shima T et al (2002) Cutting edge: Fyn is essential for tyrosine phosphorylation of Csk-binding protein/phosphoprotein associated with glycolipid-enriched microdomains in lipid rafts in resting T cells. J Immunol. 169(6):2813–2817
Kong KF, Yokosuka T, Canonigo-Balancio AJ, Isakov N, Saito T, Altman A (2011) A motif in the V3 domain of the kinase PKC-theta determines its localization in the immunological synapse and functions in T cells via association with CD28. Nat Immunol 12(11):1105–1112
Kang JA, Choi H, Yang T, Cho SK, Park ZY, Park SG (2017) PKCtheta-Mediated PDK1 Phosphorylation Enhances T Cell Activation by Increasing PDK1 Stability. Mol Cells 40(1):37–44
Qiao Q, Yang C, Zheng C et al (2013) Structural architecture of the CARMA1/Bcl10/MALT1 signalosome: nucleation-induced filamentous assembly. Mol Cell 51(6):766–779
Isakov N, Altman A (2002) Protein kinase C(theta) in T cell activation. Annu Rev Immunol 20:761–794
Gazzola A, Mannu C, Rossi M et al (2014) The evolution of clonality testing in the diagnosis and monitoring of hematological malignancies. Ther Adv Hematol. 5(2):35–47
van Dongen JJ, Langerak AW, Bruggemann M et al (2003) Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17(12):2257–2317
Szczepanski T, van der Velden VH, Raff T et al (2003) Comparative analysis of T-cell receptor gene rearrangements at diagnosis and relapse of T-cell acute lymphoblastic leukemia (T-ALL) shows high stability of clonal markers for monitoring of minimal residual disease and reveals the occurrence of second T-ALL. Leukemia 17(11):2149–2156
Iqbal J, Naushad H, Bi C et al (2016) Genomic signatures in B-cell lymphoma: How can these improve precision in diagnosis and inform prognosis? Blood Rev 30(2):73–88
Iqbal J, Wilcox R, Naushad H et al (2016) Genomic signatures in T-cell lymphoma: How can these improve precision in diagnosis and inform prognosis? Blood Rev 30(2):89–100
Iqbal J, Shen Y, Huang X et al (2015) Global microRNA expression profiling uncovers molecular markers for classification and prognosis in aggressive B-cell lymphoma. Blood 125(7):1137–1145
Iqbal J, Shen Y, Liu Y et al (2012) Genome-wide miRNA profiling of mantle cell lymphoma reveals a distinct subgroup with poor prognosis. Blood 119(21):4939–4948
Liu C, Iqbal J, Teruya-Feldstein J et al (2013) MicroRNA expression profiling identifies molecular signatures associated with anaplastic large cell lymphoma. Blood 122(12):2083–2092
Bouska A, McKeithan TW, Deffenbacher KE et al (2014) Genome-wide copy-number analyses reveal genomic abnormalities involved in transformation of follicular lymphoma. Blood 123(11):1681–1690
Bouska A, Zhang W, Gong Q et al (2017) Combined copy number and mutation analysis identifies oncogenic pathways associated with transformation of follicular lymphoma. Leukemia 31(1):83–91
Cairns RA, Iqbal J, Lemonnier F et al (2012) IDH2 mutations are frequent in angioimmunoblastic T-cell lymphoma. Blood 119(8):1901–1903
Guo S, Chan JK, Iqbal J et al (2014) EZH2 mutations in follicular lymphoma from different ethnic groups and associated gene expression alterations. Clin Cancer Res 20(12):3078–3086
Kucuk C, Hu X, Jiang B et al (2015) Global promoter methylation analysis reveals novel candidate tumor suppressor genes in natural killer cell lymphoma. Clin Cancer Res 21(7):1699–1711
McKinney M, Moffitt AB, Gaulard P et al (2017) The Genetic Basis of Hepatosplenic T-cell Lymphoma. Cancer Discov 7(4):369–379
Laurent C et al (2017) J Clinic Oncol 35(18):2008–2017
Weisenburger et al (2011) Blood, 117:3402–3408
Bowen et al (2014) Bristish J Hematol 166:202–208
de Leval L, Rickman DS, Thielen C et al (2007) The gene expression profile of nodal peripheral T-cell lymphoma demonstrates a molecular link between angioimmunoblastic T-cell lymphoma (AITL) and follicular helper T (TFH) cells. Blood 109(11):4952–4963
Martinez-Delgado B (2006) Peripheral T-cell lymphoma gene expression profiles. Hematol Oncol 24(3):113–119
Piccaluga PP, Agostinelli C, Califano A et al (2007) Gene expression analysis of peripheral T cell lymphoma, unspecified, reveals distinct profiles and new potential therapeutic targets. J Clin Invest. 117(3):823–834
Cuadros M, Dave SS, Jaffe ES et al (2007) Identification of a proliferation signature related to survival in nodal peripheral T-cell lymphomas. J Clin Oncol 25(22):3321–3329
Martinez-Delgado B, Cuadros M, Honrado E et al (2005) Differential expression of NF-kappaB pathway genes among peripheral T-cell lymphomas. Leukemia 19(12):2254–2263
Piccaluga PP, Fuligni F, De Leo A et al (2013) Molecular profiling improves classification and prognostication of nodal peripheral T-cell lymphomas: results of a phase III diagnostic accuracy study. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 31(24):3019–3025
