Abstract
The steroid hormone estrogen has potent effects in a variety of tissues across the body in both females and males. In the nuclear signaling pathway for estrogens, the hormone acts by stimulating the DNA binding and transcriptional activity of estrogen receptors (ERs), transcription factors which robustly and transiently regulate the expression of target genes. More broadly, estrogen signaling controls the ER cistrome, as well as the epigenome and the estrogen-regulated transcriptome. A host of deep sequencing-based genomic assays have provided novel insights into the mechanisms by which ERs regulate transcriptional responses. Estrogen-dependent transcriptional responses have been studied widely in breast cancer cells, primarily in the context of the ER alpha (ERα) isoform. These studies have revealed an intricate cross talk between the estrogen-ERα signaling pathway and other signaling pathways, impacting transcriptional programs and clinical outcomes in breast cancer. This chapter reviews the key features of ERα-regulated transcription and the current technological advances that have allowed for the careful dissection of these mechanisms.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Estrogen Signaling Through Estrogen Receptors
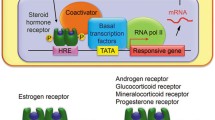
Many of the biological actions of estrogens are mediated by ERα, which functions primarily as a ligand-regulated, DNA-binding transcription factor (TF) in the nuclei of estrogen-responsive cells [1,2,3]. Natural estrogens, synthetic agonists, and synthetic antagonists promote binding of ERα to the genome. Each of these types of ligands generates distinct and overlapping ERα “cistromes” (i.e., the collection of ERα binding sites across the genome), as well as an “epigenome” (i.e., the collection of chemical modifications on chromatin and genomic DNA) and the estrogen-regulated “transcriptome” (i.e., the collection of all RNA transcripts up- or downregulated by estrogen signaling) (Table 1). Important roles for “non-genomic” or “membrane-initiated” estrogen signaling through ERs have been characterized [4], some of which may ultimately impact the transcriptome [5, 6] (Fig. 1). In this chapter, we will focus on the actions of estrogen signaling through nuclear ERα, and we describe the key concepts and molecular details of estrogen-regulated transcription.
Cytoplasmic and nuclear signaling pathways converge on nuclear ERα signaling pathways. (a) Cytoplasmic and nuclear and signaling pathways originating from the extracellular space or cytoplasmic membrane, including (from left to right) (1) direct action of estrogens on ERα in the nucleus; (2) membrane-initiated estrogen signaling through membrane-associated canonical ERα directed to the membrane by palmitoylation [7,8,9]; (3) membrane-initiated estrogen signaling through GPR30 [a.k.a. G protein-coupled estrogen receptor 1 (GPER-1)], a Gs-coupled heptahelical transmembrane receptor that binds estrogens [10]; (4) growth factor signaling (e.g., EGF and IGF-1) through growth factor receptors; and (5) proinflammatory signaling (e.g., TNFα) through cytokine receptors. The intracellular signaling pathways stimulated by the initiating events described above require kinases (e.g., Src, PI3K, AKT, PKA, JNK1, ERK1/2, IKK). (b) The intracellular signaling pathways stimulated by the initiating events described in (a) require kinases ultimately link to a variety of transcription factors [TFs; e.g., CREB, NF-κB (p65 and p50), AP-1 (Fos and Jun), and SP-1]. Many of these TFs (1) are phosphorylated by the kinases noted in (a) (indicated by the red stars) and (2) interact physically and functionally with ERα in the nucleus to promote the formation of estrogen-responsive ERα enhancers across the genome
1.1 ERα Binding Sites, Cistromes, and Transcriptional Enhancers
The binding of ligands to ERα (agonists and antagonists) stimulates receptor dimerization and binding to thousands of sites across the genome, collectively called the ERα “cistrome,” within minutes of hormone exposure [11,12,13,14,15]. While many ERα binding sites (ERBSs) contain a DNA sequence motif called the estrogen response element (ERE), the consensus of which is a 13-base pair palindrome with a 3-base pair spacer [2], others lack any semblance of such a motif and may recruit ERα indirectly through other TFs (see below) [16, 17] (Fig. 1b). Direct-binding ERBSs are pre-established and prebound (“marked”) by pioneer transcription factors, such as FoxA1 or AP2γ prior to ERα binding [18, 19]. DNA-bound ERα acts as a nucleation site and scaffold for the assembly of multiprotein complexes containing histone-modifying enzymes, ATP-dependent nucleosome-remodeling enzymes, and Mediator, an RNA polymerase II (Pol II)-interacting coregulator [20,21,22,23] (Fig. 2 and Table 2). The coordinated recruitment of these coregulator proteins and chromatin-modifying enzymes functions to establish an active transcriptional “enhancer,” leading to chromatin looping and target gene transcription [24,25,26,27,28] (Fig. 2). The transcriptional effects of estrogen signaling are rapid, on the order of minutes, resulting in transcription at both ERα enhancers and their target genes [25, 29, 30].
Features of ERα enhancers. Agonist-bound ERα dimerizes and binds to regulatory regions across the genome to promote the formation of estrogen-regulated enhancers. ERα binds to these regions with assistance from other TFs, such as pioneer factors (e.g., FoxA1) or cooperatively interacting TFs (see Fig. 1). ERα binds to genomic directly DNA through its sequence element [i.e., the estrogen response element (ERE)] or via tethering mechanisms with other DNA-binding transcription factors. ERα nucleates the formation of enhancers by recruiting direct-binding transcriptional coregulators, including members of the SRC family (SRC-1, SRC-2, or SRC-3), the Swi/Snf complex (via its Brg1 subunit), and the Mediator complex (via its Med1 subunit). These, in turn, recruit other coregulators, including histone-modifying enzymes, such as the lysine acetyltransferase p300/CBP. Ultimately, the forming enhancer recruits the RNA Pol II transcriptional machinery, which loops to target gene promoters to promote gene transcription. Enhancers are characterized “active” histone modifications (e.g., H3K27ac and H3K4me1), coregulator recruitment, an open chromatin architecture, and the production of enhancer RNAs (eRNAs). Although not depicted here for simplicity, many of the features and factors shown at the enhancer are also found at the promoter
1.2 Direct and Indirect ERα Binding Sites
The nuclear actions of ERα occur through both “classical” (direct) and “nonclassical” (indirect) binding of ERα to the genome [31]. In the classical pathway, ERα binds to EREs (short DNA motifs with the consensus AGGTCAnnnTGACCT) [32, 33]. In contrast, the nonclassical genomic pathway does not require the presence of ERE in ERα binding sites. Rather, the binding of ERα to the genome is facilitated by tethering mechanisms with other TFs, such as AP1, NF-κB, CREB, and SP1 [34,35,36,37] (Fig. 1b). Variable binding of ERα and other TFs to their genomic binding sites may also be influenced by neighboring sequence or motifs, as well as combinations of different TFs [16, 17, 38, 39]. Notably, many ERα binding sites contain “imperfect” (i.e., non-consensus) EREs, which has led to a reevaluation of the sequence determinants for ERα binding and perhaps a redefinition of the concept of classical and nonclassical binding.
1.3 Features of Active ERα Enhancers
Active ERα enhancers (i.e., those capable of promoting target gene expression) are associated with a variety of features that provide useful “marks” of enhancer activity [40]. These include (1) enrichment of specific histone modifications [e.g., H3 lysine 4 monomethyl (H3K4me1), H3 lysine 27 acetyl (H3K27ac)], coregulators (e.g., p300/CBP, Mediator), and RNA polymerase II (Pol II) [41,42,43], (2) an open chromatin structure (e.g., DNAse I hypersensitivity, nucleosome depletion by FAIRE-seq) [44, 45], (3) enhancer transcription (enhancer RNA production) [25, 45, 46], and (4) looping to target gene promoters [47] (Figs. 2 and 3). Different classes of enhancers may exhibit differential accumulation of these features in a context-dependent manner to specify distinct gene regulatory mechanisms [48]. Differential enhancer selection and activity result in cell- and context-specific gene expression. While the definition of enhancers continues to evolve, technological advances have revealed new features of enhancer function and generated renewed interest in enhancers as important regulators of tissue-specific gene expression. Some of the remaining questions address challenges in the identification, conservation, and functional annotation of these enhancers [49].
Visualizing ERα enhancer formation and E2-regulated gene transcription with genomic assays. The use of genomic assays to visualize estrogen-dependent regulation of the P2RY2 gene Fig. 3 (continued) and distal enhancer in ERα-positive MCF-7 breast cancer cells (from top to bottom): (1) GRO-seq measures of actively transcribing RNA polymerases across the genome, shown for a 40 min time course of treatment with 17β-estradiol (E2); (2) RNA-seq measures the steady-state levels of RNAs, including mRNAs; and (3) ChIP-seq for ERα (±E2, 45 min), FoxA1 (+E2, 45 min), H3K4me1 (+E2, 45 min), and H3K27ac (+E2, 45 min) shows the enrichment of these factors and chromatin modifications at various loci. The P2RY2 gene annotation (with exons and introns) and a size marker (in kb) are shown. Adapted from [25]
2 Coregulators for Estrogen Receptors
The binding of ERα to genomic regulatory regions promotes the recruitment of a broad array of coregulator proteins and coregulator complexes (Table 2). Some of these coregulators interact directly with ERα (e.g., the steroid receptor coactivators, SRC1, 2, and 3; the Mediator complex via Med1; and the Swi/Snf complex via Brg1), while others interact indirectly through scaffolding coregulators (e.g., p300 and CBP, which interact with ERα through SRCs proteins) [50,51,52,53]. These coregulators serve a variety of functions, including (1) the posttranslational modification of histones and other transcription-related proteins (p300 and CBP, which are protein acetyltransferases) [54], (2) chromatin remodeling (e.g., Swi/Snf complex) [55], and (3) chromatin looping (e.g., Mediator complex) [56,57,58,59]. These events ultimately facilitate looping events that associate enhancers to target promoters and induce changes in RNA polymerase II (Pol II) occupancy or activity [60]. While SRCs, p300/CBP (sometimes referred to collectively), Swi/Snf, and Mediator function with ERα, they also support the transcriptional activity of many other TFs. Thus, the ligand activation and specific ERBSs are critical determinants of their function in the estrogen signaling pathway.
