Abstract
The centromere is the genetic locus that specifies the site of kinetochore assembly, where the chromosome will attach to the kinetochore microtubule. The pericentromere is the physical region responsible for the geometry of bi-oriented sister kinetochores in metaphase. In budding yeast the 125 bp point centromere is sufficient to specify kinetochore assembly. The flanking region is enriched (3X) in cohesin and condensin relative to the remaining chromosome arms. The enrichment spans about 30–50 kb around each centromere. We refer to the flanking chromatin as the pericentromere in yeast. In mammals, a 5–10 Mb region dictates where the kinetochore is built. The kinetochore interacts with a very small fraction of DNA on the surface of the centromeric region. The remainder of the centromere lies between the sister kinetochores. This is typically called centromere chromatin. The chromatin sites that directly interface to microtubules cannot be identified due to the repeated sequence within the mammalian centromere. However in both yeast and mammals, the total amount of DNA between the sites of microtubule attachment in metaphase is highly conserved. In yeast the 16 chromosomes are clustered into a 250 nm diameter region, and 800 kb (16 × 50 kb) or ~1 Mb of DNA lies between sister kinetochores. In mammals, 5–10 Mb lies between sister kinetochores. In both organisms the sister kinetochores are separated by about 1 μm. Thus, centromeres of different organisms differ in how they specify kinetochore assembly, but there may be important centromere chromatin functions that are conserved throughout phylogeny. Recently, centromeric chromatin has been reconstituted in vitro using alpha satellite DNA revealing unexpected features of centromeric DNA organization, replication, and response to stress. We will focus on the conserved features of centromere in this review.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Characteristics of Centromere Chromatin—Primary Folding of a Nucleosome
1.1 Centromere-Specific Histone H3 Variant, Cse4 Yeast, CENP-A Mammals
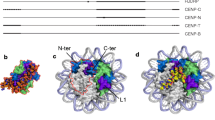
Centromeres contain an atypical Histone H3, known as CENP-A, or Cse4 in budding yeast (Earnshaw et al. 2013). The functional role of CENP-A at the centromere is considerably less clear. A variety of studies reveal that CENP-A containing nucleoprotein complexes can adopt a number of conformations, including tetramers (Dalal et al. 2007; Shivaraju et al. 2012), hemisomes (Henikoff et al. 2014) or octamers (Camahort et al. 2009; Hasson et al. 2013); canonical left-handed octamers (Dechassa et al. 2011; Hasson et al. 2013), right-handed hemisomes or octamers (Furuyama and Henikoff 2009; Diaz-Ingelmo et al. 2015), and hexamers (Mizuguchi et al. 2007; Xiao et al. 2011). In budding yeast, with its point centromere and purportedly single Cse4-containing nucleosome (Lawrimore et al. 2011; Shivaraju et al. 2012; Aravamudhan et al. 2013; Haase et al. 2013; Wisniewski et al. 2014), there is controversy over the number of Cse4 molecules and the handedness of DNA wrap. At the heart of such controversies is the quest to determine critical features responsible for establishing a functional kinetochore. For Cse4 and CENP-A, the ability to swap centromere-specific domains with canonical Histone H3 reveals crucial molecular determinants unique to the centromere nucleosome. These include the Histone Fold Domain and the essential N-terminus (END) in Cse4 (Keith et al. 1999; Chen et al. 2000), the CENP-A targeting domain (CATD) within the HFD, essential N- and C-termini (Black et al. 2004; Logsdon et al. 2015), and insight into distinct requirements for establishment versus maintenance of the CENP-A nucleosome at the centromere (Logsdon et al. 2015). In addition to studies at the level of isolated nucleosomes, the centromere nucleosome must also be considered in the context of flanking chromatin. It is the intent of this review to focus on the higher order structures that lead to a bolus of CENP-A on the surface of the chromosome where they can interact and engage microtubules.
1.2 Centromere DNA Directs the Sequence Invariant Position of Cse4
The budding yeast centromere is unique in that the microtubule-DNA interphase is known at base pair resolution. There is a single position invariant Cse4 protein(s) at centromere DNA elements CDEI, II, and III. There are additional Cse4 molecules not confined to a specific sequence (between 3 and 5/chromosome), that can be observed as a cloud surrounding the centromere core (Haase et al. 2013). While evidence for and against the Cse4 cloud exists (Aravamudhan et al. 2013; Haase et al. 2013; Wisniewski et al. 2014), the position variable molecules can be eliminated in specific mutants (pat1, xrn1) as evidenced by the loss of the cloud and confirmed by a reduction in chromatin immunoprecipitation (Haase et al. 2013). We will focus on the sequence invariant molecules as the function of the position variable molecules is not known.
The centromere DNA elements are recognized by a unique DNA-binding complex, Centromere Binding Factor (CBF3, composed of Ndc10 (2), Cep3 (2), Ctf13, Skp1) (Espelin et al. 1997; Russell et al. 1999). Ndc10 is distantly related to the tyrosine DNA recombinases, such as CRE (Cho and Harrison 2012; Perriches and Singleton 2012). Ndc10 may be central to the controversy surrounding the Cse4 nucleosome, as well as the structure of the inner centromere. Ndc10 binds CDEII in the absence of other CBF3 components, as well as A + T rich domains elsewhere in the chromosome (Espelin et al. 2003). Thus Ndc10 is promiscuous in its DNA-binding properties, and is not restricted to kinetochores in vivo (Espelin et al. 2003). Within the kinetochore, Ndc10 localizes to the inner centromere proximal to the spindle axis, where it binds DNA as a dimer and has been proposed to promote DNA bending and/or looping (Pietrasanta et al. 1999). Ndc10 is essential for forming the yeast kinetochore and specifically in recruiting CENP-A (Cse4, Pearson et al. 2003).