Ballester B, Ramuz O, Gisselbrecht C et al (2006) Gene expression profiling identifies molecular subgroups among nodal peripheral T-cell lymphomas. Oncogene 25(10):1560–1570
O’Shea JJ, Paul WE (2010) Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science 327(5969):1098–1102
Wang T, Feldman AL, Wada DA et al (2014) GATA-3 expression identifies a high-risk subset of PTCL, NOS with distinct molecular and clinical features. Blood 123(19):3007–3015
Heavican TB, Yu J, Bouska A, Greiner TC, Lachel CM, Wang C, Dave BJ, Amador CC, Fu K, Vose JM, Weisenburger DD, Gascoyne RD, Hartmann S, Pedersen MBJ, Wilcox R, Teh BT, Lim ST, Ong CK, Seto M, Berger F, Rosenwald A, Ott G, Campo E, Rimsza LM, Jaffe ES, Braziel RM, d’Amore FA, Inghirami G, Bertoni F, Staudt L, McKeithan TW, Pileri SA, Chan WC, Iqbal J (2016) Molecular subgroups of peripheral T-cell lymphoma evolve by distinct genetic pathways. In: 58th ASH Annual Meeting and Exposition, San Diego, CA
Schatz JH, Horwitz SM, Teruya-Feldstein J et al (2015) Targeted mutational profiling of peripheral T-cell lymphoma not otherwise specified highlights new mechanisms in a heterogeneous pathogenesis. Leukemia 29(1):237–241
Abate F, da Silva-Almeida AC, Zairis S et al (2017) Activating mutations and translocations in the guanine exchange factor VAV1 in peripheral T-cell lymphomas. Proc Natl Acad Sci U S A. 114(4):764–769
Yoo HY, Sung MK, Lee SH et al (2014) A recurrent inactivating mutation in RHOA GTPase in angioimmunoblastic T cell lymphoma. Nat Genet 46(4):371–375
Laginestra MA, Piccaluga PP, Fuligni F et al (2014) Pathogenetic and diagnostic significance of microRNA deregulation in peripheral T-cell lymphoma not otherwise specified. Blood Cancer J. 4:259
Dobay MP, Lemonnier F, Missiaglia E et al (2017) Integrative clinicopathological and molecular analyses of angioimmunoblastic T-cell lymphoma and other nodal lymphomas of follicular helper T-cell origin. Haematologica 102(4):e148–e151
Piccaluga et al (2007) Cancer Res 15, 67(22):10703–10710
Crotty S (2014) Immunity 41(4):529–542
Fazilleau N, McHeyzer-Williams LJ, Rosen H, McHeyzer-Williams MG (2009) The function of follicular helper T cells is regulated by the strength of T cell antigen receptor binding. Nat Immunol 10(4):375–384
Hatzi K, Nance JP, Kroenke MA et al (2015) BCL6 orchestrates Tfh cell differentiation via multiple distinct mechanisms. J Exp Med 212(4):539–553
Dupuis J, Boye K, Martin N et al (2006) Expression of CXCL13 by neoplastic cells in angioimmunoblastic T-cell lymphoma (AITL): a new diagnostic marker providing evidence that AITL derives from follicular helper T cells. Am J Surg Pathol 30(4):490–494
Miysohi et al (2012) Am J Clin Pathol 137(6):879–89
Bisig B, Thielen C, Herens C et al (2012) c-Maf expression in angioimmunoblastic T-cell lymphoma reflects follicular helper T-cell derivation rather than oncogenesis. Histopathology 60(2):371–376
Murakami YI, Yatabe Y, Sakaguchi T et al (2007) c-Maf expression in angioimmunoblastic T-cell lymphoma. Am J Surg Pathol 31(11):1695–1702
Iqbal J et al (2007) Leukemia 21(11):2332–2343
Iqbal J, Weisenburger DD, Greiner TC et al (2010) Molecular signatures to improve diagnosis in peripheral T-cell lymphoma and prognostication in angioimmunoblastic T-cell lymphoma. Blood 115(5):1026–1036
Grogg KL, Attygalle AD, Macon WR, Remstein ED, Kurtin PJ, Dogan A (2005) Angioimmunoblastic T-cell lymphoma: a neoplasm of germinal-center T-helper cells? Blood 106(4):1501–1502
Schlegelberger B, Himmler A, Godde E, Grote W, Feller AC, Lennert K (1994) Cytogenetic findings in peripheral T-cell lymphomas as a basis for distinguishing low-grade and high-grade lymphomas. Blood 83(2):505–511
Nelson M, Horsman DE, Weisenburger DD et al (2008) Cytogenetic abnormalities and clinical correlations in peripheral T-cell lymphoma. Br J Haematol 141(4):461–469
Lepretre S, Buchonnet G, Stamatoullas A et al (2000) Chromosome abnormalities in peripheral T-cell lymphoma. Cancer Genet Cytogenet 117(1):71–79
Lakkala-Paranko T, Franssila K, Lappalainen K et al (1987) Chromosome abnormalities in peripheral T-cell lymphoma. Br J Haematol 66(4):451–460
Lemonnier F, Couronne L, Parrens M et al (2012) Recurrent TET2 mutations in peripheral T-cell lymphomas correlate with TFH-like features and adverse clinical parameters. Blood 120(7):1466–1469
Sakata-Yanagimoto M, Enami T, Yoshida K et al (2014) Somatic RHOA mutation in angioimmunoblastic T cell lymphoma. Nat Genet 46(2):171–175
Odejide O, Weigert O, Lane AA et al (2014) A targeted mutational landscape of angioimmunoblastic T-cell lymphoma. Blood 123(9):1293–1296