2.1 Coregulators and Noncoding RNAs
Details about the specific functions of coregulators, especially as they relate to enhancer formation and activity, are becoming clearer. Some of these functions require interactions with noncoding RNAs. For example, SRC-1 interacts with SRA (steroid receptor RNA activator) to facilitate ERα-dependent transcriptional outcomes [61]. SRA was subsequently found to enhance the activities of the AF-1 and AF-2 domains of ERα [62, 63]. p300/CBP is thought to be the key enzyme that acetylates histone H3 at lysine 27 (H3K27), a mark of active enhancers [42, 64, 65]. Interestingly, interactions with short noncoding RNAs produced from enhancers (i.e., eRNAs) may stimulate p300 catalytic activity at enhancers [66]. Interaction of Mediator with enhancer-derived noncoding RNAs [e.g., long noncoding RNAs (lncRNAs) or eRNAs] via the Med12 subunit promotes enhancer-promoter looping [59]. These are a few of the details about coregulatory functions that have emerged recently, but a number of gaps in our knowledge remain. For example, we still know little about the temporal aspects of coregulator function, although recent studies have shown that coregulators may have distinct and changing roles during the time course of enhancer function [67].
3 Molecular Aspects of Estrogen-Dependent Transcriptional Outcomes
ERα regulates gene expression by increasing or decreasing transcription of its target genes, which include protein-coding mRNA genes, as well as noncoding RNA genes for microRNAs, lncRNAs, transfer RNAs (tRNAs), and ribosomal RNAs (rRNAs) [29]. In turn, the products of these transcriptional targets are involved in various cellular functions, including the regulation of a secondary “wave” of target genes and their RNA products. Ultimately, estrogen-regulated transcriptional responses lead to changes in the transcriptome by altering RNA Pol I, II, and III transcription, as well as the proteome by altering the levels of mRNAs (via microRNAs), ribosome biogenesis (via rRNAs and ribosomal proteins), and translation (via tRNAs and translation regulatory proteins) [29] (Fig. 4).
Broad effects of estrogen signaling on the transcriptome. Estrogen signaling via ERα regulates transcription from all three RNA polymerases (Pols I, II, and III) in ERα-positive MCF-7 breast cancer cells. The products from these estrogen-regulated transcription events include eRNAs, mRNAs, lncRNAs, microRNAs, rRNAs, and tRNAs. Estrogen signaling regulates (1) mRNA expression by controlling Pol II activity at target gene promoters via eRNAs and lncRNAs, as well as mRNA stability via microRNAs; (2) ribosome biogenesis by enhancing the production of rRNAs, as well as mRNAs encoding ribosomal proteins; and (3) protein translation by enhancing the production of tRNAs, as well as promoting ribosome biogenesis as noted above. The functional interactions between the RNA products from estrogen-regulated transcription have profound effects on the cell and collectively promote mitogenic responses [29, 30]
3.1 Primary and Secondary Transcriptional Responses
Defining the hierarchy of the estrogen transcriptional responses has helped delineate the mechanism of estrogen action. To this end, estrogen transcriptional responsive genes can be categorized as primary response and secondary response genes. Primary response genes are those that are regulated as an immediate response to cellular signaling without the requirement for protein synthesis [68, 69]. Recently, the concept of “direct” target genes has emerged to describe target genes whose promoters are associated with TF binding at proximal or distal enhancers [11, 15, 26]. Secondary response genes are regulated following protein synthesis. The transcriptional regulation of secondary response genes may involve transcriptional cross talk between ERα with the products of the primary response (Fig. 4).
3.2 Pol II Recruitment and Promoter-Proximal Pausing
The concepts of “primary/immediate” and “direct” target genes often describe two distinct aspects of the mechanism of transcriptional regulation and are often incorrectly used synonymously. The distinction arises from the underlying mechanisms of Pol II regulation at ERα target promoters [70]. The prevailing view of the regulation commences with ligand-dependent ERα enhancer formation, which subsequently mediates the recruitment of the Pol II machinery to promoters. The rate-limiting step in this mechanism is the recruitment of Pol II [70, 71] (Fig. 3). However, more recent studies in the fly and mammalian systems have revealed that Pol II is preloaded at many promoters across the genome prior to stimulation, initiating the synthesis of a short transcript and subsequently “pausing” ~20–40 base pairs downstream of the TSS [72] (Fig. 5). This mechanism may allow rapid transcriptional responses to stimuli by bypassing the rate-limiting step of the aforementioned mechanism or possibly generate synchronous transcriptional responses to stimuli for a population of genes or cells. In the context of estrogen-regulated transcription, regulation of Pol II loading and release of Pol II from pause sites can be regulated by nuclear ERα action, as well as membrane-initiated estrogen signaling.
Robust and transient ERα-regulated transcription. Metaplots showing the average read profiles of GRO-seq data for estrogen-stimulated genes in ERα-positive MCF-7 breast cancer cells over a 160 min time course of E2 treatment. The data are aligned relative to the transcriptional start sites (TSSs) of the genes. For the genes shown here with maximal transcription at 40 min of E2 treatment, the GRO-seq data reveal the following: (a) Strong polymerase pause peak immediately downstream of the TSS prior to E2 treatment. (b) Additional E2-dependent Pol II recruitment (loading), as well as enhanced pausing, after 10 min of E2 treatment. (c) Robust transition from pausing to elongation across the gene body. (d) Attenuation of transcriptional activity are resetting to basal levels. Adapted from [29]
In the subsequent sections, we have summarized the approaches employed to interrogate these aspects of transcriptional regulation and highlight the most significant findings from recent studies of estrogen-dependent transcription through ERα.
4 Overview of Genomic Analyses of Estrogen-Regulated Transcription
The advent of microarray-based genomic technologies in the late 1980s and early 1990s, leading to the groundbreaking paper by Pat Brown and colleagues in 1995 reporting the results of the first microarray expression analysis [73], ushered in the age of genomics. The fields of molecular biology and biomedical sciences have never looked back. The past decade has seen a rapid expansion of “genomic” methods for the analysis of signal-regulated transcription, including estrogen-regulated transcription through ERα. Today, a plethora of genomic technologies, now based on next-generation or deep sequencing technologies, are available to scientists to explore the features of the genome, transcriptome, and cistrome of their favorite biological system in a facile and robust manner. As scientists have learned how to apply these technologies, we have seen how they can be used effectively for discovery-based experiments, as well as to test specific hypotheses on a global scale [74].
4.1 Aspects of Estrogen-Regulated Transcription Queried on a Genomic Scale
Today, nearly every aspect of signal-regulated transcription by a diverse array of TFs, including ERα and other nuclear receptors, can be probed with a genomic approach . These include TF binding, chromatin opening, enhancer assembly, enhancer-promoter looping, and transcriptional outcomes (Table 3). More broadly, downstream actions, such as protein translation, can also be probed as well through Ribo-seq and ribosome profiling [75] (Table 3). Through these approaches, a clearer picture of the molecular details of signal-regulated transcription has emerged.
4.2 Current Challenges and Opportunities in Genomic Analyses of Transcription
While our understanding of the estrogen-regulated transcriptome has advanced quite rapidly in the age of genomics, there are a number of challenges that must be overcome to increase our understanding further. These challenges include (1) single cell analyses of not only the transcriptome, but also of cistromes, chromatin accessibility, and looping, (2) allele-specific transcriptional effects, and (3) analyses in tissues and pathological samples. Overcoming some of these challenges will require the development of new methodologies.
The age of genomics and high-throughput screening (e.g., siRNA, shRNA, or CRISPR screens; [76, 77]) has also brought many new opportunities. The possibility to conduct perturbation-response experiments and assays of kinetics on a genome-wide scale should bring even greater understanding of the molecular mechanisms of estrogen-regulated transcription. While there has been a predominant focus on upregulation, genomic assays provide an opportunity to explore downregulation or repression on a global scale.
5 Global Views of the Estrogen-Regulated Transcriptome
Numerous studies over the past two decades have examined the mechanisms and outcomes of estrogen-regulated transcription using various genomic approaches, such as gene expression microarrays, as well as genome-wide chromatin immunoprecipitation (ChIP) assays (ChIP-chip and ChIP-seq) [78]. Although they have provided a wealth of knowledge about the biology of estrogen signaling in the nucleus, these studies have not always provided a consistent view of the primary or immediate estrogen-regulated gene set. This information is critical for mechanistic studies when trying to relate estrogen signaling to specific molecular events at target gene promoters. Like expression microarrays, which have produced discrepancies in the numbers of estrogen-regulated genes within a given cell type [78, 79], genomic ChIP assays of ERα and Pol II have not produced a clear picture of the estrogen-regulated gene set. This is due, in part, to the difficulty in assigning ERα binding events to specific gene regulatory outcomes [15, 80]. New technologies for assessing enhancer-promoter looping events (e.g., ChIA-PET) and for measuring nascent transcription (e.g., GRO-seq) have helped to overcome some of these problems [72, 81]. A limitation of many of the earlier analyses is that they focused on the effects of estrogen signaling on the expression of annotated Pol II transcripts (i.e., transcripts encoding proteins and microRNAs), without considering potential effects on unannotated transcripts, or Pol I and Pol III transcripts. In this section, we describe studies that have employed genomic approaches to query the estrogen-regulated transcriptome.
5.1 Expression Microarrays and RNA-seq
The earliest attempts to understand the global effects of estrogen signaling on the transcriptome were performed using microarrays (reviewed in [82]). Microarray expression profiling relies on the hybridization of differentially labeled fluorescent cDNA probes to control/reference samples and experimental samples to determine relative steady-state expression levels of thousands of annotated mRNAs across different conditions and treatments [73]. Breast cancer was one of the first and has been one of the most widely studied, biological models analyzed with this technique [83]. In fact, the expression profiles of breast cancers were used to identify and define a set of distinct molecular subtypes, a classification system that is still used today [84]. Using information from expression microarrays, molecular taxonomy was used to group breast cancers into four major subtypes, luminal A, luminal B, basal-like, and HER2+, related to expressed molecular markers [85]. These breast cancer molecular subtypes exhibit distinct clinical and pathological phenotypes [86].
The most commonly employed model for studying the estrogen-regulated transcriptome is the ERα-positive, luminal A adenocarcinoma cell line, MCF-7. Microarray analyses using these cells helped to provide an initial view of some of the transcriptional responses to estrogen [79]. Expression microarray experiments in MCF-7 cells have been performed with a wide range of estrogen treatment times (e.g., 0, 3, 6, 12, and 24 h). They have yielded estimates for the number of estrogen-responsive target genes ranging from ~100 to ~1500 [12, 78, 79, 87]. In these experiments, the genes regulated at the earliest time points were considered to be immediate, direct, or primary transcriptional targets, while the genes regulated at the later time points were considered to be late, indirect, or secondary targets. Several attempts have been made to define the primary estrogen-regulated transcriptome by employing cycloheximide, a potent mRNA translation inhibitor, to prevent the secondary effects of estrogen-regulated transcription [87]. However, this approach is limited by the toxic effects of cycloheximide and its inability to account for the effects of noncoding transcriptional products, such as microRNAs and lncRNAs. As discussed below, it is likely that even in the earliest of these time points (i.e., 3 h), the regulated genes reported include indirect or secondary effects of estrogen treatment.