The function of the variant centromere histone complex is not well understood. There are differing biological consequences imparted from the mechanics of DNA, whether it is wrapped in a canonical left- or non-canonical right-handed direction around the nucleosome. The determination of left- versus right-handed DNA wraps around the nucleosome can be assessed from indirect measurements of the number of supercoils in small circular plasmids. Circular centromere-containing plasmids contain fewer negative supercoils than acentric plasmids, indicative of a positive (right-handed wrap) supercoil around the centromere core (Bloom et al. 1983, 1984; Furuyama and Henikoff 2009; Diaz-Ingelmo et al. 2015). There are several alternative interpretations of the change in linking number observed in plasmids containing the centromere-specific Cse4 nucleosome, all of which are dependent on CDEIII and/or Ndc10 (Furuyama and Henikoff 2009; Gkikopoulos et al. 2011; Diaz-Ingelmo et al. 2015). One possibility is that that the functional centromere contains a left-handed wrap around Cse4, flanked by two DNAaseI hypersensitive nucleosome-depleted regions (~70 bp each) (Bloom and Carbon 1982). Loss of centromere function in ndc10 mutants leads to loss of the nuclease hypersensitivity (Saunders et al. 1988). If these sites become occupied by a nucleosome with canonical histones, plasmid DNA isolated from these mutants will appear to have additional negative superhelical turns (and hence plasmid from wild-type cells will a net change of +1 in linking number). Alternatively, if Ndc10 introduces a right-handed turn at the base of a DNA loop, this will also influence the net linking number in a positive fashion (Cho and Harrison 2012; Diaz-Ingelmo et al. 2015).
1.3 Primary Loop at the Cse4 Nucleosome
The looping function of Ndc10 and hypersensitivity of flanking sites (70 bp) to DNAase I provides important insight into the three-dimensional structure of the yeast centromere. One can build a unifying model that incorporates Ndc10 looping and dimerization function, the observed change in linking number and extent of micrococcal nuclease digestion (120 bp core (Cole et al. 2011; Krassovsky et al. 2012) and a larger 150–220 bp protected structure (Bloom and Carbon 1982; Funk et al. 1989; Gkikopoulos et al. 2011)). At the base of the CDEIII, Ndc10 binding results in a right-handed DNA wrap of the ends of a ~80 bp duplex that encircles the Cse4 nucleosome (Henikoff et al. 2014). This accounts for the core 80–120 bp seen in extensive MNase digests employed in ChIP seq studies (Cole et al. 2011; Krassovsky et al. 2012). DNA exiting the Ndc10 loop continues to wrap around the Cse4 nucleosome to what will be the surface of the chromosome (middle panel Fig. 1). This leaves about 70 bp of DNA surface exposed as evidenced by extreme DNAase I hypersensitivity. Considering the concentration of nucleosomes adjacent to CDEI-III, it is surprising that stochastic nucleosome dynamics [sliding as well as protein exchange (Verdaasdonk et al. 2012)] does not result in the nuclease hypersensitive sites becoming occupied by histones. These sites are kept nucleosome-free through the action of the ATP-dependent SWI/SNF chromatin remodeling complex, Snf2 (Gkikopoulos et al. 2011). Deletion of Snf2 results in a reduction in the nuclease hypersensitivity flanking CDEI-III, with no change in the flanking nucleosomes or other hypersensitive DNAase I sites.
A DNA basket on the surface of centromeres in metaphase. Left The pericentromere region of all 16 chromosomes in metaphase in budding yeast. The 125 bp CEN region (pink nucleosome at right-most edge of the nucleosome fiber, depicted as yellow DNA wrapped around red histones) lie at the apex of the pericentric chromatin loops (11 nm fiber, yellow strands). The centromere DNA containing loops extend perpendicular to the chromosome axis (Lawrimore et al. 2016). Middle panels Top: end-on view of the Cse4 containing nucleosomes, one from each centromere of the 16 sister chromatids are cylindrically arranged and lie on the surface of the chromosome. Bottom: side view of centromere DNA in metaphase. 80 bp of DNA is in direct contact with the Cse4-containing core (pink), flanking DNA follows a trajectory toward the kinetochore (yellow strands away from the pericentromere). DNA devoid of histones reflect the DNAase I hypersensitive regions (Bloom and Carbon 1982; Bloom et al. 1983) exiting and entering the Cse4 containing nucleosome that protrude from the surface of the chromosome to make a basket. Far right The Cse4 containing nucleosomes are proximal to the pericentric chromatin (yellow DNA strands, bottom), while the DNAase I hypersensitive sites protrude from the chromosome surface toward the kinetochore (top). Sister kinetochores lie ~800 nm away on the opposite surface of the sister strands
In a three-dimensional model of the centromere, these nucleosome-free DNAase I sites of all 16 centromeres are clustered at the chromosome/microtubule interface. The 16 kinetochores are arrayed in a cylinder (~ 250 dia.) around the central spindle resulting in the confinement of DNA loops from all 16 chromosomes to the vicinity of the spindle. Clustering the DNAase I sites to the chromosome surface is a mechanism to distinguish the centromere from the chromosome arms. The active chromatin remodeling mechanism that prevents occlusion of these sites by histones ensures this unique attribute of centromeres. Exposure of such a large patch of naked DNA (~150 bp × 16–2500 bp) is likely to contribute to kinetochore microtubule capture and stabilization mechanisms.