Dawlaty MM, Ganz K, Powell BE, et al Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9(2):166–175
Dawlaty MM, Breiling A, Le T, et al. Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev Cell 24(3):310–323
Wu H, Zhang Y (2011) Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev 25(23):2436–2452
Hill PW, Amouroux R, Hajkova P (2014) DNA demethylation, Tet proteins and 5-hydroxymethylcytosine in epigenetic reprogramming: an emerging complex story. Genomics 104(5):324–333
Losman JA, Looper RE, Koivunen P et al (2013) (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 339(6127):1621–1625
Koivunen P, Lee S, Duncan CG et al (2012) Transformation by the (R)-enantiomer of 2-hydroxyglutarate linked to EGLN activation. Nature 483(7390):484–488
Li Z, Cai X, Cai CL et al (2011) Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118(17):4509–4518
Sasaki M, Knobbe CB, Munger JC et al (2012) IDH1(R132H) mutation increases murine haematopoietic progenitors and alters epigenetics. Nature 488(7413):656–659
Akbay EA, Moslehi J, Christensen CL et al (2014) D-2-hydroxyglutarate produced by mutant IDH2 causes cardiomyopathy and neurodegeneration in mice. Genes Dev 28(5):479–490
Chen C, Liu Y, Lu C et al (2013) Cancer-associated IDH2 mutants drive an acute myeloid leukemia that is susceptible to Brd4 inhibition. Genes Dev 27(18):1974–1985
Xu W, Yang H, Liu Y et al Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell 19(1):17–30
Palomero T, Couronne L, Khiabanian H et al Recurrent mutations in epigenetic regulators, RHOA and FYN kinase in peripheral T cell lymphomas. Nature genetics 46(2):166–170
Nagata Y, Kontani K, Enami T et al (2016) Variegated RHOA mutations in adult T-cell leukemia/lymphoma. Blood 127(5):596–604
Yoo HY, Sung MK, Lee SH, et al A recurrent inactivating mutation in RHOA GTPase in angioimmunoblastic T cell lymphoma. Nature genetics 46(4):371–375
Abdel-Wahab O, Levine RL (2013) Mutations in epigenetic modifiers in the pathogenesis and therapy of acute myeloid leukemia. Blood 121(18):3563–3572
Rohr J, Guo S, Hu D et al (2014) CD28 Mutations in Peripheral T-Cell Lymphomagenesis and Progression. Blood 124(21):1681–1681
Rohr J, Guo S, Huo J et al (2015) Recurrent activating mutations of CD28 in peripheral T-cell lymphomas. Leukemia
Wang C, McKeithan TW, Gong Q et al (2015) IDH2R172 mutations define a unique subgroup of patients with angioimmunoblastic T-cell lymphoma. Blood 126(15):1741–1752
DiNardo CD, Propert KJ, Loren AW et al Serum 2-hydroxyglutarate levels predict isocitrate dehydrogenase mutations and clinical outcome in acute myeloid leukemia. Blood 121(24):4917–4924
Cheminant M, Bruneau J, Kosmider O et al (2014) Efficacy of 5-Azacytidine in a TET2 mutated angioimmunoblastic T cell lymphoma. Br J Haematol
Pro B, Horwitz SM, Prince HM et al (2016) Romidepsin induces durable responses in patients with relapsed or refractory angioimmunoblastic T-cell lymphoma. Hematol Oncol
Borroto A, Gil D, Delgado P et al (2000) Rho regulates T cell receptor ITAM-induced lymphocyte spreading in an integrin-independent manner. Eur J Immunol 30(12):3403–3410
Rougerie P, Delon J Rho GTPases: masters of T lymphocyte migration and activation. Immunol Lett 142(1–2):1-13
Cleverley SC, Costello PS, Henning SW, Cantrell DA (2000) Loss of Rho function in the thymus is accompanied by the development of thymic lymphoma. Oncogene 19(1):13–20
Cortes JR, Ambesi-Impiombato A, Couronne L, Kim CS, West Z, Belver L, da Silva Almeida AC, Bhagat G, Bernard OA, Ferrando AA, PalomeroT (2016) Role and Mechanisms of Rhoa G17 V in the Pathogenesis of AITL. ASH Meeting, San Diego
Li Z, Dong X, Wang Z et al (2005) Regulation of PTEN by Rho small GTPases. Nat Cell Biol 7(4):399–404
Morris SW, Kirstein MN, Valentine MB et al (1995) Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin’s lymphoma. Science 267(5196):316–317
Mason DY, Bastard C, Rimokh R et al (1990) CD30-positive large cell lymphomas (‘Ki-1 lymphoma’) are associated with a chromosomal translocation involving 5q35. Br J Haematol 74(2):161–168
Morris SW, Kirstein MN, Valentine MB et al (1994) Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin’s lymphoma. Science 263(5151):1281–1284
Rimokh R, Magaud JP, Berger F et al (1989) A translocation involving a specific breakpoint (q35) on chromosome 5 is characteristic of anaplastic large cell lymphoma (‘Ki-1 lymphoma’). Br J Haematol 71(1):31–36
Agnelli L, Mereu E, Pellegrino E et al (2012) Identification of a 3-gene model as a powerful diagnostic tool for the recognition of ALK-negative anaplastic large-cell lymphoma. Blood 120(6):1274–1281