Many of the salient features of the results from the experiments with MCF-7 cells are conserved across estrogen-dependent biological systems. However, the results from different studies using the same cell type exhibit variability due to a variety of factors, including the cell growth and estrogen treatment conditions, the subpopulations of MCF-7 clones, and the specific microarray platforms. Moreover, technical limitations associated with the microarray approach, such as relatively high level of noise, low sensitivity for rare or low-abundance transcripts, narrow dynamic range, and biased detection of transcript variants, gene fusions, single nucleotide variants, and indels (small insertions and deletions), have limited the utility of microarrays for gene expression analyses. However, the low cost and facile application of microarray technology have made expression microarrays a useful diagnostic tool for the clinic [88].
More recent developments in next generation sequencing technologies have given way to more powerful techniques for gene expression profiling, including RNA-seq, which overcome many of the limitations encountered by microarrays. RNA expression analyses using RNA-seq allow for sensitive and unbiased profiling of the steady-state estrogen-regulated transcriptome. From the gene expression perspective, many of the conclusions about the estrogen-regulated transcriptome from RNA-seq studies mirror those made using expression microarrays. But, RNA-seq studies have facilitated a greater exploration of the effects of estrogen on noncoding RNA expression, splicing, and ribosome loading, revealing new facets of the molecular response to estrogen signaling.
5.2 GRO-seq and Derivatives
While gene expression profiling with microarrays and RNA-seq have provided an abundance of information regarding estrogen responses, these approaches have fallen short in defining the primary and immediate responses at the genomic level. Microarray and RNA-seq can reveal the steady-state levels of mRNA (or other RNAs) and, therefore, require the accumulation of transcripts over time. Our understanding of estrogen action at the genomic level has expanded with more recent developments in next generation sequencing technologies that measure active or ongoing transcription with higher sensitivity and temporal resolution. One such method, global run-on coupled with massively parallel sequencing (GRO-seq), has been used in breast cancer cells and other biological systems to identify new features of estrogen-regulated transcription [25, 29, 89].
GRO-seq is a direct sequencing method that provides a “map” of the position and orientation of all engaged RNA polymerases (Pols I, II, and III) across the genome at extremely high resolution. It has helped to (1) identify primary response genes, (2) establish a hierarchy of the estrogen-regulated transcriptional network, (3) reveal mechanisms of the estrogen-regulated transcriptional response, and (4) identify novel unannotated and noncoding transcripts. Derivatives of GRO-seq, such as precision run-on sequencing (PRO-seq) and PRO-cap (or 5′-GRO-seq), allow (1) mapping of the 3′ end of nascent RNAs in the Pol II active site with nucleotide resolution and (2) mapping of the 5′ end of nascent RNAs with nucleotide resolution [90], respectively (Table 3). These methods have revealed new facets of estrogen-regulated transcription, which have not been discerned using other techniques.
Using GRO-seq and short time courses of estrogen treatment in MCF-7 cells, the Kraus lab has made a number of observations about the estrogen-regulated transcriptional response, reported in a series of papers over the past 8 years. Three adjectives that describe this response are rapid, extensive, and transient [29]. Here we summarize some of these findings.
Estrogen-Dependent Transcriptional Responses Are Rapid
The initial GRO-seq experiments in MCF-7 cells recalibrated our understanding about the time scale of estrogen-dependent transcription experiments. While the steady-state RNA experiments with expression microarrays and RNA-seq examined responses on the order of hours, and sometime even days, the direct transcriptional readout from GRO-seq experiments showed that the responses occurred on the order of minutes [29] (Fig. 3). In fact, the transcriptional outcomes of nuclear estrogen signaling occur at least as fast, if not faster, than the so-called “rapid” membrane-initiated actions. Using GRO-seq and with detailed time courses of estrogen treatment, we were able to determine the rates of Pol II transcription for estrogen-regulated genes [91]. Elongation rates varied as much as fourfold at different genomic loci. Gene body elongation rates correlated with the density of Pol II on the gene, resulting in higher rates of transcript production at genes with higher Pol II densities. These studies also revealed that estrogen stimulates gene expression by increasing Pol II initiation, whereas other signaling pathways (e.g., TNFα) reduce Pol II residence time at promoter proximal pause sites [91]. Collectively, these studies provided new insights into the mechanisms of estrogen-regulated transcription.
Estrogen-Dependent Transcriptional Responses Are Extensive
The early expression microarray and RNA-seq experiments focused on the Pol II mRNA transcriptome or subsets thereof. Given that GRO-seq can readily detect transcription by all three RNA polymerases, it provides an opportunity to explore the transcriptome more broadly. The initial GRO-seq experiments in MCF-7 cells showed that estrogen signaling not only regulates the Pol II transcriptome but also the Pol I and Pol III transcriptomes as well [29]. The expression of every major class of RNA, including rRNAs and tRNAs, is altered by estrogen signaling (Fig. 4). By developing computational tools for analyzing GRO-seq data, such as groHMM [92], we were able to annotate novel transcripts in MCF-7 cells, including previously unannotated lncRNAs and eRNAs (discussed in more detail below) [29, 93].
Estrogen-Dependent Transcriptional Responses Are Transient
An additional observation about the estrogen-regulated transcriptome from the initial GRO-seq experiments was that upregulation was generally transient, with the majority of upregulated genes showing maximal transcription at 40 min and then declining thereafter [29]. In contrast, the downregulated genes stayed down for longer periods of time. Together, these results illustrate that estrogen-dependent transcriptional responses are transient and that upregulation and downregulation may occur by fundamentally different mechanisms [29].
6 LncRNAs and eRNAs Revealed by GRO-seq
As noted above, GRO-seq has been a useful tool for annotating new transcription units, including those that produce lncRNAs and eRNAs.
6.1 LncRNAs
We found that a computational approach integrating both GRO-seq and RNA-seq increased the sensitivity for detecting low-abundance lncRNAs [93]. Integration of these data with genomic information about histone modifications and factor binding at lncRNA gene promoters provided new insights about lncRNA gene structure and regulation, as well as lncRNA transcript stability, regulation, and function. For example, we observed that many ERα binding sites occur in lncRNAs gene promoters, which are also marked with histone modifications that are typical of transcriptional enhancers [93]. Functional analysis of selected lncRNAs with altered expression in breast cancers, such as lncRNA67 and lncRNA152, revealed novel roles for these lncRNAs in cell proliferation, regulation of an E2F-dependent cell-cycle gene expression program, and estrogen-dependent mitogenic growth. These studies illustrated the power of GRO-seq data, when combined with appropriate computational tools, to annotate novel transcription units across the genome.
6.2 Enhancer Transcription and eRNAs
Recent studies, including work from the Kraus lab , have shown that many enhancers overlap with sites of Pol II loading and the production of enhancer RNAs (“eRNAs”) [29, 94,95,96,97,98,99]. GRO-seq has been a powerful approach for detecting, analyzing, and annotating enhancer transcription and eRNAs [25, 67]. Enhancer transcription mirrors the kinetics of the emergence of other enhancer features, as well as enhancer activation and target gene transcription [29, 100, 101] (Fig. 3). A common signature of enhancer transcription is the production of short (i.e., ~1–2 kb) eRNAs that are transcribed bidirectionally [95] and can be readily detected by GRO-seq [25, 29, 96,97,98, 101]. The role of transcription in enhancer function is unknown, but the act of transcription may help to create an open chromatin environment that promotes enhancer function [46, 102]. Alternatively, the stable accumulation of eRNAs may play a functional or structural role and may facilitate gene looping (reviewed in [103]).
7 Integrating the Estrogen-Regulated Transcriptome with Other Aspects of Gene Regulation
In the sections above, we described fundamental aspects of the estrogen-regulated transcriptome and genomic methods to analyze it. The RNA Pol II transcriptome is of particular interest to the study of hormone signaling because it comprises a set of mRNAs that encode the proteome, as well as functional RNAs that can regulate the expression or abundance of mRNAs (e.g., lncRNAs and microRNAs). As noted above, the promoters of the estrogen-regulated genes that produce mRNAs, lncRNAs, and microRNAs are controlled by transcriptional enhancers that nucleate at ERα binding sites and communicate with the promoters through higher-order chromatin looping mechanisms. Thus, a full understanding of the regulation of the RNA Pol II transcriptome requires the integration of transcriptome data with other genomic data (e.g., including cistrome, epigenome, chromatin accessibility, and chromatin looping data), which allows the identification of active enhancers and their target genes.
7.1 Identifying Active Estrogen-Regulated Enhancers from Genomic Data
Deep sequencing technologies used to study transcriptomes and epigenomes have revealed that the genome is pervasively transcribed [29, 104, 105] and that the epigenome is remarkably plastic [41, 106]. These studies have also identified transcriptional enhancers as the key regulatory elements that control the cell type-specific biology of essentially all biological systems examined to date [48, 107]. However, the cistrome for a given TF is not synonymous with the set of active enhancers nucleated by the TF, since many TF binding sites are not functional as enhancers. For ERα, the number of ERE-like sequences is greater than the number of ERBSs, and the number of ERBSs is greater than the number of functionally active ERα enhancers [25]. Thus, identifying active enhancers, rather than TF binding sites, is important for understanding the estrogen-dependent regulation of gene expression.
As discussed above, our current understanding of the features of active enhancers has been derived from an integration of a variety of genomic techniques and complemented with specific mechanistic assays. Active enhancers , including those nucleated by agonist-occupied ERα, share several common features [48, 107, 108]. For example, enhancers are typically (1) located in open regions of chromatin (as assessed by DNase-seq) [109, 110]; (2) enriched with a common set of histone modifications (as assessed by ChIP-seq), including H3K4me1 and H3K27ac [41, 106]; (3) enriched with the coregulators p300 and CBP (as assessed by ChIP-seq) [41,42,43]; (4) bound by RNA Pol II (as assessed by ChIP-seq); and (5) actively transcribed, producing enhancer RNAs (“eRNAs”) (as assessed by RNA-seq, GRO-seq, and derivatives) [29, 94, 95] and loop to target gene promoters (as assessed by 3C-based methods) [47]. Ongoing transcription at enhancers, as assessed by GRO-seq and derivatives, can be used for the prediction of active enhancers [25, 29, 48, 92, 96, 111]. Integration of data from multiple genomic methods using computational pipelines can provide an effective way to identify active enhancers [25, 67, 112]. Of course, these genomic studies should always be followed up with additional locus-specific perturbation studies to mechanistically define the biological functions of selected enhancers that were identified with the genomic approaches.