The DNAase I hypersensitive sites are conserved in other fungi (Kluyveromyces lactis) harboring point centromeres (Heus et al. 1993a), even though the centromere sequences have diverged (Heus et al. 1993b). Centromeres from one organism do not confer segregation function in related species. In K. lactis, CDEII is about double the size of that in budding yeast S. cerevisiae. There could be two wraps of DNA around a Cse4 core, or there could be double the number of Cse4 molecules at the site of microtubule attachment. In either case, the model of extreme DNAase I sites on the chromosome surface may be applicable to other fungi as well as larger eukaryotes.
1.4 Pericentromeric Loop—Secondary Folding into Bottle Brush
Sister Cse4-containing nucleosomes are separated by 800 nm in metaphase. Each kinetochore lies at the apex of a DNA loop that extends on average about 400 nm from the sister chromatid axis in metaphase. The centromere loops have been found in 3C maps from yeast (Yeh et al. 2008; Duan et al. 2010), and Hi-C maps of lymphoblastoid cells in mammals (Dai et al. 2016). The size of the loops can be estimated from the region over which the structural maintenance of chromosome proteins (SMC’s cohesin and condensin) are enriched. Loops from each chromosome are in turn looped, adopting loops within loop organization (Lawrimore et al. 2016) (Fig. 2). The emerging principle from the fractal organization of DNA loops is that stiffness is achieved within the pericentric chromatin. A high concentration of loops builds tension within the pericentromere and exerts a pushing force on centromere DNA at the apex of the loops (Cse4 and CDEI, II and III, including the hypersensitive sites). This pushing force is responsible for positioning the centromere to the chromosome surface. The concentration of loops is established through the energy-consuming process of loading and enriching cohesin in the pericentromere at the time of DNA replication. Once the pericentromere chromatin is enriched in SMC proteins, the fluctuations of the strands impose a tension force on the primary loop, where the centromere lies at the apex. This configuration, known as a bottle brush, is a common physical property of polymers that on solid substrates can generate nanonewtons of force, enough to break covalent bonds (Panyukov et al. 2009a, b).
Configuration of pericentric chromatin loops surrounding the spindle axis in the budding yeast. The blue spheres are spindle pole bodies, the green rods are kinetochore microtubules. The interpolar microtubules can be seen as blue rods extending about ¾ the length of the spindle from each pole. The yellow strands are pericentric chromatin. The primary loop (horizontal) is attached to a kinetochore microtubule. Chromosome arms (not shown) would extend north and south, from approximately the middle of the pericentromere. Condensin is at the base of the each of the radial subloops (purple staples). Cohesin are the red rings. The position of cohesin is the most thermodynamically favorable and matches the position observed experimentally with the size and number of loops modeled herein
The size and number of secondary loops has been estimated through the use of a chromosome dynamics simulator (Lawrimore et al. 2016). The bead-spring polymer model finds the thermodynamically favorable state of a string of beads with hinge forces connecting the beads tuned to give the strand the bending rigidity determined for DNA. In this model, cohesin is concentrated in the pericentromere and adopts a uniform and homogeneous distribution that surrounds the central spindle, but is radially displaced (Yeh et al. 2008; Stephens et al. 2011). There is no molecular mechanism that accounts for the position or appearance of a cohesin barrel. We have found that the size and number of DNA loops dictate the experimentally observed position. Loops that are approx. 10 kb give rise to a cohesin barrel that matches the dimension and homogeneity found in vivo (Lawrimore et al. 2016). The size of these loops is comparable to the 5 kb chromosome interaction domains found in yeast from Micro-C (Hsieh et al. 2015).
An alternative view of cohesin function at the centromere is provided in Hu et al. (2015). To reconcile the position of cohesin relative to the spindle axis with the canonical function of juxtaposing sister chromatids, Hu et al. propose that pericentric cohesin lies on average 5 kb from the centromere core and is restricted to sister chromatid interactions as observed in the arms. Stephens et al. (2011) found that a lacO array 6.8 kb from the centromere was separated greater than 65% of the time in wild-type cells, inconsistent with such a proposal. Furthermore, the radial distance of cohesin from the spindle, as well as the homogeneity of the barrel demands a physical explanation. Rather than holding sister chromatids in the pericentromere, intra-chromatid linkages as proposed in Lawrimore et al. (2016) account for the position and dynamics of pericentric cohesin. Behavior of the centromere linked lacO array as it extends and retracts thermodynamically, as well as in response to MT pulling and pushing forces also favors intra-chromosomal loops diagrammed in the model shown in Fig. 2. The intra-chromatid linkages contribute to the pushing mechanism predicted from the bottle brush, providing a novel view of centromere function in promoting sister separation and kinetochore tension at the metaphase plate (Lawrimore et al. 2015, 2016). Whether non-sister chromosome interactions are mediated by cohesin within the pericentromere remain controversial. Biophysical studies of cohesin diffusion on DNA indicates that the physiological pore size is much smaller than observed in electron micrographs of spread molecules (Stigler et al. 2016). Based upon the fluctuations of radially displaced LacO spots and correlated movement between pericentromeres of non-sister chromatids, models of cohesin rings that encompass a single chromatid and interact with cohesin rings from other chromosomes account for experimental observations (Stephens et al. 2011, 2013a; b). Interestingly, in experimental attempts to reduce the concentration of cohesin in the cell, the only region of the chromosome where cohesin was refractory to reduction was in the centromere (Heidinger-Pauli et al. 2010). Thus there are mechanisms to ensure the concentration remains in centromere, indicating this may be the critical site for the fidelity of chromosome segregation.