Piva R, Pellegrino E, Mattioli M et al (2006) Functional validation of the anaplastic lymphoma kinase signature identifies CEBPB and BCL2A1 as critical target genes. J Clin Invest. 116(12):3171–3182
Matsuyama H, Suzuki HI, Nishimori H et al (2011) miR-135b mediates NPM-ALK-driven oncogenicity and renders IL-17-producing immunophenotype to anaplastic large cell lymphoma. Blood 118(26):6881–6892
Abate F, Todaro M, van der Krogt JA et al (2014) A novel patient-derived tumorgraft model with TRAF1-ALK anaplastic large-cell lymphoma translocation. Leukemia
Parrilla Castellar ER, Jaffe ES, Said JW et al (2014) ALK-negative anaplastic large cell lymphoma is a genetically heterogeneous disease with widely disparate clinical outcomes. Blood 124(9):1473–1480
Crescenzo R, Abate F, Lasorsa E et al (2015) Convergent mutations and kinase fusions lead to oncogenic STAT3 activation in anaplastic large cell lymphoma. Cancer Cell 27(4):516–532
Chiarle R, Voena C, Ambrogio C, Piva R, Inghirami G (2008) The anaplastic lymphoma kinase in the pathogenesis of cancer. Nat Rev Cancer 8(1):11–23
Spaccarotella E, Pellegrino E, Ferracin M et al (2014) STAT3-mediated activation of microRNA cluster 17–92 promotes proliferation and survival of ALK-positive anaplastic large cell lymphoma. Haematologica 99(1):116–124
Merkel O, Hamacher F, Laimer D et al (2010) Identification of differential and functionally active miRNAs in both anaplastic lymphoma kinase (ALK)+ and ALK- anaplastic large-cell lymphoma. Proc Natl Acad Sci U S A. 107(37):16228–16233
Portis T, Grossman WJ, Harding JC, Hess JL, Ratner L (2001) Analysis of p53 inactivation in a human T-cell leukemia virus type 1 Tax transgenic mouse model. J Virol 75(5):2185–2193
Vernin C, Thenoz M, Pinatel C et al (2014) HTLV-1 bZIP factor HBZ promotes cell proliferation and genetic instability by activating OncomiRs. Cancer Res 74(21):6082–6093
Wright DG, Marchal C, Hoang K et al (2016) Human T-cell leukemia virus type-1-encoded protein HBZ represses p53 function by inhibiting the acetyltransferase activity of p300/CBP and HBO1. Oncotarget. 7(2):1687–1706
Sasaki H, Nishikata I, Shiraga T et al (2005) Overexpression of a cell adhesion molecule, TSLC1, as a possible molecular marker for acute-type adult T-cell leukemia. Blood 105(3):1204–1213
Pise-Masison CA, Radonovich M, Dohoney K et al (2009) Gene expression profiling of ATL patients: compilation of disease related genes and evidence for TCF-4 involvement in BIRC5 gene expression and cell viability. Blood
Zinzani et al (2016) Haematologica 101(10):e407–410
Ogura M et al (2014) J Clin Oncol 10, 32(11):1157–1163
Kataoka K, Nagata Y, Kitanaka A et al (2015) Integrated molecular analysis of adult T cell leukemia/lymphoma. Nat Genet 47(11):1304–1315
Yoshie O, Fujisawa R, Nakayama T et al (2002) Frequent expression of CCR4 in adult T-cell leukemia and human T-cell leukemia virus type 1-transformed T cells. Blood 99(5):1505–1511
Harasawa H, Yamada Y, Hieshima K et al (2006) Survey of chemokine receptor expression reveals frequent co-expression of skin-homing CCR4 and CCR10 in adult T-cell leukemia/lymphoma. Leuk Lymphoma 47(10):2163–2173
Li J, Lu E, Yi T, Cyster JG (2016) EBI2 augments Tfh cell fate by promoting interaction with IL-2-quenching dendritic cells. Nature 533(7601):110–114
Fujii K, Ishimaru F, Nakase K et al (2003) Over-expression of short isoforms of Helios in patients with adult T-cell leukaemia/lymphoma. Br J Haematol 120(6):986–989
Zhang Z, Swindle CS, Bates JT, Ko R, Cotta CV, Klug CA (2007) Expression of a non-DNA-binding isoform of Helios induces T-cell lymphoma in mice. Blood 109(5):2190–2197
Sato H, Oka T, Shinnou Y et al (2010) Multi-step aberrant CpG island hyper-methylation is associated with the progression of adult T-cell leukemia/lymphoma. Am J Pathol 176(1):402–415
Nosaka K, Maeda M, Tamiya S, Sakai T, Mitsuya H, Matsuoka M (2000) Increasing methylation of the CDKN2A gene is associated with the progression of adult T-cell leukemia. Cancer Res 60(4):1043–1048
Fujikawa D, Nakagawa S, Hori M et al (2016) Polycomb-dependent epigenetic landscape in adult T-cell leukemia. Blood 127(14):1790–1802
Krejsgaard T, Lindahl LM, Mongan NP et al (2016) Malignant inflammation in cutaneous T-cell lymphoma—a hostile takeover. Seminars in immunopathology
Macias ES, Pereira FA, Rietkerk W, Safai B (2011) Superantigens in dermatology. J Am Acad Dermatol 64(3):455–472; Quiz 473–454
Suga H, Sugaya M, Miyagaki T et al (2014) Skin barrier dysfunction and low antimicrobial peptide expression in cutaneous T-cell lymphoma. Clin Cancer Res 20(16):4339–4348
Thode C, Woetmann A, Wandall HH et al (2015) Malignant T cells secrete galectins and induce epidermal hyperproliferation and disorganized stratification in a skin model of cutaneous T-cell lymphoma. J Invest Dermatol. 135(1):238–246