7.2 Identifying the Target Genes of ERα Enhancers from Genomic Data
The next challenge after identifying active enhancers is to identify the target genes that they regulate. This has been typically done in one of two ways. The first is with a “nearest neighboring gene” approach , which is based on the assumption that genes closest to an active enhancer have the greatest likelihood of being regulated by that enhancer. Although this assumption is not always true and excludes some nuances related to enhancer-promoter communication and regulation, it can be a useful approach that reveals verifiable aspects of target gene regulation. In fact, the nearest neighboring genes of breast cancer subtype-specific enhancers show predictable patterns of expression in human breast cancer patient samples of the same type [25, 67, 112], supporting the biological significance of this approach. The second is by using assays of enhancer-promoter looping, such as 3C-based sequencing approaches or ChIA-PET, which provide an indication of physical communication between enhancers and their target promoters. These approaches have shown that ERα binding sites that loop to target gene promoters are more likely to be enriched for other features of active enhancers than those that do not loop [25].
Although promoters that are looped to by an enhancer are likely to be true targets of the enhancer, looping does not always specify an active transcriptional outcome. For example, the ERα L540Q mutant (leucine 540 mutated to glutamine) [113,114,115] still promotes enhancer-promoter looping, even though it is transcriptionally impaired [67]. Furthermore, inhibition of enhancer transcription, a mark of active enhancers, by the transcription inhibitor flavopiridol does not prevent enhancer-promoter looping [25]. These results suggest that although definitive detection of enhancer-promoter loops is an effective way to identify bona fide target genes, the functional outcomes of looping may not be straightforward or always lead to productive transcriptional outcomes.
7.3 Enhancer Landscapes and Transcriptional Outcomes in Breast Cancers
Recent studies have explored unbiased approaches for the identification of functional enhancers in breast cancer cells that do not start with a TF cistrome [112]. In this regard, a recent study identified regulatory elements in cell lines representing the five distinct molecular subtypes of breast cancer by characterizing the epigenomic and transcriptomic profiles of the cells. The profiles of histone marks (i.e., H3K27ac and H3K4me1) from ChIP-seq were integrated with a measure enhancer transcription, as defined by GRO-seq. Putative TFs acting on these enhancers were identified by exploring the underlying sequence motifs in the defined enhancer regions and by integrating expression levels of the TFs defined by RNA-seq in each cell line. This integrative analysis produced as its outcome a “total functional score enhancer elements (TFSEE)” which ultimately allowed for the identification of subtype-specific enhancers and their cognate TFs that could play a functional role in biology of breast cancers [112]. One advantage of this approach is that it requires no prior knowledge of the TF of interest in a given cell type. The TFSEE approach can be further employed to evaluate the contribution of subtype-specific enhancers to therapeutic responses for the understanding of clinical outcomes.
8 Plasticity of ERα Transcription: Cross Talk of Nuclear Signaling Through ERα with Membrane-Initiated Signal Pathways
Genomic ERα action is modulated by diverse membrane-initiated signaling cascades. Membrane signaling can initiate from various extracellular stimuli, including growth factors, cytokines, chemokines, and estrogen via membrane-bound receptors (Fig. 1). These membrane receptors stimulate downstream signaling that activates kinase cascades and impacts transcriptional responses by inducing regulatory posttranslation modification of ERα or by stimulating the activation of other TFs. The following sections will describe key features in the cross talk between genomic ERα activity and membrane-initiated signals.
8.1 Estrogen and cAMP Signaling
Cytoplasmic estrogen signaling may be mediated by a small pool of cytoplasmic membrane-associated ERs (Fig. 1). Among these is the G protein-coupled receptor 30 (GPR30) [116]. Estrogen signaling via GPR30 stimulates heterotrimeric G proteins, which in turn activate adenylate cyclase activity to increase intracellular cAMP. Estrogen-dependent increases in cAMP stimulate transcription driven by cAMP response elements (CREs) [117]. In the murine uterine and human breast cancer models, intracellular cAMP stimulates ERα phosphorylation and transcriptional activity [118]. Furthermore, increases in intracellular cAMP activate protein kinase A (PKA), which in turn translocates to the nucleus to activate the cAMP response element-binding protein (CREB). Functional interactions between ERα and CREB induce a transcriptional response involving a complex containing the CREB-binding protein (CBP). Interestingly, transcriptional cross talk between ERα and CREB may not rely on regulatory regions containing the CREs but rather may be dependent on regulatory regions containing EREs [37]. Although this mechanism remains to be fully evaluated on a genomic scale, these studies demonstrated that extensive cross talk between estrogen and cAMP signaling can regulate ERα transcriptional activity.
8.2 Estrogen and Growth Factor Signaling
Growth factors are powerful mitogens that promote cellular proliferation across normal and disease tissues, especially in the breast and the reproductive tract [119,120,121]. Activation of transmembrane growth factor receptors by ligands, such as the epidermal growth factor (EGF), stimulates receptor dimerization and intracellular kinase activity. The activated downstream signaling kinases include mitogen-activated protein kinase or extracellular-regulated kinase (MAPK/ERK), protein kinase B (PKB/AKT), and c-Jun N-terminal kinases (JNK) [122] (Fig. 1). The EGF signaling pathway is extensively linked with estrogen signaling [123]. The activation of ERα by EGF involves the direct phosphorylation of ERα by MAPK on serine 118. This phosphorylation is required for full activity of the AF-1 and estrogen-mediated transcription [124]. In MCF-7 cells, estrogen stimulation elicits a rapid increase in intracellular calcium, an important intracellular second messenger, which in turn results in the activation of MAPK [125]. Moreover, ERα interacts with ERK2 and JNK1 , downstream effectors in the MAPK pathway, across the genome in regulatory regions [5, 126]. The cross regulation between MAPK and ERα can be a critical component of transcriptional regulation in proliferative biological systems.
8.3 Estrogen and Proinflammatory Signaling
Transcriptional regulation by estrogen can be modulated by cross talk with proinflammatory signaling pathways . The contribution of these pathways to estrogen-regulated transcription has significant impact in breast cancer biology and can have either pro-proliferative or antiproliferative effects depending on the tumor context. Activation of proinflammatory pathways can occur by a variety of chemokines and cytokines, including tumor necrosis alpha (TNFα). The effects of proinflammatory signaling on estrogen-regulated transcription were originally attributed to the TNFα-induced downregulation of ERα protein, resulting in the inhibition of ERα-dependent transcription [127]. Activation of proinflammatory signaling results in the activation of NF-κB by the translocation of the p65 and p50 subunits to the nucleus for target gene regulation (Fig. 1). Extensive cross talk between ERα and NF-κB was revealed in genomic experiments, which showed that TNFα and estrogen signaling acts to redistribute ERα and NF-κB across the genome in MCF-7 cells, resulting in altered cistromes [89]. The redistribution of ERα is driven by the redistribution of the pioneer factor FoxA1 in response to TNFα signaling, which brings ERα to new binding sites across the genome that are only revealed in the presence of TNFα signaling . The activation of these latent ERα binding sites into active enhancers in the presence of TNFα regulates a unique set of target genes that are not modulated by either agent alone and are strongly associated with clinical outcomes in breast cancer [89]. These findings suggest an important role for the early and transient effects of proinflammatory signaling on estrogen-regulated transcription.
9 Insights into Aberrant Estrogen-Regulated Transcription in Breast Cancers
Luminal/ER-positive breast cancers are the most heterogeneous subtype according to gene expression, mutation spectrum, copy number mutations, and patient outcomes [85]. Gene amplification of certain loci, such as ESR1 (encoding ERα) and NCOA3 (encoding SRC3), has a causal role in tumorigenesis [128,129,130]. The hypothesis is that amplification of these genes results in an increased dosage of the expressed proteins, which have oncogenic roles. A similar effect has been observed with the gene encoding the EGF transmembrane tyrosine kinase receptor HER2 (ERBB2). Amplifications of ERBB2 resulting in increased HER2 expression are observed in approximately 20% of breast cancers and are associated with poor prognosis, increased risk for disease progression, and decreased overall survival [131]. Notably, HER2-elevated breast cancers exhibit distinct transcriptional signatures in ER-positive and ER-negative breast cancers, suggesting an important modulatory interaction with estrogen signaling and distinct responses to antiestrogen therapy [132]. Other studies have shown that gene amplification events influence other aspects of transcription, ultimately modulating gene expression outcomes. The following sections highlight some notable examples of altered ERα-mediated transcription in breast cancers.
9.1 Genomic Amplification of Regulatory Elements
Copy number gains do not always correlate with upregulated expression of the genes in the amplified loci [85]. Moreover, many amplified or rearranged regions of the cancer genome do not contain protein- or microRNA-encoding genes that are aberrantly expressed in cancers [133]. Recent evidence suggests that the genetic, structural, or epigenetic disruption of DNA regulatory elements, such as enhancers, represents a major contribution to breast cancer initiation and recurrence. Amplification of EREs is found in ER-positive luminal breast cancers. This can result in deregulated transcription via long-range chromatin interactions with target genes outside of the amplified regions, which can lead to cancer development and tamoxifen resistance [134]. Chronic exposure of normal breast progenitor cells to estrogenic chemicals further results in progressive accumulation of these amplified response elements. The amplification of regulatory regions allows for the synchronized transcriptional control of several genes located on distinct chromosomes through long-range chromatin interactions [135]. This model of transcriptional regulation differs from the prevalent models of enhancer-promoter regulation, which typically assume a 1:1 relationship between regulatory elements and their targets. Assessing sufficiency and necessity of regulatory regions for transcriptional outputs in all of these models remains a challenge and an area of active research. The mechanisms underlying these chromosomal alterations and the functional characterization of these aberrations remain to be fully explored. Nonetheless, estrogen-dependent accumulation and modulation of chromatin interactions are thought to be a driving force of genomic instability driving breast cancer tumorigenesis.
9.2 Gain-of-Function ERα Mutations
Mutations in the gene encoding ERα, ESR1, are frequently detected in ER-positive metastatic breast cancers. These mutations are clustered in a “hotspot” which produce mutants in the ligand-binding domain of the expressed ERα. The most common of these mutations, Y537S and D538G, which appear as therapy-related mutations, increase association of ERα with coregulators in the absence of ligand by stabilizing the agonist conformation of helix 12 [136, 137]. These mutations lead to ligand-independent and enhanced ligand-dependent ERα activity that promotes tumor growth and partial resistance to endocrine therapy and may potentially enhance metastatic capacity [136, 137]. However, there is still a gap in our understanding of the molecular mechanisms and genomic effects of enhanced transcription by these ERα mutants, which are thought to involve selective interactions with coregulators or other transcription factors that modulate chromatin binding by ERα. The coregulators SRC3 (NCOA3) and p300 are essential for growth of the breast cancer cells expressing the Y537S mutant [138]. Given that recruitment of p300 may be the rate-limiting step for full ERα enhancer activation [67], it is possible that the enhanced transcriptional output of the Y537S involves a stabilization or over-activation of ERα-enhancers. Understanding the relevant mechanisms would help to identify targetable features of these gain-of-function mutants in endocrine-resistant breast cancer tumors with clinical utility.