1.5 Loops on Mammalian DNA and Role of Satellite Repeats in Promoting DNA Looping
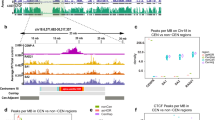
Recently, centromeric chromatin was reconstituted in cell-free extracts derived from Xenopus laevis eggs (Aze et al. 2016). For these experiments artificial chromosomes (BACs) containing large inserts (130–160 kb) of centromeric alpha satellite DNA sequences of different human centromeres were used. BACs were chosen for their large size, which allowed efficient formation of nuclear structures in egg extract (Aze et al. 2016). Electron microscopy (EM) based structural analysis of centromeric DNA isolated from egg extract highlighted the presence of numerous single-stranded DNA bubbles. These structures were due to intrinsic resistance of centromeric chromatin to psoralen-mediated cross-linking, a procedure required to prevent melting and branch migration of DNA preparations for EM analysis. Poor cross-linking was due to the presence of positively supercoiled DNA, which is known to prevent psoralen-mediated cross-linking (Bermudez et al. 2010). Formation of positively supercoiled DNA required Topoisomerase I activity. Chromatin composition analysis also revealed the enrichment of condensin, which can promote the formation of positively supercoiled DNA in closed plasmids in the presence of Topoisomerase I (Hirano 2012). These findings suggest that positive supercoils observed in centromeric DNA assembled in egg extract is an active phenomenon linked to the presence of Topoisomerase I and condensin.
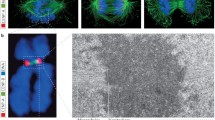
When partially digested chromatin still retaining condensin was analyzed instead of naked DNA EM revealed the presence of long double stranded DNA loops embedded in a protein matrix (Fig. 3). The average size of the loops was around 1–1.5 kb but some of the loops reached 2.5–3 kb in size. Although the individual components of the protein matrix could not be resolved, electron dense particles corresponding to residual protein material left after partial proteolysis were spotted at the base of the loops, indicating a possible role in their formation or stabilization (Fig. 3). Consistent with this hypothesis removal of residual proteins by complete digestion dissolved the loop structure (Aze et al. 2016).
Electron micrograph showing partially digested chromatin isolated from Xenopus laevis egg extract incubated with human alpha satellite DNA. Loops of double stranded DNA filaments running parallel to each other embedded in a protein matrix can be appreciated. Electron dense particles can be noticed at the base of some of the loops. Bar corresponds to 500 base pairs
Formation of loops observed on centromeric chromatin isolated from interphase egg extracts required DNA replication and active Topoisomerase I, although Topoisomerase II role could not be excluded. The presence of positively supercoiled DNA associated to the loops suggested that these structures formed behind replication forks as their presence ahead of them would heavily disturb fork progression (Branzei and Foiani 2010). Condensins play a major role in mitosis-dependent chromosome condensation in eukaryotic cells. A role for condensin during DNA replication has been documented in bacteria, where their activity is required for DNA loop formation. In this case condensins appear to resolve replicated replication origins by promoting the juxtaposition of DNA, drawing sister replication origins away from each other (Wang et al. 2015). To this end condensins encircle double-stranded DNA at their loading site and slide along it, tethering the two arms of double-stranded DNA together. If applied to mammalian cells this process could disentangle and individualize replicated sections of centromeric DNA promoting at same time DNA condensation, which could start at centromeres in interphase and then spread towards flanking regions of the chromosome during the mitotic phase. This process could help to disentangle highly repeated centromeric DNA sequences after replication and prevent their recombination.
The loops uncovered in this system could be related to the organization of centromeric chromatin in mitosis. The EM pictures evoke the bottlebrush structure proposed for the loop arrangement in yeast centromere. The smaller size of the loops identified in interphase by EM compared to the ones predicted by biophysical studies might reflect an early stage of centromere reorganization. The loops assembled in interphase could indeed be the basic unit of larger structures, which might grow further in mitosis. The requirement of Topoisomerase I and possibly Topoisomerase II activities for their formation and/or for their stabilization indicates the occurrence of rotational processes along DNA axes during loop formation and the possible presence of loop entanglements that stabilize these structures. Overall these features could be compatible with the formation of the centromeric spring.