Dobbeling U, Dummer R, Laine E, Potoczna N, Qin JZ, Burg G (1998) Interleukin-15 is an autocrine/paracrine viability factor for cutaneous T-cell lymphoma cells. Blood 92(1):252–258
Leroy S, Dubois S, Tenaud I et al (2001) Interleukin-15 expression in cutaneous T-cell lymphoma (mycosis fungoides and Sezary syndrome). The British journal of dermatology. 144(5):1016–1023
Netchiporouk E, Litvinov IV, Moreau L, Gilbert M, Sasseville D, Duvic M (2014) Deregulation in STAT signaling is important for cutaneous T-cell lymphoma (CTCL) pathogenesis and cancer progression. Cell Cycle 13(21):3331–3335
Mishra A, La Perle K, Kwiatkowski S et al (2016) Mechanism, Consequences, and Therapeutic Targeting of Abnormal IL15 Signaling in Cutaneous T-cell Lymphoma. Cancer Discov 6(9):986–1005
McKenzie RC, Jones CL, Tosi I, Caesar JA, Whittaker SJ, Mitchell TJ (2012) Constitutive activation of STAT3 in Sezary syndrome is independent of SHP-1. Leukemia 26(2):323–331
van der Fits L, Out-Luiting JJ, van Leeuwen MA et al (2012) Autocrine IL-21 stimulation is involved in the maintenance of constitutive STAT3 activation in Sezary syndrome. J Invest Dermatol. 132(2):440–447
Takahashi N, Sugaya M, Suga H et al (2016) Thymic Stromal Chemokine TSLP Acts through Th2 Cytokine Production to Induce Cutaneous T-cell Lymphoma. Cancer Res 76(21):6241–6252
Tuzova M, Richmond J, Wolpowitz D et al (2015) CCR4+ T cell recruitment to the skin in mycosis fungoides: potential contributions by thymic stromal lymphopoietin and interleukin-16. Leuk Lymphoma 56(2):440–449
Geskin LJ, Viragova S, Stolz DB, Fuschiotti P (2015) Interleukin-13 is overexpressed in cutaneous T-cell lymphoma cells and regulates their proliferation. Blood 125(18):2798–2805
Ohmatsu H, Humme D, Gulati N et al (2014) IL32 is progressively expressed in mycosis fungoides independent of helper T-cell 2 and helper T-cell 9 polarization. Cancer Immunol Res 2(9):890–900
Suga H, Sugaya M, Miyagaki T et al (2014) The role of IL-32 in cutaneous T-cell lymphoma. J Invest Dermatol. 134(5):1428–1435
Lauenborg B, Christensen L, Ralfkiaer U et al (2015) Malignant T cells express lymphotoxin alpha and drive endothelial activation in cutaneous T cell lymphoma. Oncotarget. 6(17):15235–15249
Michel L, Jean-Louis F, Begue E, Bensussan A, Bagot M (2013) Use of PLS3, Twist, CD158 k/KIR3DL2, and NKp46 gene expression combination for reliable Sezary syndrome diagnosis. Blood 121(8):1477–1478
Wong HK, Gibson H, Hake T et al (2015) Promoter-Specific Hypomethylation Is Associated with Overexpression of PLS3, GATA6, and TWIST1 in the Sezary Syndrome. J Invest Dermatol. 135(8):2084–2092
Huang Y, Su MW, Jiang X, Zhou Y (2015) Evidence of an oncogenic role of aberrant TOX activation in cutaneous T-cell lymphoma. Blood 125(9):1435–1443
Dulmage BO, Akilov O, Vu JR, Falo LD, Geskin LJ (2015) Dysregulation of the TOX-RUNX3 pathway in cutaneous T-cell lymphoma. Oncotarget
Haider A, Steininger A, Ullmann R et al (2016) Inactivation of RUNX3/p46 Promotes Cutaneous T-Cell Lymphoma. J Invest Dermatol. 136(11):2287–2296
Choi J, Goh G, Walradt T et al (2015) Genomic landscape of cutaneous T cell lymphoma. Nat Genet 47(9):1011–1019
da Silva Almeida AC, Abate F, Khiabanian H et al (2015) The mutational landscape of cutaneous T cell lymphoma and Sezary syndrome. Nat Genet 47(12):1465–1470
Kiel MJ, Sahasrabuddhe AA, Rolland DC et al (2015) Genomic analyses reveal recurrent mutations in epigenetic modifiers and the JAK-STAT pathway in Sezary syndrome. Nat Commun. 6:8470
Wang L, Ni X, Covington KR et al (2015) Genomic profiling of Sezary syndrome identifies alterations of key T cell signaling and differentiation genes. Nat Genet 47(12):1426–1434
Mathur R, Alver BH, San Roman AK et al (2017) ARID1A loss impairs enhancer-mediated gene regulation and drives colon cancer in mice. Nat Genet 49(2):296–302
Watanabe R, Ui A, Kanno S et al (2014) SWI/SNF factors required for cellular resistance to DNA damage include ARID1A and ARID1B and show interdependent protein stability. Cancer Res 74(9):2465–2475
Williamson CT, Miller R, Pemberton HN et al (2016) ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat Commun. 7:13837
Guo C, Chen LH, Huang Y et al (2013) KMT2D maintains neoplastic cell proliferation and global histone H3 lysine 4 monomethylation. Oncotarget. 4(11):2144–2153
Kaikkonen MU, Spann NJ, Heinz S et al (2013) Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol Cell 51(3):310–325
Hidaka T, Nakahata S, Hatakeyama K et al (2008) Down-regulation of TCF8 is involved in the leukemogenesis of adult T-cell leukemia/lymphoma. Blood 112(2):383–393