10 Summary, Conclusions, and Perspectives
Estrogen-regulated transcription through ERα contributes in important ways to the biology breast cancer, as well as other developmental and physiological systems that depend on estrogen signaling. Tumorigenic estrogen-regulated transcription is induced by genetic mutations that affect (1) ERα expression, posttranslational modifications, and coregulator interactions, (2) regulatory element function, or (3) alterations to chromatin conformation that modify the genomic landscape. Our ability to identify these alterations and measure their contribution to the estrogen-regulated transcriptome has greatly improved in the last decade. As discoveries such as these continue to be made, our understanding of the mechanisms governing estrogen-regulated transcriptional responses will continue to evolve.
The historical molecular techniques that have provided valuable insights into the process of estrogen-regulated transcription have been replaced with more robust and sensitive deep sequencing-based genomic technologies. Approaches that interrogate the nascent transcriptional response to cellular stimuli, such as GRO-seq, have uncovered novel insights into the mechanism of estrogen transcriptional responses, as well as the ERα enhancers that control them. These studies have revealed that estrogen regulates transcription by RNA Pol I, II and III at annotated and unannotated genomic regions, controlling the expression of nearly every class of transcript described to date. Notably, intergenic transcripts produced from ERα binding sites (i.e., eRNAs) represent a novel feature of ERα enhancer biology that is shared with enhancers formed by other TFs. ERα enhancer RNAs have been used as measure of enhancer activity to understand the plasticity of the ERα enhancers in the context of various stimuli. Overall, the picture that has emerged of estrogen-regulated transcription is that it is rapid, robust, extensive, and transient.
While our understanding of the estrogen-regulated transcriptome has advanced dramatically with the use of genomics, a number of challenges and gaps in our understanding remain. These include methodological challenges, such as (1) single cell analyses of transcriptomes, cistromes, chromatin accessibility, and looping, (2) assessing allele-specific transcriptional effects, and (3) analyses in tissues and pathological samples. They also include knowledge gaps related to (1) the kinetics of the transcription process (e.g., enhancer formation, looping, target gene activation), (2) the decommissioning of active enhancers as signaling wanes, and (3) active and passive repression by the ERα. Successful resolution of these and other methodological challenges and knowledge gaps will help to advance our understanding of the estrogen-regulated transcriptome and its relevance to human physiology and disease.
References
Couse JF, Korach KS (1999) Estrogen receptor null mice: what have we learned and where will they lead us? Endocr Rev 20(3):358–417. https://doi.org/10.1210/edrv.20.3.0370
Warner M, Nilsson S, Gustafsson JA (1999) The estrogen receptor family. Curr Opin Obstet Gynecol 11(3):249–254
Welboren WJ, Sweep FC, Span PN, Stunnenberg HG (2009) Genomic actions of estrogen receptor alpha: what are the targets and how are they regulated? Endocr Relat Cancer 16(4):1073–1089. https://doi.org/10.1677/ERC-09-0086
Fan W, Chang J, Fu P (2015) Endocrine therapy resistance in breast cancer: current status, possible mechanisms and overcoming strategies. Future Med Chem 7(12):1511–1519. https://doi.org/10.4155/fmc.15.93
Madak-Erdogan Z, Lupien M, Stossi F, Brown M, Katzenellenbogen BS (2011) Genomic collaboration of estrogen receptor alpha and extracellular signal-regulated kinase 2 in regulating gene and proliferation programs. Mol Cell Biol 31(1):226–236. https://doi.org/10.1128/MCB.00821-10
Madak-Erdogan Z, Gong P, Katzenellenbogen BS (2016) Differential utilization of nuclear and extranuclear receptor signaling pathways in the actions of estrogens, SERMs, and a tissue-selective estrogen complex (TSEC). J Steroid Biochem Mol Biol 158:198–206. https://doi.org/10.1016/j.jsbmb.2015.12.008
Adlanmerini M, Solinhac R, Abot A, Fabre A, Raymond-Letron I, Guihot AL, Boudou F, Sautier L, Vessieres E, Kim SH, Liere P, Fontaine C, Krust A, Chambon P, Katzenellenbogen JA, Gourdy P, Shaul PW, Henrion D, Arnal JF, Lenfant F (2014) Mutation of the palmitoylation site of estrogen receptor alpha in vivo reveals tissue-specific roles for membrane versus nuclear actions. Proc Natl Acad Sci U S A 111(2):E283–E290. https://doi.org/10.1073/pnas.1322057111
Meitzen J, Luoma JI, Boulware MI, Hedges VL, Peterson BM, Tuomela K, Britson KA, Mermelstein PG (2013) Palmitoylation of estrogen receptors is essential for neuronal membrane signaling. Endocrinology 154(11):4293–4304. https://doi.org/10.1210/en.2013-1172
Acconcia F, Ascenzi P, Bocedi A, Spisni E, Tomasi V, Trentalance A, Visca P, Marino M (2005) Palmitoylation-dependent estrogen receptor alpha membrane localization: regulation by 17beta-estradiol. Mol Biol Cell 16(1):231–237. https://doi.org/10.1091/mbc.E04-07-0547
Filardo EJ, Thomas P (2012) Minireview: G protein-coupled estrogen receptor-1, GPER-1: its mechanism of action and role in female reproductive cancer, renal and vascular physiology. Endocrinology 153(7):2953–2962. https://doi.org/10.1210/en.2012-1061
Carroll JS, Liu XS, Brodsky AS, Li W, Meyer CA, Szary AJ, Eeckhoute J, Shao W, Hestermann EV, Geistlinger TR, Fox EA, Silver PA, Brown M (2005) Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122(1):33–43. https://doi.org/10.1016/j.cell.2005.05.008
Carroll JS, Meyer CA, Song J, Li W, Geistlinger TR, Eeckhoute J, Brodsky AS, Keeton EK, Fertuck KC, Hall GF, Wang Q, Bekiranov S, Sementchenko V, Fox EA, Silver PA, Gingeras TR, Liu XS, Brown M (2006) Genome-wide analysis of estrogen receptor binding sites. Nat Genet 38(11):1289–1297. https://doi.org/10.1038/ng1901
Lin CY, Vega VB, Thomsen JS, Zhang T, Kong SL, Xie M, Chiu KP, Lipovich L, Barnett DH, Stossi F, Yeo A, George J, Kuznetsov VA, Lee YK, Charn TH, Palanisamy N, Miller LD, Cheung E, Katzenellenbogen BS, Ruan Y, Bourque G, Wei CL, Liu ET (2007) Whole-genome cartography of estrogen receptor alpha binding sites. PLoS Genet 3(6):e87. https://doi.org/10.1371/journal.pgen.0030087
Hewitt SC, Li L, Grimm SA, Chen Y, Liu L, Li Y, Bushel PR, Fargo D, Korach KS (2012) Research resource: whole-genome estrogen receptor alpha binding in mouse uterine tissue revealed by ChIP-seq. Mol Endocrinol 26(5):887–898. https://doi.org/10.1210/me.2011-1311
Welboren WJ, van Driel MA, Janssen-Megens EM, van Heeringen SJ, Sweep FC, Span PN, Stunnenberg HG (2009) ChIP-Seq of ERalpha and RNA polymerase II defines genes differentially responding to ligands. EMBO J 28(10):1418–1428. https://doi.org/10.1038/emboj.2009.88
Stender JD, Kim K, Charn TH, Komm B, Chang KC, Kraus WL, Benner C, Glass CK, Katzenellenbogen BS (2010) Genome-wide analysis of estrogen receptor alpha DNA binding and tethering mechanisms identifies Runx1 as a novel tethering factor in receptor-mediated transcriptional activation. Mol Cell Biol 30(16):3943–3955. https://doi.org/10.1128/MCB.00118-10
Heldring N, Isaacs GD, Diehl AG, Sun M, Cheung E, Ranish JA, Kraus WL (2011) Multiple sequence-specific DNA-binding proteins mediate estrogen receptor signaling through a tethering pathway. Mol Endocrinol 25(4):564–574. https://doi.org/10.1210/me.2010-0425
Tan SK, Lin ZH, Chang CW, Varang V, Chng KR, Pan YF, Yong EL, Sung WK, Cheung E (2011) AP-2gamma regulates oestrogen receptor-mediated long-range chromatin interaction and gene transcription. EMBO J 30(13):2569–2581. https://doi.org/10.1038/emboj.2011.151
Hurtado A, Holmes KA, Ross-Innes CS, Schmidt D, Carroll JS (2011) FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat Genet 43(1):27–33. https://doi.org/10.1038/ng.730
Acevedo ML, Kraus WL (2004) Transcriptional activation by nuclear receptors. Essays Biochem 40:73–88
Biddie SC, John S, Hager GL (2010) Genome-wide mechanisms of nuclear receptor action. Trends Endocrinol Metab 21(1):3–9. https://doi.org/10.1016/j.tem.2009.08.006
Glass CK, Rosenfeld MG (2000) The coregulator exchange in transcriptional functions of nuclear receptors. Genes Dev 14(2):121–141
Lonard DM, O’Malley BW (2012) Nuclear receptor coregulators: modulators of pathology and therapeutic targets. Nat Rev Endocrinol 8(10):598–604. https://doi.org/10.1038/nrendo.2012.100
Foulds CE, Feng Q, Ding C, Bailey S, Hunsaker TL, Malovannaya A, Hamilton RA, Gates LA, Zhang Z, Li C, Chan D, Bajaj A, Callaway CG, Edwards DP, Lonard DM, Tsai SY, Tsai MJ, Qin J, O’Malley BW (2013) Proteomic analysis of coregulators bound to ERalpha on DNA and nucleosomes reveals coregulator dynamics. Mol Cell 51(2):185–199. https://doi.org/10.1016/j.molcel.2013.06.007
Hah N, Murakami S, Nagari A, Danko CG, Kraus WL (2013) Enhancer transcripts mark active estrogen receptor binding sites. Genome Res 23(8):1210–1223. https://doi.org/10.1101/gr.152306.112