CENP-A is a key factor able to trigger assembly of kinethocore proteins in vitro (Weir et al. 2016) and in Xenopus egg cytoplasm (Guse et al. 2011). However, the links between centromeric DNA and CENP-A as far as chromatin structure is concerned are largely unclear. Interestingly, CENP-A could be selectively loaded onto naked centromeric DNA containing alpha satellite in Xenopus egg extract (Aze et al. 2016). The ability of satellite DNA to induce loading of centromeric proteins was consistent with work performed on human artificial chromosomes (HACs), which are made with satellite DNA and are able attract centromere and kinetochore proteins when introduced into cells (Nakano et al. 2008; Bergmann et al. 2011; Kouprina et al. 2013). Replicative features such as invariant inter-origin distance between centromeric and non-centromeric DNA in Xenopus were also consistent with data obtained from studies on HAC replication in intact cells (Erliandri et al. 2014). These two systems could help to better define important aspects of centromeric chromatin assembly in the future.
It is generally assumed that CENP-A is loaded onto centromere chromatin between the end of mitosis and subsequent G1 phase of the cell cycle. An epigenetic mechanism is involved in the deposition of new CENP-A onto chromatin regions with preexisting CENP-A. However, in vitro results suggested that de novo CENP-A loading can also take place in interphase and on naked centromeric DNA made of alpha satellite DNA. The reason why satellite DNA could trigger such a complex chain reaction of events is not known. Similar to the ability of CENP-A to induce kinetochore assembly (Guse et al. 2011) CENP-A could be an important trigger for centromeric loop formation. Centromeric chromatin reconstitution was achieved with DNA sequences containing the CENP-B box, a 17 bp element, which functions as a binding site for CENP-B protein (Fachinetti et al. 2015). CENP-B protein has been recently shown to be required for the stable loading of CENP-A on alpha satellite DNA (Fachinetti et al. 2015). Although satellite DNA has not been described yet in X. laevis, repetitive sequences containing a conserved CENP-B box able to bind X. laevis CENP-A have been isolated (Edwards and Murray 2005). Therefore, it is possible that the presence of CENP-B box facilitates recruitment of CENP-A onto human centromeric DNA in egg extract possibly through CENP-B. Consistent with this hypothesis proteomic analysis of centromeric chromatin assembled in egg extract revealed the selective loading of a number of centromeric proteins (Aze et al. 2016) among which a protein with weak homology to CENP-B. However, further studies are needed to confirm that this protein is the true CENP-B ortholog.
Other proteins found enriched in centromeric chromatin in many species such as condensins might be responsible for the centromere loop formation on satellite DNA. How condensins are specifically loaded on satellite DNA is not known. There could be a structural code in the repetitive DNA able to attract condensins and other centromeric proteins promoting the formation of complex structures. The existence of a structural code conserved from yeast to mammals has been proposed and it is related to an internal dyadic symmetry of individual tandem repeats and yeast centromeric sequences, which could give rise to mismatched hairpins (Koch 2000; Jonstrup et al. 2008). Some of these structures have been demonstrated in vitro but their significance in vivo has never been explored. Condensins might have affinity for hairpins and could recognize these structures. With the advent of techniques such as Crisper/Cas9 satellite DNA could be modified to test the in vivo relevance of this structural code. Condensins have also been implicated in the assembly of CENP-A chromatin in Xenopus and human cells (Samoshkin et al. 2009; Bernad et al. 2011) but their precise role in the process is still unclear. Recent work has shown that condensin subunit Cut3 in fission yeast mediates the organization of pericentromeric tandem repeats into a specific higher order structure, which helps to restrict CENP-A loading to centromeres (He et al. 2016). Interestingly, formation of neocentromeres triggered by CENP-A binding to extra-centromeric sites which do not contain centromeric repeats frequently occurs in genomic areas enriched for duplicated sequences (Marshall et al. 2008). Therefore, the tandem repeats or closely duplicated sequences themselves might act as signal to recruit condensins and CENP-A, giving rise to series of complex events required to build a mature centromere.
2 Mechanisms of Loop Formation
Several mechanisms of chromosome and centromere loop formation have been proposed in the literature. One is that natural fluctuations of the polymer chains lead to encounters that are stabilized by SMC proteins cohesin and condensin (Vasquez et al. 2016). Using bead-springs to model chromosome arms as polymer chains we have shown that domains of high interaction (intra-chromosome loops) arise naturally from polymer thermodynamics without the need for other mechanical or chemical potentials interacting via entropic potentials, such as the chromatin spring and excluded volume forces. These regions are not static and vary from cell to cell. In the centromere, such loops could be stabilized through the action of proteins such as topoisomerases, and SMCs that increase the lifetime of a given loop. A second model that has been proposed is the activity of loop extrusion enzymes (Alipour and Marko 2012). In this model, SMC proteins act as a machine that threads DNA into a loop as the enzymes translocate along the helix. There is considerable interest in this model from recent studies using 3C and Hi-C techniques (Fudenberg et al. 2016; Goloborodko et al. 2016a, b).