Papadopoulou V, Postigo A, Sanchez-Tillo E, Porter AC, Wagner SD (2010) ZEB1 and CtBP form a repressive complex at a distal promoter element of the BCL6 locus. Biochem J 427(3):541–550
Wang J, Lee S, Teh CE, Bunting K, Ma L, Shannon MF (2009) The transcription repressor, ZEB1, cooperates with CtBP2 and HDAC1 to suppress IL-2 gene activation in T cells. Int Immunol 21(3):227–235
Migliazza A, Lombardi L, Rocchi M et al (1994) Heterogeneous chromosomal aberrations generate 3’ truncations of the NFKB2/lyt-10 gene in lymphoid malignancies. Blood 84(11):3850–3860
Neri A, Fracchiolla NS, Migliazza A, Trecca D, Lombardi L (1996) The involvement of the candidate proto-oncogene NFKB2/lyt-10 in lymphoid malignancies. Leuk Lymphoma 23(1–2):43–48
Ungewickell A, Bhaduri A, Rios E et al (2015) Genomic analysis of mycosis fungoides and Sezary syndrome identifies recurrent alterations in TNFR2. Nat Genet 47(9):1056–1060
Ralfkiaer U, Hagedorn PH, Bangsgaard N et al (2011) Diagnostic microRNA profiling in cutaneous T-cell lymphoma (CTCL). Blood 118(22):5891–5900
Sandoval J, Diaz-Lagares A, Salgado R et al (2015) MicroRNA expression profiling and DNA methylation signature for deregulated microRNA in cutaneous T-cell lymphoma. J Invest Dermatol. 135(4):1128–1137
Ralfkiaer U, Lindahl LM, Litman T et al (2014) MicroRNA expression in early mycosis fungoides is distinctly different from atopic dermatitis and advanced cutaneous T-cell lymphoma. Anticancer Res 34(12):7207–7217
Narducci MG, Arcelli D, Picchio MC et al (2011) MicroRNA profiling reveals that miR-21, miR486 and miR-214 are upregulated and involved in cell survival in Sezary syndrome. Cell Death Dis 2:e151
Carreon JD, Morton LM, Devesa SS et al (2008) Incidence of lymphoid neoplasms by subtype among six Asian ethnic groups in the United States, 1996-2004. Cancer Causes Control 19(10):1171–1181
Morton LM, Wang SS, Devesa SS, Hartge P, Weisenburger DD, Linet MS (2006) Lymphoma incidence patterns by WHO subtype in the United States, 1992-2001. Blood 107(1):265–276
Chan JK, Sin VC, Wong KF et al (1997) Nonnasal lymphoma expressing the natural killer cell marker CD56: a clinicopathologic study of 49 cases of an uncommon aggressive neoplasm. Blood 89(12):4501–4513
Matano S, Nakamura S, Nakamura S et al (1999) Monomorphic agranular natural killer cell lymphoma/leukemia with no Epstein-Barr virus association. Acta Haematol 101(4):206–208
Martin AR, Chan WC, Perry DA, Greiner TC, Weisenburger DD (1995) Aggressive natural killer cell lymphoma of the small intestine. Mod Pathol 8(5):467–472
Yagita M, Huang CL, Umehara H et al (2000) A novel natural killer cell line (KHYG-1) from a patient with aggressive natural killer cell leukemia carrying a p53 point mutation. Leukemia 14(5):922–930
Chen IM, Whalen M, Bankhurst A et al (2004) A new human natural killer leukemia cell line, IMC-1. A complex chromosomal rearrangement defined by spectral karyotyping: functional and cytogenetic characterization. Leuk Res 28(3):275–284
Kulwichit W, Edwards RH, Davenport EM, Baskar JF, Godfrey V, Raab-Traub N (1998) Expression of the Epstein-Barr virus latent membrane protein 1 induces B cell lymphoma in transgenic mice. Proc Natl Acad Sci U S A. 95(20):11963–11968
Pratt ZL, Kuzembayeva M, Sengupta S, Sugden B (2009) The microRNAs of Epstein-Barr Virus are expressed at dramatically differing levels among cell lines. Virology 386(2):387–397
Klinke O, Feederle R, Delecluse HJ (2014) Genetics of Epstein-Barr virus microRNAs. Semin Cancer Biol 26:52–59
Vereide DT, Seto E, Chiu YF et al (2014) Epstein-Barr virus maintains lymphomas via its miRNAs. Oncogene 33(10):1258–1264
Huang WT, Lin CW (2014) EBV-encoded miR-BART20-5p and miR-BART8 inhibit the IFN-gamma-STAT1 pathway associated with disease progression in nasal NK-cell lymphoma. Am J Pathol 184(4):1185–1197
Motsch N, Alles J, Imig J et al (2012) MicroRNA profiling of Epstein-Barr virus-associated NK/T-cell lymphomas by deep sequencing. PLoS ONE 7(8):e42193
Iqbal J, Kucuk C, Deleeuw RJ et al (2009) Genomic analyses reveal global functional alterations that promote tumor growth and novel tumor suppressor genes in natural killer-cell malignancies. Leukemia 23(6):1139–1151
Huang Y, de Reynies A, de Leval L et al (2010) Gene expression profiling identifies emerging oncogenic pathways operating in extranodal NK/T-cell lymphoma, nasal type. Blood 115(6):1226–1237
Nakashima Y, Tagawa H, Suzuki R et al (2005) Genome-wide array-based comparative genomic hybridization of natural killer cell lymphoma/leukemia: different genomic alteration patterns of aggressive NK-cell leukemia and extranodal Nk/T-cell lymphoma, nasal type. Genes Chromosomes Cancer 44(3):247–255