Fullwood MJ, Liu MH, Pan YF, Liu J, Xu H, Mohamed YB, Orlov YL, Velkov S, Ho A, Mei PH, Chew EG, Huang PY, Welboren WJ, Han Y, Ooi HS, Ariyaratne PN, Vega VB, Luo Y, Tan PY, Choy PY, Wansa KD, Zhao B, Lim KS, Leow SC, Yow JS, Joseph R, Li H, Desai KV, Thomsen JS, Lee YK, Karuturi RK, Herve T, Bourque G, Stunnenberg HG, Ruan X, Cacheux-Rataboul V, Sung WK, Liu ET, Wei CL, Cheung E, Ruan Y (2009) An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462(7269):58–64. https://doi.org/10.1038/nature08497
Levine M, Cattoglio C, Tjian R (2014) Looping back to leap forward: transcription enters a new era. Cell 157(1):13–25. https://doi.org/10.1016/j.cell.2014.02.009
Plank JL, Dean A (2014) Enhancer function: mechanistic and genome-wide insights come together. Mol Cell 55(1):5–14. https://doi.org/10.1016/j.molcel.2014.06.015
Hah N, Danko CG, Core L, Waterfall JJ, Siepel A, Lis JT, Kraus WL (2011) A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell 145(4):622–634. https://doi.org/10.1016/j.cell.2011.03.042
Hah N, Kraus WL (2014) Hormone-regulated transcriptomes: lessons learned from estrogen signaling pathways in breast cancer cells. Mol Cell Endocrinol 382(1):652–664. https://doi.org/10.1016/j.mce.2013.06.021
McDevitt MA, Glidewell-Kenney C, Jimenez MA, Ahearn PC, Weiss J, Jameson JL, Levine JE (2008) New insights into the classical and non-classical actions of estrogen: evidence from estrogen receptor knock-out and knock-in mice. Mol Cell Endocrinol 290(1–2):24–30. https://doi.org/10.1016/j.mce.2008.04.003
Klein-Hitpass L, Schorpp M, Wagner U, Ryffel GU (1986) An estrogen-responsive element derived from the 5′ flanking region of the Xenopus vitellogenin A2 gene functions in transfected human cells. Cell 46(7):1053–1061
Klinge CM (2001) Estrogen receptor interaction with estrogen response elements. Nucleic Acids Res 29(14):2905–2919
Safe S (2001) Transcriptional activation of genes by 17 beta-estradiol through estrogen receptor-Sp1 interactions. Vitam Horm 62:231–252
Ray A, Prefontaine KE, Ray P (1994) Down-modulation of interleukin-6 gene expression by 17 beta-estradiol in the absence of high affinity DNA binding by the estrogen receptor. J Biol Chem 269(17):12940–12946
Kushner PJ, Agard DA, Greene GL, Scanlan TS, Shiau AK, Uht RM, Webb P (2000) Estrogen receptor pathways to AP-1. J Steroid Biochem Mol Biol 74(5):311–317
Lazennec G, Thomas JA, Katzenellenbogen BS (2001) Involvement of cyclic AMP response element binding protein (CREB) and estrogen receptor phosphorylation in the synergistic activation of the estrogen receptor by estradiol and protein kinase activators. J Steroid Biochem Mol Biol 77(4–5):193–203
Deplancke B, Alpern D, Gardeux V (2016) The genetics of transcription factor DNA binding variation. Cell 166(3):538–554. https://doi.org/10.1016/j.cell.2016.07.012
Fiorito E, Katika MR, Hurtado A (2013) Cooperating transcription factors mediate the function of estrogen receptor. Chromosoma 122(1–2):1–12. https://doi.org/10.1007/s00412-012-0392-7
Pradeepa MM (2017) Causal role of histone acetylations in enhancer function. Transcription 8(1):40–47. https://doi.org/10.1080/21541264.2016.1253529
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, Wang W, Weng Z, Green RD, Crawford GE, Ren B (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39(3):311–318. https://doi.org/10.1038/ng1966
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, Boyer LA, Young RA, Jaenisch R (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A 107(50):21931–21936. https://doi.org/10.1073/pnas.1016071107
Visel A, Blow MJ, Li Z, Zhang T, Akiyama JA, Holt A, Plajzer-Frick I, Shoukry M, Wright C, Chen F, Afzal V, Ren B, Rubin EM, Pennacchio LA (2009) ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457(7231):854–858. https://doi.org/10.1038/nature07730
Magnani L, Stoeck A, Zhang X, Lanczky A, Mirabella AC, Wang TL, Gyorffy B, Lupien M (2013) Genome-wide reprogramming of the chromatin landscape underlies endocrine therapy resistance in breast cancer. Proc Natl Acad Sci U S A 110(16):E1490–E1499. https://doi.org/10.1073/pnas.1219992110
Melgar MF, Collins FS, Sethupathy P (2011) Discovery of active enhancers through bidirectional expression of short transcripts. Genome Biol 12(11):R113. https://doi.org/10.1186/gb-2011-12-11-r113
Natoli G, Andrau JC (2012) Noncoding transcription at enhancers: general principles and functional models. Annu Rev Genet 46:1–19. https://doi.org/10.1146/annurev-genet-110711-155459
Liu MH, Cheung E (2014) Estrogen receptor-mediated long-range chromatin interactions and transcription in breast cancer. Mol Cell Endocrinol 382(1):624–632. https://doi.org/10.1016/j.mce.2013.09.019
Heinz S, Romanoski CE, Benner C, Glass CK (2015) The selection and function of cell type-specific enhancers. Nat Rev Mol Cell Biol 16(3):144–154. https://doi.org/10.1038/nrm3949
Pennacchio LA, Bickmore W, Dean A, Nobrega MA, Bejerano G (2013) Enhancers: five essential questions. Nat Rev Genet 14(4):288–295. https://doi.org/10.1038/nrg3458
Yi P, Wang Z, Feng Q, Pintilie GD, Foulds CE, Lanz RB, Ludtke SJ, Schmid MF, Chiu W, O’Malley BW (2015) Structure of a biologically active estrogen receptor-coactivator complex on DNA. Mol Cell 57(6):1047–1058. https://doi.org/10.1016/j.molcel.2015.01.025
Metivier R, Penot G, Hubner MR, Reid G, Brand H, Kos M, Gannon F (2003) Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell 115(6):751–763
Kamei Y, Xu L, Heinzel T, Torchia J, Kurokawa R, Gloss B, Lin SC, Heyman RA, Rose DW, Glass CK, Rosenfeld MG (1996) A CBP integrator complex mediates transcriptional activation and AP-1 inhibition by nuclear receptors. Cell 85(3):403–414
Acevedo ML, Kraus WL (2003) Mediator and p300/CBP-steroid receptor coactivator complexes have distinct roles, but function synergistically, during estrogen receptor alpha-dependent transcription with chromatin templates. Mol Cell Biol 23(1):335–348
Kim MY, Hsiao SJ, Kraus WL (2001) A role for coactivators and histone acetylation in estrogen receptor alpha-mediated transcription initiation. EMBO J 20(21):6084–6094. https://doi.org/10.1093/emboj/20.21.6084
Belandia B, Orford RL, Hurst HC, Parker MG (2002) Targeting of SWI/SNF chromatin remodelling complexes to estrogen-responsive genes. EMBO J 21(15):4094–4103
Chen W, Roeder RG (2011) Mediator-dependent nuclear receptor function. Semin Cell Dev Biol 22(7):749–758. https://doi.org/10.1016/j.semcdb.2011.07.026
Malik S, Roeder RG (2010) The metazoan Mediator co-activator complex as an integrative hub for transcriptional regulation. Nat Rev Genet 11(11):761–772. https://doi.org/10.1038/nrg2901
Plaschka C, Nozawa K, Cramer P (2016) Mediator architecture and RNA polymerase II interaction. J Mol Biol 428(12):2569–2574. https://doi.org/10.1016/j.jmb.2016.01.028
Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA, Shiekhattar R (2013) Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 494(7438):497–501. https://doi.org/10.1038/nature11884
Foulds CE, Feng Q, Ding C, Bailey S, Hunsaker TL, Malovannaya A, Hamilton RA, Gates LA, Zhang Z, Li C, Chan D, Bajaj A, Callaway CG, Edwards DP, Lonard DM, Tsai SY, Tsai MJ, Qin J, O’Malley BW (2013) Proteomic analysis of coregulators bound to ERα on DNA and nucleosomes reveals coregulator dynamics. Mol Cell 51(2):185–199. https://doi.org/10.1016/j.molcel.2013.06.007
Lanz RB, McKenna NJ, Onate SA, Albrecht U, Wong J, Tsai SY, Tsai MJ, O’Malley BW (1999) A steroid receptor coactivator, SRA, functions as an RNA and is present in an SRC-1 complex. Cell 97(1):17–27
Coleman KM, Lam V, Jaber BM, Lanz RB, Smith CL (2004) SRA coactivation of estrogen receptor-alpha is phosphorylation-independent, and enhances 4-hydroxytamoxifen agonist activity. Biochem Biophys Res Commun 323(1):332–338. https://doi.org/10.1016/j.bbrc.2004.08.090
Deblois G, Giguere V (2003) Ligand-independent coactivation of ERalpha AF-1 by steroid receptor RNA activator (SRA) via MAPK activation. J Steroid Biochem Mol Biol 85(2–5):123–131
Jin Q, Yu LR, Wang L, Zhang Z, Kasper LH, Lee JE, Wang C, Brindle PK, Dent SY, Ge K (2011) Distinct roles of GCN5/PCAF-mediated H3K9ac and CBP/p300-mediated H3K18/27ac in nuclear receptor transactivation. EMBO J 30(2):249–262. https://doi.org/10.1038/emboj.2010.318
Tie F, Banerjee R, Stratton CA, Prasad-Sinha J, Stepanik V, Zlobin A, Diaz MO, Scacheri PC, Harte PJ (2009) CBP-mediated acetylation of histone H3 lysine 27 antagonizes Drosophila Polycomb silencing. Development 136(18):3131–3141. https://doi.org/10.1242/dev.037127
Bose DA, Donahue G, Reinberg D, Shiekhattar R, Bonasio R, Berger SL (2017) RNA binding to CBP stimulates histone acetylation and transcription. Cell 168(1–2):135–149 e122. https://doi.org/10.1016/j.cell.2016.12.020
Murakami S, Nagari A, Kraus WL (2017) Dynamic assembly and activation of estrogen receptor alpha enhancers through coregulator switching. Genes Dev 31(15):1535–1548. https://doi.org/10.1101/gad.302182.117
Herschman HR (1991) Primary response genes induced by growth factors and tumor promoters. Annu Rev Biochem 60:281–319. https://doi.org/10.1146/annurev.bi.60.070191.001433
Winkles JA (1998) Serum- and polypeptide growth factor-inducible gene expression in mouse fibroblasts. Prog Nucleic Acid Res Mol Biol 58:41–78