Chromatin remodeling proteins have also been shown to be loop extruding motors (De Cian et al. 2012; Sun et al. 2013), and may impart this function at the centromere. The centromere is enriched in several members of various chromatin remodeling protein families. Chromatin remodelers are able to mobilize nucleosomes, function to translocate linker DNA over the nucleosome, maintain nucleosome-free regions, such as around promoters, and loop DNA to tune topological domains. The yeast Isw1 complex (Imitation switch) is of particular interest as it has been shown in vitro to function as an inchworm that will form and propagate intra-stand DNA looping (Fig. 4) (De Cian et al. 2012). Isw1 is a member of the SWI/SNF ATPase complex and interacts with the centromere DNA-binding factor CBF1 (Moreau et al. 2003). Cbf1 is required to recruit Isw1 to sites of transcription (Moreau et al. 2003), and may recruit Isw1 to centromere as well. Other members of this family of ATPases, including Snf2 (Gkikopoulos et al. 2011) and PICH (Plk1-interacting checkpoint helicase) (Baumann et al. 2007) have been shown to have a direct role in centromere function. The actin containing chromatin remodeling ATPases have been implicated in chromatin structure of the pericentric domain (Chambers et al. 2012). These enzymes are not essential for centromere function, but mutations lead to decreased segregation fidelity. Their role in sculpting chromatin loops is consistent with the experimental evidence and the formation of loops may be a unifying function that enhances segregation fidelity. Loops also exhibit features observed in RNA processing and lariat intermediates. The base of secondary loops in the budding yeast centromere is proximal to the spindle, where condensin and dyskerin (CBF5) are concentrated (Snider et al. 2014). Dyskerin is a pseudouridine synthetase associated with the snoRNP complex (Zebarjadian et al. 1999). In other guises, these RNPs are able to make lariats (loops) in processing introns from primary transcripts. While the function on RNA and ssDNA is targeted to single-strand nucleic acid, perhaps the pericentromeric dyskerin-containing complexes have evolved a DNA based looping mechanism.
Isw1a, a member of the SWI/SNF chromatin remodeling family functions as a loop extruding enzyme in vitro, adapted from (De Cian et al. 2012). Isw1 interacts with the yeast Cbf1 factor where it functions to maintain nucleosome-free regions at promoters. This and other ATPases at centromere could collectively promote pericentromeric looping
The Ndc10 protein of the CBF3 complex may also promote looping in the pericentromere. Ndc10 binds CDEIII where it recruits Cse4 (Pearson et al. 2003), but Ndc10 also localizes to pericentric chromatin along the spindle axis in live cells (Bouck and Bloom 2005). Based upon its homology to tyrosine site-specific recombinases, Ndc10 may be able to join distal sites to which it is bound (Jayaram et al. 2015). If chromatin-bound Ndc10 oligomerizes through its dimerization domain (Cho and Harrison 2012; Perriches and Singleton 2012), DNA loops will emerge.
3 Centromere Inactivation and de Novo Activation
If cohesin is not holding sister centromeres together, considering they are separated by 400–800 nm in metaphase, what other function might these molecules have. Using transcription as a mechanism to functionally inactivate the centromere we have found that cohesin contributes to the conformation of pericentric chromatin that is favorable for kinetochore assembly (Tsabar et al. 2016). It is unlikely that cohesin recruits kinetochore proteins as there are no direct interactions, and in vivo the pericentric cohesion barrel is well separated from the kinetochore/microtubule attachment complex (Yeh et al. 2008). It has been suggested that proteins such as Sgo1 contribute to the bias that favors sister centromeres to face opposite poles (Fernius and Hardwick 2007; Indjeian and Murray 2007). The barrel of pericentric cohesin could be the physical manifestation of such a mechanism. By assembling cohesin between sister centromeres, the centromeres will be inherently pushed apart, thereby favoring them to lie on the surface of the chromosome.
Chromatin remodeling complexes may also be critical in centromere architecture. Durand-Dubief (Durand-Dubief et al. 2012) found that like cohesin, the Swi/Snf-like chromatin remodeling complex, Fun30 contributes significantly to centromere function when centromeres are transcriptionally inactivated. In fun30 mutants, cells containing a chromosome with a transcriptionally inactivated centromere as the only centromere are completely inviable. Fun30 is therefore required to build a proper architecture that can compensate for centromere inactivation. Fun30 shares a phenotype with another centromere component Chl4. Chl4 is a non-essential kinetochore protein, associated with the inner kinetochore (Iml3 of the Ctf19 complex that interacts with Ctf19, Ctf3 and Mif2). Both Chl4 and Fun30, as well as Ctf19 are required for de novo centromere formation (Mythreye and Bloom 2003; Laha et al. 2011; Durand-Dubief et al. 2012).
4 Centromere DNA Replication and Response to Stress
Repetitive DNA sequences are generally unstable and prone to recombination (Branzei and Foiani 2010). The formation of positively supercoiled DNA loops might help to compact centromeric chromatin preventing inappropriate recombination. However, this structure could act as barrier to DNA replication machinery affecting the progression of replication fork. In this case the compact chromatin structure could act as double edge sword, protecting repetitive DNA from recombination on one hand and imposing replication roadblocks to replication fork progression on the other hand. In addition centromeric repetitive sequences can form secondary DNA structures such as hairpins with misaligned and mismatched bases during DNA replication, when the double-stranded DNA is unwound. Such hairpins have been described in vitro for individual satellite DNA repeats, which harbor internal dyadic symmetry conserved from yeast to primates, promoting in vitro self-annealing (Koch 2000; Jonstrup et al. 2008). Therefore chromatin and DNA conformation could be the source of replication stress ultimately provoking DNA breakage in the centromere. Inappropriate control of such stress together with the abnormal forces exerted in mitosis on centromere regions could lead to centromere DNA breakage. Consistent with this idea is that centromeres appear to be hotspots for chromosomal breakage and rearrangements in mammalian and yeast cells (Simi et al. 1998; McFarlane and Humphrey 2010). Induction of replication stress has also been linked to the formation of acentric broken chromosomes in human cells, in which the centromeric protein staining is completely lost (Burrell et al. 2013).