Siu LL, Chan V, Chan JK, Wong KF, Liang R, Kwong YL (2000) Consistent patterns of allelic loss in natural killer cell lymphoma. Am J Pathol 157(6):1803–1809
Siu LL, Wong KF, Chan JK, Kwong YL (1999) Comparative genomic hybridization analysis of natural killer cell lymphoma/leukemia. Recognition of consistent patterns of genetic alterations. Am J Pathol 155(5):1419–1425
Ko YH, Choi KE, Han JH, Kim JM, Ree HJ (2001) Comparative genomic hybridization study of nasal-type NK/T-cell lymphoma. Cytometry 46(2):85–91
Kucuk C, Iqbal J, Hu X et al (2011) PRDM1 is a tumor suppressor gene in natural killer cell malignancies. Proc Natl Acad Sci USA 108(50):20119–20124
Karube K, Nakagawa M, Tsuzuki S et al (2011) Identification of FOXO3 and PRDM1 as tumor-suppressor gene candidates in NK-cell neoplasms by genomic and functional analyses. Blood 118(12):3195–3204
Kucuk C, Hu X, Iqbal J et al (2013) HACE1 is a tumor suppressor gene candidate in natural killer cell neoplasms. Am J Pathol 182(1):49–55
Zhang L, Anglesio MS, O’Sullivan M et al (2007) The E3 ligase HACE1 is a critical chromosome 6q21 tumor suppressor involved in multiple cancers. Nat Med 13(9):1060–1069
Quintanilla-Martinez L, Kremer M, Keller G et al (2001) p53 Mutations in nasal natural killer/T-cell lymphoma from Mexico: association with large cell morphology and advanced disease. Am J Pathol 159(6):2095–2105
Frappier L (2012) Contributions of Epstein-Barr nuclear antigen 1 (EBNA1) to cell immortalization and survival. Viruses. 4(9):1537–1547
Li M, Chen D, Shiloh A et al (2002) Deubiquitination of p53 by HAUSP is an important pathway for p53 stabilization. Nature 416(6881):648–653
Saridakis V, Sheng Y, Sarkari F et al (2005) Structure of the p53 binding domain of HAUSP/USP7 bound to Epstein-Barr nuclear antigen 1 implications for EBV-mediated immortalization. Mol Cell 18(1):25–36
Oka T, Ouchida M, Koyama M et al (2002) Gene silencing of the tyrosine phosphatase SHP1 gene by aberrant methylation in leukemias/lymphomas. Cancer Res 62(22):6390–6394
Siu LL, Chan JK, Wong KF, Kwong YL (2002) Specific patterns of gene methylation in natural killer cell lymphomas: p73 is consistently involved. Am J Pathol 160(1):59–66
Deyoung MP, Ellisen LW (2007) p63 and p73 in human cancer: defining the network. Oncogene 26(36):5169–5183
Candi E, Agostini M, Melino G, Bernassola F (2014) How the TP53 family proteins TP63 and TP73 contribute to tumorigenesis: regulators and effectors. Hum Mutat 35(6):702–714
Lanier LL (2008) Up on the tightrope: natural killer cell activation and inhibition. Nat Immunol 9(5):495–502
Han Y, Amin HM, Frantz C et al (2006) Restoration of shp1 expression by 5-AZA-2’-deoxycytidine is associated with downregulation of JAK3/STAT3 signaling in ALK-positive anaplastic large cell lymphoma. Leukemia 20(9):1602–1609
Chim CS, Fung TK, Cheung WC, Liang R, Kwong YL (2004) SOCS1 and SHP1 hypermethylation in multiple myeloma: implications for epigenetic activation of the Jak/STAT pathway. Blood 103(12):4630–4635
Chen KF, Su JC, Liu CY et al (2012) A novel obatoclax derivative, SC-2001, induces apoptosis in hepatocellular carcinoma cells through SHP-1-dependent STAT3 inactivation. Cancer Lett 321(1):27–35
Kim DJ, Tremblay ML, Digiovanni J (2010) Protein tyrosine phosphatases, TC-PTP, SHP1, and SHP2, cooperate in rapid dephosphorylation of Stat3 in keratinocytes following UVB irradiation. PLoS ONE 5(4):e10290
Gupta SC, Phromnoi K, Aggarwal BB (2013) Morin inhibits STAT3 tyrosine 705 phosphorylation in tumor cells through activation of protein tyrosine phosphatase SHP1. Biochem Pharmacol 85(7):898–912
Kucuk C, Jiang B, Hu X et al (2015) Activating mutations of STAT5B and STAT3 in lymphomas derived from gammadelta-T or NK cells. Nat Commun. 6:6025
Takakuwa T, Dong Z, Nakatsuka S et al (2002) Frequent mutations of Fas gene in nasal NK/T cell lymphoma. Oncogene 21(30):4702–4705
Shen L, Liang AC, Lu L et al (2002) Frequent deletion of Fas gene sequences encoding death and transmembrane domains in nasal natural killer/T-cell lymphoma. Am J Pathol 161(6):2123–2131
Coppo P, Gouilleux-Gruart V, Huang Y et al (2009) STAT3 transcription factor is constitutively activated and is oncogenic in nasal-type NK/T-cell lymphoma. Leukemia 23(9):1667–1678
Yu H, Kortylewski M, Pardoll D (2007) Crosstalk between cancer and immune cells: role of STAT3 in the tumour microenvironment. Nat Rev Immunol 7(1):41–51
Yu H, Pardoll D, Jove R (2009) STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer 9(11):798–809
Wang T, Niu G, Kortylewski M et al (2004) Regulation of the innate and adaptive immune responses by Stat-3 signaling in tumor cells. Nat Med 10(1):48–54