Kininis M, Isaacs GD, Core LJ, Hah N, Kraus WL (2009) Postrecruitment regulation of RNA polymerase II directs rapid signaling responses at the promoters of estrogen target genes. Mol Cell Biol 29(5):1123–1133. https://doi.org/10.1128/MCB.00841-08
Liu X, Kraus WL, Bai X (2015) Ready, pause, go: regulation of RNA polymerase II pausing and release by cellular signaling pathways. Trends Biochem Sci 40(9):516–525. https://doi.org/10.1016/j.tibs.2015.07.003
Core LJ, Waterfall JJ, Lis JT (2008) Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322(5909):1845–1848. https://doi.org/10.1126/science.1162228
Schena M, Shalon D, Davis RW, Brown PO (1995) Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270(5235):467–470
Kraus WL (2015) Editorial: would you like a hypothesis with those data? Omics and the age of discovery science. Mol Endocrinol 29(11):1531–1534. https://doi.org/10.1210/me.2015-1253
Calviello L, Ohler U (2017) Beyond read-counts: Ribo-seq data analysis to understand the functions of the transcriptome. Trends Genet 33(10):728–744. https://doi.org/10.1016/j.tig.2017.08.003
Shalem O, Sanjana NE, Zhang F (2015) High-throughput functional genomics using CRISPR-Cas9. Nat Rev Genet 16(5):299–311. https://doi.org/10.1038/nrg3899
Root DE, Hacohen N, Hahn WC, Lander ES, Sabatini DM (2006) Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat Methods 3(9):715–719. https://doi.org/10.1038/nmeth924
Kininis M, Kraus WL (2008) A global view of transcriptional regulation by nuclear receptors: gene expression, factor localization, and DNA sequence analysis. Nucl Recept Signal 6:e005
Cheung E, Kraus WL (2010) Genomic analyses of hormone signaling and gene regulation. Annu Rev Physiol 72:191–218. https://doi.org/10.1146/annurev-physiol-021909-135840
Carroll JS, Brown M (2006) Estrogen receptor target gene: an evolving concept. Mol Endocrinol 20(8):1707–1714. https://doi.org/10.1210/me.2005-0334
Zhang J, Poh HM, Peh SQ, Sia YY, Li G, Mulawadi FH, Goh Y, Fullwood MJ, Sung WK, Ruan X, Ruan Y (2012) ChIA-PET analysis of transcriptional chromatin interactions. Methods 58(3):289–299. https://doi.org/10.1016/j.ymeth.2012.08.009
Welboren WJ, Stunnenberg HG, Sweep FC, Span PN (2007) Identifying estrogen receptor target genes. Mol Oncol 1(2):138–143. https://doi.org/10.1016/j.molonc.2007.04.001
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752. https://doi.org/10.1038/35021093
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Lonning PE, Borresen-Dale AL (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A 98(19):10869–10874. https://doi.org/10.1073/pnas.191367098
Cancer Genome Atlas N (2012) Comprehensive molecular portraits of human breast tumours. Nature 490(7418):61–70. https://doi.org/10.1038/nature11412
van’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536. https://doi.org/10.1038/415530a
Lin CY, Strom A, Vega VB, Kong SL, Yeo AL, Thomsen JS, Chan WC, Doray B, Bangarusamy DK, Ramasamy A, Vergara LA, Tang S, Chong A, Bajic VB, Miller LD, Gustafsson JA, Liu ET (2004) Discovery of estrogen receptor alpha target genes and response elements in breast tumor cells. Genome Biol 5(9):R66. https://doi.org/10.1186/gb-2004-5-9-r66
Dai X, Li T, Bai Z, Yang Y, Liu X, Zhan J, Shi B (2015) Breast cancer intrinsic subtype classification, clinical use and future trends. Am J Cancer Res 5(10):2929–2943
Franco HL, Nagari A, Kraus WL (2015) TNFalpha signaling exposes latent estrogen receptor binding sites to alter the breast cancer cell transcriptome. Mol Cell 58(1):21–34. https://doi.org/10.1016/j.molcel.2015.02.001
Mahat DB, Kwak H, Booth GT, Jonkers IH, Danko CG, Patel RK, Waters CT, Munson K, Core LJ, Lis JT (2016) Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq). Nat Protoc 11(8):1455–1476. https://doi.org/10.1038/nprot.2016.086
Danko CG, Hah N, Luo X, Martins AL, Core L, Lis JT, Siepel A, Kraus WL (2013) Signaling pathways differentially affect RNA polymerase II initiation, pausing, and elongation rate in cells. Mol Cell 50(2):212–222. https://doi.org/10.1016/j.molcel.2013.02.015
Chae M, Danko CG, Kraus WL (2015) groHMM: a computational tool for identifying unannotated and cell type-specific transcription units from global run-on sequencing data. BMC Bioinformatics 16:222. https://doi.org/10.1186/s12859-015-0656-3
Sun M, Gadad SS, Kim DS, Kraus WL (2015) Discovery, annotation, and functional analysis of long noncoding RNAs controlling cell-cycle gene expression and proliferation in breast cancer cells. Mol Cell 59(4):698–711. https://doi.org/10.1016/j.molcel.2015.06.023
De Santa F, Barozzi I, Mietton F, Ghisletti S, Polletti S, Tusi BK, Muller H, Ragoussis J, Wei CL, Natoli G (2010) A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol 8(5):e1000384. https://doi.org/10.1371/journal.pbio.1000384
Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S, Markenscoff-Papadimitriou E, Kuhl D, Bito H, Worley PF, Kreiman G, Greenberg ME (2010) Widespread transcription at neuronal activity-regulated enhancers. Nature 465(7295):182–187. https://doi.org/10.1038/nature09033
Wang D, Garcia-Bassets I, Benner C, Li W, Su X, Zhou Y, Qiu J, Liu W, Kaikkonen MU, Ohgi KA, Glass CK, Rosenfeld MG, Fu XD (2011) Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474(7351):390–394. https://doi.org/10.1038/nature10006
Lam MT, Cho H, Lesch HP, Gosselin D, Heinz S, Tanaka-Oishi Y, Benner C, Kaikkonen MU, Kim AS, Kosaka M, Lee CY, Watt A, Grossman TR, Rosenfeld MG, Evans RM, Glass CK (2013) Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature 498(7455):511–515. https://doi.org/10.1038/nature12209
Li W, Notani D, Ma Q, Tanasa B, Nunez E, Chen AY, Merkurjev D, Zhang J, Ohgi K, Song X, Oh S, Kim HS, Glass CK, Rosenfeld MG (2013) Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498(7455):516–520. https://doi.org/10.1038/nature12210
Hsieh CL, Fei T, Chen Y, Li T, Gao Y, Wang X, Sun T, Sweeney CJ, Lee GS, Chen S, Balk SP, Liu XS, Brown M, Kantoff PW (2014) Enhancer RNAs participate in androgen receptor-driven looping that selectively enhances gene activation. Proc Natl Acad Sci U S A 111(20):7319–7324. https://doi.org/10.1073/pnas.1324151111
Arner E, Daub CO, Vitting-Seerup K, Andersson R, Lilje B, Drablos F, Lennartsson A, Ronnerblad M, Hrydziuszko O, Vitezic M, Freeman TC, Alhendi AM, Arner P, Axton R, Baillie JK, Beckhouse A, Bodega B, Briggs J, Brombacher F, Davis M, Detmar M, Ehrlund A, Endoh M, Eslami A, Fagiolini M, Fairbairn L, Faulkner GJ, Ferrai C, Fisher ME, Forrester L, Goldowitz D, Guler R, Ha T, Hara M, Herlyn M, Ikawa T, Kai C, Kawamoto H, Khachigian LM, Klinken SP, Kojima S, Koseki H, Klein S, Mejhert N, Miyaguchi K, Mizuno Y, Morimoto M, Morris KJ, Mummery C, Nakachi Y, Ogishima S, Okada-Hatakeyama M, Okazaki Y, Orlando V, Ovchinnikov D, Passier R, Patrikakis M, Pombo A, Qin XY, Roy S, Sato H, Savvi S, Saxena A, Schwegmann A, Sugiyama D, Swoboda R, Tanaka H, Tomoiu A, Winteringham LN, Wolvetang E, Yanagi-Mizuochi C, Yoneda M, Zabierowski S, Zhang P, Abugessaisa I, Bertin N, Diehl AD, Fukuda S, Furuno M, Harshbarger J, Hasegawa A, Hori F, Ishikawa-Kato S, Ishizu Y, Itoh M, Kawashima T, Kojima M, Kondo N, Lizio M, Meehan TF, Mungall CJ, Murata M, Nishiyori-Sueki H, Sahin S, Nagao-Sato S, Severin J, de Hoon MJ, Kawai J, Kasukawa T, Lassmann T, Suzuki H, Kawaji H, Summers KM, Wells C, Consortium F, Hume DA, Forrest AR, Sandelin A, Carninci P, Hayashizaki Y (2015) Transcribed enhancers lead waves of coordinated transcription in transitioning mammalian cells. Science 347(6225):1010–1014. https://doi.org/10.1126/science.1259418
Fang B, Everett LJ, Jager J, Briggs E, Armour SM, Feng D, Roy A, Gerhart-Hines Z, Sun Z, Lazar MA (2014) Circadian enhancers coordinate multiple phases of rhythmic gene transcription in vivo. Cell 159(5):1140–1152. https://doi.org/10.1016/j.cell.2014.10.022
Mousavi K, Zare H, Dell’orso S, Grontved L, Gutierrez-Cruz G, Derfoul A, Hager GL, Sartorelli V (2013) eRNAs promote transcription by establishing chromatin accessibility at defined genomic loci. Mol Cell 51(5):606–617. https://doi.org/10.1016/j.molcel.2013.07.022
Li W, Notani D, Rosenfeld MG (2016) Enhancers as non-coding RNA transcription units: recent insights and future perspectives. Nat Rev Genet 17(4):207–223. https://doi.org/10.1038/nrg.2016.4
Lander ES (2011) Initial impact of the sequencing of the human genome. Nature 470(7333):187–197. https://doi.org/10.1038/nature09792
Clark MB, Amaral PP, Schlesinger FJ, Dinger ME, Taft RJ, Rinn JL, Ponting CP, Stadler PF, Morris KV, Morillon A, Rozowsky JS, Gerstein MB, Wahlestedt C, Hayashizaki Y, Carninci P, Gingeras TR, Mattick JS (2011) The reality of pervasive transcription. PLoS Biol 9(7):e1000625.; discussion e1001102. https://doi.org/10.1371/journal.pbio.1000625
Heintzman ND, Hon GC, Hawkins RD, Kheradpour P, Stark A, Harp LF, Ye Z, Lee LK, Stuart RK, Ching CW, Ching KA, Antosiewicz-Bourget JE, Liu H, Zhang X, Green RD, Lobanenkov VV, Stewart R, Thomson JA, Crawford GE, Kellis M, Ren B (2009) Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459(7243):108–112. https://doi.org/10.1038/nature07829