Surprisingly, reconstitution of centromeric chromatin revealed that centromeric DNA was efficiently replicated and replication efficiency was comparable to non-repetitive sequences (Aze et al. 2016). Therefore repetitive DNA did not cause apparent problems to the replication apparatus. These findings indicated the possible presence of specialized factors bound to centromeric chromatin that could help resolve replication roadblocks. Mass spectrometry analysis of the proteome associated with replicating centromeric DNA revealed the enrichment of several DNA repair and DNA structural proteins among which MSH2-6, the MRE11-RAD50 complex, HMGB1-3, XRCC1, XRCC5/DNA-PK, PARP1, ERCC6L/PICH helicase and MUS81 endonuclease (Aze et al. 2016). Some of these proteins were independently found on mouse centromeres (Saksouk et al. 2014). The accumulation of many of these repair factors was abolished by Geminin, an inhibitor of DNA replication fork assembly indicating that centromeric repair factors were loaded in a replication-dependent fashion. In contrast, other common replication players were underrepresented, such as the single-stranded DNA binding (ssDNA) complex RPA and ATR activator TopBP1. Accumulation of these proteins on DNA following induction of stalled replication forks was also diminished. Taken together these results indicated that centromeric DNA replication likely requires accessory DNA repair factors for accurate and efficient replication. This hypothesis was directly tested for the MSH2-6 complex, whose absence compromised centromeric replication. Furthermore, induction of replication stress by inhibition of DNA polymerases induced further recruitment of the MSH2-6 complex. In the same conditions accumulation of RPA, which usually follows formation of ssDNA induced by fork uncoupling triggered by polymerase arrest was not observed. These findings suggested that ssDNA arising at centromeric stalled forks does not get exposed to RPA and likely forms secondary structures containing mismatched bases, which attract MSH2-6.
Reduced RPA accumulation was also responsible for decreased levels of TopBP1 protein and suppression of ATR-dependent CHK1 phosphorylation following induction of stalled replication forks (Aze et al. 2016). Suppression of the ATR checkpoint was required for the efficient replication of repetitive centromeric DNA as forced activation of the checkpoint by interference with topoisomerase activity was indeed sufficient to selectively inhibit centromeric DNA replication. These results revealed two unexpected features of centromeric chromatin. One is that checkpoint suppression is an active phenomenon due to the topological arrangement of the centromeric chromatin in positively supercoiled loops, whose disruption restores the sensitivity to checkpoint activation. The second is that local suppression of ATR signaling facilitates replication of centromeric repetitive DNA, which would otherwise trigger continuous activation of ATR inhibiting replication origin firing.
The physiological roles of checkpoint suppression and sensitivity of centromeric DNA replication to its activation are unclear at the moment. One possibility is that ATR-dependent inhibition of centromeric DNA replication could play a role in preventing unscheduled chromosome segregation in response to stalled forks elsewhere on the chromosome. This could confer a selective advantage to chromosomes bearing DNA repeats at centromeres. It is worth noticing that similar to centromeric DNA other repetitive DNA regions such as the telomeres suppress checkpoint activation, form large loops and are organized in positively supercoiled domains (Benarroch-Popivker et al. 2016) indicating that there might be similarities between telomeres and centromeres in the way DNA is organized and respond to stress.
5 Centromere Breakage and Repair
Replication stress, topological constraints and pulling forces exerted on centromeres during chromosomes segregation might all cause centromere double strand break (DSBs). Recent evidence obtained in cancer cell lines documented increased incidence of DSBs at centromeres following replication stress induced by aphidicolin (Crosetto et al. 2013). The occurrence of DSBs at centromeres following replication stress could explain the centromeric enrichment of DNA repair such as DNA-PK, PARP1, MRE11 and MUS81 (Aze et al. 2016). These proteins are all involved at some level in DSB processing and repair, suggesting the occurrence of ongoing DSB repair at centromeres. Alternatively, DSB repair proteins might accumulate following induction of reversed forks (RFs), which form after the annealing of nascent DNA strands at stalled replication intermediates and mimic the occurrence of DSBs being double stranded (Errico et al. 2014). RFs might frequently occur on centromeric DNA due to their repetitive nature and might require nuclease such as MRE11 or MUS81 to be removed. DSBs might also occur during incomplete DNA decatenation, which requires Topoisomerase II as it is prevented by Topoisomerase II inhibitors (Liu et al. 2014). Interference with DNA decatenation induces formation at centromeres of ultrafine bridges (UFBs), mitotic DNA structures visible in the anaphase of mitosis due to the pulling of incompletely replicated or processed DNA intermediates during chromosome segregation (Liu et al. 2014), which could generate centromeric DSBs.