Koo GC, Tan SY, Tang T et al (2012) Janus kinase 3-activating mutations identified in natural killer/T-cell lymphoma. Cancer Discov 2(7):591–597
Bouchekioua A, Scourzic L, de Wever O et al (2014) JAK3 deregulation by activating mutations confers invasive growth advantage in extranodal nasal-type natural killer cell lymphoma. Leukemia 28(2):338–348
Kimura H, Karube K, Ito Y et al (2014) Rare occurrence of JAK3 mutations in natural killer cell neoplasms in Japan. Leuk Lymphoma 55(4):962–963
Jiang L, Gu ZH, Yan ZX, et al. Exome sequencing identifies somatic mutations of DDX3X in natural killer/T-cell lymphoma. Nature genetics. 2015:Epub ahead of print
Dobashi A, Tsuyama N, Asaka R et al (2016) Frequent BCOR aberrations in extranodal NK/T-Cell lymphoma, nasal type. Genes Chromosomes Cancer 55(5):460–471
Lee S, Park HY, Kang SY et al (2015) Genetic alterations of JAK/STAT cascade and histone modification in extranodal NK/T-cell lymphoma nasal type. Oncotarget. 6(19):17764–17776
Miyazaki K, Yamaguchi M, Imai H et al (2009) Gene expression profiling of peripheral T-cell lymphoma including gammadelta T-cell lymphoma. Blood 113(5):1071–1074
Travert M, Huang Y, de Leval L et al (2012) Molecular features of hepatosplenic T-cell lymphoma unravels potential novel therapeutic targets. Blood 119(24):5795–5806
Moffitt AB, Ondrejka SL, McKinney M et al (2017) Enteropathy-associated T cell lymphoma subtypes are characterized by loss of function of SETD2. J Exp Med
Davis RE, Brown KD, Siebenlist U, Staudt LM (2001) Constitutive nuclear factor kappaB activity is required for survival of activated B cell-like diffuse large B cell lymphoma cells. J Exp Med 194(12):1861–1874
Bild AH, Yao G, Chang JT et al (2006) Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439(7074):353–357
Rosenwald A, Wright G, Leroy K et al (2003) Molecular diagnosis of primary mediastinal B cell lymphoma identifies a clinically favorable subgroup of diffuse large B cell lymphoma related to Hodgkin lymphoma. J Exp Med 198(6):851–862
Savage KJ, Monti S, Kutok JL et al (2003) The molecular signature of mediastinal large B-cell lymphoma differs from that of other diffuse large B-cell lymphomas and shares features with classical Hodgkin lymphoma. Blood 102(12):3871–3879
Zinzani PL, Tani M, Musuraca G et al (2006) Phase II study of proteasome inhibitor bortezomib (Velcade®) in patients with relapsed/refractory T-cell lymphoma: preliminary results. Blood 108a
Piva R, Ruggeri B, Williams M et al (2007) CEP-18770: a novel orally-active proteasome inhibitor with a tumor-selective pharmacological profile competitive with bortezomib. Blood
Feldman A, Sun D, Law M (2007) Syk Tyrosine Kinase is Overexpressed in the Majority of Peripheral T and NK-cell Lymphomas, and Re. Blood 110:690a
Wilcox RA, Sun DX, Novak A, Dogan A, Ansell SM, Feldman AL (2010) Inhibition of Syk protein tyrosine kinase induces apoptosis and blocks proliferation in T-cell non-Hodgkin’s lymphoma cell lines. Leukemia 24(1):229–232
Piccaluga PP, Rossi M, Agostinelli C et al (2014) Platelet-derived growth factor alpha mediates the proliferation of peripheral T-cell lymphoma cells via an autocrine regulatory pathway. Leukemia 28(8):1687–1697
Andrae J, Gallini R, Betsholtz C (2008) Role of platelet-derived growth factors in physiology and medicine. Genes Dev 22(10):1276–1312
Perry AM, Molina-Kirsch H, Nathwani BN et al (2011) Classification of non-Hodgkin lymphomas in Guatemala according to the World Health Organization system. Leuk Lymphoma 52(9):1681–1688
Wang C, Collins M, Kuchroo VK (2015) Effector T cell differentiation: are master regulators of effector T cells still the masters? Curr Opin Immunol 37:6–10
Rudiger T, Weisenburger DD, Anderson JR et al (2002) Peripheral T-cell lymphoma (excluding anaplastic large-cell lymphoma): results from the Non-Hodgkin’s Lymphoma Classification Project. Ann Oncol 13(1):140–149
Vose J, Armitage J, Weisenburger D, International TCLP (2008) International peripheral T-cell and natural killer/T-cell lymphoma study: pathology findings and clinical outcomes. J Clin Oncol 26(25):4124–4130
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Iqbal, J., Amador, C., McKeithan, T.W., Chan, W.C. (2019). Molecular and Genomic Landscape of Peripheral T-Cell Lymphoma. In: Querfeld, C., Zain, J., Rosen, S. (eds) T-Cell and NK-Cell Lymphomas. Cancer Treatment and Research, vol 176. Springer, Cham. https://doi.org/10.1007/978-3-319-99716-2_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-99716-2_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-99715-5
Online ISBN: 978-3-319-99716-2
eBook Packages: MedicineMedicine (R0)