Shlyueva D, Stampfel G, Stark A (2014) Transcriptional enhancers: from properties to genome-wide predictions. Nat Rev Genet 15(4):272–286. https://doi.org/10.1038/nrg3682
Calo E, Wysocka J (2013) Modification of enhancer chromatin: what, how, and why? Mol Cell 49(5):825–837. https://doi.org/10.1016/j.molcel.2013.01.038
Crawford GE, Holt IE, Whittle J, Webb BD, Tai D, Davis S, Margulies EH, Chen Y, Bernat JA, Ginsburg D, Zhou D, Luo S, Vasicek TJ, Daly MJ, Wolfsberg TG, Collins FS (2006) Genome-wide mapping of DNase hypersensitive sites using massively parallel signature sequencing (MPSS). Genome Res 16(1):123–131. https://doi.org/10.1101/gr.4074106
Sheffield NC, Thurman RE, Song L, Safi A, Stamatoyannopoulos JA, Lenhard B, Crawford GE, Furey TS (2013) Patterns of regulatory activity across diverse human cell types predict tissue identity, transcription factor binding, and long-range interactions. Genome Res 23(5):777–788. https://doi.org/10.1101/gr.152140.112
Core LJ, Martins AL, Danko CG, Waters CT, Siepel A, Lis JT (2014) Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat Genet 46(12):1311–1320. https://doi.org/10.1038/ng.3142
Franco HL, Nagari A, Malladi VS, Li W, Xi Y, Richardson D, Allton KL, Tanaka K, Li J, Murakami S, Keyomarsi K, Bedford MT, Shi X, Li W, Barton MC, Dent SYR, Kraus WL (2018) Enhancer transcription reveals subtype-specific gene expression programs controlling breast cancer pathogenesis. Genome Res 28(2):159–170. https://doi.org/10.1101/gr.226019.117
Ince BA, Zhuang Y, Wrenn CK, Shapiro DJ, Katzenellenbogen BS (1993) Powerful dominant negative mutants of the human estrogen receptor. J Biol Chem 268(19):14026–14032
Schodin DJ, Zhuang Y, Shapiro DJ, Katzenellenbogen BS (1995) Analysis of mechanisms that determine dominant negative estrogen receptor effectiveness. J Biol Chem 270(52):31163–31171
Acevedo ML, Lee KC, Stender JD, Katzenellenbogen BS, Kraus WL (2004) Selective recognition of distinct classes of coactivators by a ligand-inducible activation domain. Mol Cell 13(5):725–738
Prossnitz ER, Arterburn JB, Smith HO, Oprea TI, Sklar LA, Hathaway HJ (2008) Estrogen signaling through the transmembrane G protein-coupled receptor GPR30. Annu Rev Physiol 70:165–190. https://doi.org/10.1146/annurev.physiol.70.113006.100518
Aronica SM, Kraus WL, Katzenellenbogen BS (1994) Estrogen action via the cAMP signaling pathway: stimulation of adenylate cyclase and cAMP-regulated gene transcription. Proc Natl Acad Sci U S A 91(18):8517–8521
Aronica SM, Katzenellenbogen BS (1993) Stimulation of estrogen receptor-mediated transcription and alteration in the phosphorylation state of the rat uterine estrogen receptor by estrogen, cyclic adenosine monophosphate, and insulin-like growth factor-I. Mol Endocrinol 7(6):743–752. https://doi.org/10.1210/mend.7.6.7689695
Ignar-Trowbridge DM, Nelson KG, Bidwell MC, Curtis SW, Washburn TF, McLachlan JA, Korach KS (1992) Coupling of dual signaling pathways: epidermal growth factor action involves the estrogen receptor. Proc Natl Acad Sci U S A 89(10):4658–4662
Hayashi S, Sakamoto T, Inoue A, Yoshida N, Omoto Y, Yamaguchi Y (2003) Estrogen and growth factor signaling pathway: basic approaches for clinical application. J Steroid Biochem Mol Biol 86(3–5):433–442
Ribeiro JR, Freiman RN (2014) Estrogen signaling crosstalk: implications for endocrine resistance in ovarian cancer. J Steroid Biochem Mol Biol 143:160–173. https://doi.org/10.1016/j.jsbmb.2014.02.010
Wee P, Wang Z (2017) Epidermal growth factor receptor cell proliferation signaling pathways. Cancers (Basel) 9(5). https://doi.org/10.3390/cancers9050052
Gee JM, Robertson JF, Gutteridge E, Ellis IO, Pinder SE, Rubini M, Nicholson RI (2005) Epidermal growth factor receptor/HER2/insulin-like growth factor receptor signalling and oestrogen receptor activity in clinical breast cancer. Endocr Relat Cancer 12(Suppl 1):S99–S111. https://doi.org/10.1677/erc.1.01005
Kato S, Endoh H, Masuhiro Y, Kitamoto T, Uchiyama S, Sasaki H, Masushige S, Gotoh Y, Nishida E, Kawashima H, Metzger D, Chambon P (1995) Activation of the estrogen receptor through phosphorylation by mitogen-activated protein kinase. Science 270(5241):1491–1494
Improta-Brears T, Whorton AR, Codazzi F, York JD, Meyer T, McDonnell DP (1999) Estrogen-induced activation of mitogen-activated protein kinase requires mobilization of intracellular calcium. Proc Natl Acad Sci U S A 96(8):4686–4691
Sun M, Isaacs GD, Hah N, Heldring N, Fogarty EA, Kraus WL (2012) Estrogen regulates JNK1 genomic localization to control gene expression and cell growth in breast cancer cells. Mol Endocrinol 26(5):736–747. https://doi.org/10.1210/me.2011-1158
Lee SH, Nam HS (2008) TNF alpha-induced down-regulation of estrogen receptor alpha in MCF-7 breast cancer cells. Mol Cells 26(3):285–290
Holst F (2016) Estrogen receptor alpha gene amplification in breast cancer: 25 years of debate. World J Clin Oncol 7(2):160–173. https://doi.org/10.5306/wjco.v7.i2.160
Gutierrez C, Schiff R (2011) HER2: biology, detection, and clinical implications. Arch Pathol Lab Med 135(1):55–62. https://doi.org/10.1043/2010-0454-RAR.1
Gojis O, Rudraraju B, Gudi M, Hogben K, Sousha S, Coombes RC, Cleator S, Palmieri C (2010) The role of SRC-3 in human breast cancer. Nat Rev Clin Oncol 7(2):83–89. https://doi.org/10.1038/nrclinonc.2009.219
Menard S, Fortis S, Castiglioni F, Agresti R, Balsari A (2001) HER2 as a prognostic factor in breast cancer. Oncology 61(Suppl 2):67–72. https://doi.org/10.1159/000055404
Marchio C, Natrajan R, Shiu KK, Lambros MB, Rodriguez-Pinilla SM, Tan DS, Lord CJ, Hungermann D, Fenwick K, Tamber N, Mackay A, Palacios J, Sapino A, Buerger H, Ashworth A, Reis-Filho JS (2008) The genomic profile of HER2-amplified breast cancers: the influence of ER status. J Pathol 216(4):399–407. https://doi.org/10.1002/path.2423
Hampton OA, Den Hollander P, Miller CA, Delgado DA, Li J, Coarfa C, Harris RA, Richards S, Scherer SE, Muzny DM, Gibbs RA, Lee AV, Milosavljevic A (2009) A sequence-level map of chromosomal breakpoints in the MCF-7 breast cancer cell line yields insights into the evolution of a cancer genome. Genome Res 19(2):167–177. https://doi.org/10.1101/gr.080259.108
Hsu PY, Hsu HK, Lan X, Juan L, Yan PS, Labanowska J, Heerema N, Hsiao TH, Chiu YC, Chen Y, Liu Y, Li L, Li R, Thompson IM, Nephew KP, Sharp ZD, Kirma NB, Jin VX, Huang TH (2013) Amplification of distant estrogen response elements deregulates target genes associated with tamoxifen resistance in breast cancer. Cancer Cell 24(2):197–212. https://doi.org/10.1016/j.ccr.2013.07.007
Hsu PY, Hsu HK, Hsiao TH, Ye Z, Wang E, Profit AL, Jatoi I, Chen Y, Kirma NB, Jin VX, Sharp ZD, Huang TH (2016) Spatiotemporal control of estrogen-responsive transcription in ERalpha-positive breast cancer cells. Oncogene 35(18):2379–2389. https://doi.org/10.1038/onc.2015.298
Merenbakh-Lamin K, Ben-Baruch N, Yeheskel A, Dvir A, Soussan-Gutman L, Jeselsohn R, Yelensky R, Brown M, Miller VA, Sarid D, Rizel S, Klein B, Rubinek T, Wolf I (2013) D538G mutation in estrogen receptor-alpha: a novel mechanism for acquired endocrine resistance in breast cancer. Cancer Res 73(23):6856–6864. https://doi.org/10.1158/0008-5472.CAN-13-1197
Toy W, Shen Y, Won H, Green B, Sakr RA, Will M, Li Z, Gala K, Fanning S, King TA, Hudis C, Chen D, Taran T, Hortobagyi G, Greene G, Berger M, Baselga J, Chandarlapaty S (2013) ESR1 ligand-binding domain mutations in hormone-resistant breast cancer. Nat Genet 45(12):1439–1445. https://doi.org/10.1038/ng.2822
Jeselsohn R, Bergholz JS, Pun M, Cornwell M, Liu W, Nardone A, Xiao T, Li W, Qiu X, Buchwalter G, Feiglin A, Abell-Hart K, Fei T, Rao P, Long H, Kwiatkowski N, Zhang T, Gray N, Melchers D, Houtman R, Liu XS, Cohen O, Wagle N, Winer EP, Zhao J, Brown M (2018) Allele-specific chromatin recruitment and therapeutic vulnerabilities of ESR1 activating mutations. Cancer Cell 33(2):173–186 e175. https://doi.org/10.1016/j.ccell.2018.01.004
Acknowledgments
We are grateful for critical comments and suggestions from members of the Kraus lab, including Shino Murakami, Anusha Nagari, and J. Tyler Piazza. We are also thankful for the contributions of past Kraus lab members, whose studies furthering our understanding of the molecular mechanisms of estrogen-regulated transcription are highlighted in this review, including Nasun Hah, Charles Danko, Shino Murakami, Hector Franco, and Anusha Nagari. The estrogen-related studies in the Kraus lab are supported by grants from the NIH/NIDDK and the Cancer Prevention and Research Institute of Texas (CPRIT) to W.L.K. Y.M.V. is supported by a Postdoctoral Research Fellowship from the Lalor Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Vasquez, Y.M., Kraus, W.L. (2019). The Estrogen-Regulated Transcriptome: Rapid, Robust, Extensive, and Transient. In: Zhang, X. (eds) Estrogen Receptor and Breast Cancer. Cancer Drug Discovery and Development. Humana Press, Cham. https://doi.org/10.1007/978-3-319-99350-8_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-99350-8_5
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-99349-2
Online ISBN: 978-3-319-99350-8
eBook Packages: MedicineMedicine (R0)