DSBs at centromeres could be rapidly rejoined through different DSB repair pathways. Centromeric DNA repair events might be facilitated by the presence of tandem repeats. Localization of homologous recombination (HR) and non-homologous end joining (NHEJ) repair proteins on DSBs centromeres has recently been confirmed in mammalian cells (Tsouroula et al. 2016). Interestingly, differently from other regions of the chromosome centromeric DSBs induced by Crispr/Cas9 were shown to bind both NHEJ and HR in all phases of the cell cycle, suggesting that both pathways are active at all times. Among the HR pathways single-strand annealing (SSA) and synthesis-dependent strand annealing (SDSA), might be particularly facilitated by repetitive sequences (Paques et al. 1998; Paques and Haber 1999). In SSA one end of the DSB with exposed ssDNA anneals to the other end and this event is facilitated by the presence of duplicated sequences. During SDSA the annealing phase is followed by replication-mediated extension of the invading strand, which uses the homologous sequence as template. Multiple direct repeats, as found in human centromeres, might also contribute to generate large ring structures via recombination between distant homologous direct repeats. Pairing between homologous sequences might stabilize such large rings. Abnormal metabolism and resolution of recombination products between these direct repeats would instead release the ring from the chromosome, forming extra-chromosomal circular molecules. Many extra-chromosomal circles of centromeric DNA have been observed to accumulate after replication stress in several species (Cohen et al. 2003; Cohen and Segal 2009).
Although studies of satellite-less neocentromeres have overshadowed the possible importance of repetitive DNA in established centromeres these observations indicate that the centromere might require repetitive DNA to ensure its optimal function, particularly under stress. It is likely that DNA repeats are actively maintained notwithstanding their problematic replication for a number of advantageous features such as promoting efficient repair of centromeres in case of breakage and functioning as checkpoint sensitive zone of the replication checkpoint. Unregulated recombination between repeated sequences could lead to the complete or partial loss of centromeric DNA following replication stress. Furthermore, abnormal resolution of SSA and SDSA repair events could promote contraction and expansion cycles of the intervening DNA repeats, respectively (Paques et al. 1998; Paques and Haber 1999). Loss of repetitive DNA could result in the shortening of centromeres in older cells, which have undergone several cell cycles. Centromere deterioration has been indeed observed in aging women although it is not clear if this depends on DNA replication (Nakagome et al. 1984).
6 Repetitive DNA and Loops Generate Centromere Force
The bottlebrush centromere provides a mechanistic understanding for the role of repeat DNA in the centromere and addresses several outstanding problems (Lawrimore et al. 2016). First, the organization of repeats into DNA loops via another SMC family member cohesin, has been well documented in the nucleolus (Harris et al. 2014). Second, the bottlebrush provides a physical basis for how a floppy DNA chain can be converted into a stiff (relative to an entropic chain) spring. A fluctuating chain in a thermal bath will find the most entropically favored state, that of a random coil, as a random coil chain is the most disordered. The addition of side chains relative to the primary chain (or primary axis), limits the ability of the primary chain to adopt a random coil, through limiting the number of states the primary chain can adopt. Additional side chains further restrict the motion of the primary axis until a point where they generate tension along the axis. In this fashion, enthalpic energy put into making chains, results in entropic forces of the brush where side chain fluctuation amplifies tension along the primary axis. Thus chromatin loops significantly change the state of the centromere from a floppy chromatin polymer into a stiff chromatin network.
The chromatin loops provide a mechanism for buffering changes in tension resulting from microtubule dynamics. Kinetochore microtubules are in a constant state of flux, and rare persistent growth and/or shortening events could result in large local changes in tension at individual kinetochores. DNA loops compensate for large changes in kinetochore microtubule length through their ability to convert between looped and stretched states (Stephens et al. 2013a). The looped state is radially displaced from the spindle axis. Increased tension at the kinetochore exerts a pulling force at the base of the loop, switching the loop to the stretched state and increasing the length of DNA along the spindle axis. The increase in axial DNA compensates for kinetochore microtubule shortening. Likewise persistent microtubule growth events will bias the DNA into the looped state, decreasing the length of axial DNA. Thus pericentric loops buffer the system such that tension can be maintained on a landscape of dynamic kinetochore microtubules.
The bottlebrush also helps us understand how the centromere retains morphology (and cohesin its uniform barrel structure) in a dynamic system where individual chromatin chains are fluctuating and microtubules are constantly prodding and probing the kinetochore resulting in its deformation surface of the centromere (Magidson et al. 2016). Chromosomes, like slip link gels are known to retain their elastic and tensile moduli properties over several orders of magnitude (Okumura and Ito 2001; Granick and Rubinstein 2004; Bloom and Joglekar 2010). It has been known for almost half a century that chromosomes expand and contract upon removal and return of mono- and divalent ions (Lezzi and Gilbert 1970), and depending on the treatment retain their original structure (e.g. such as their macroscopic banding pattern). The concentration of ring complexes such as cohesin and condensin in the pericentromere may be indicative of the slip-link property of centromeres. These complexes are able to compact chromatin, and in the case of cohesin, is able to compact DNA against a force of 0.45 pN (Sun et al. 2013).
7 Repeat Stability and Chromosome Loss
Loss of centromeric DNA repeats due to incorrect duplication or repair might ultimately cause impairment of microtubule attachment to chromosome. Replication stress has indeed been shown to be a primary cause of chromosome number abnormalities and instability (CIN) in cancer cells (Burrell et al. 2013). Complete loss of centromeric DNA following replication stress could indeed lead to segregation errors due to lack of functional kinetochores in case of acentric chromosomes, which are frequently encountered in CIN+ cells. This would suggest a pre-mitotic origin of some chromosome segregation errors (Burrell et al. 2013). This view was recently challenged by observations that the occurrence of acentric chromosomes is limited compared to whole lagging chromosomes caused by primary mitotic segregation errors in CIN+ cells (Bakhoum et al. 2014). However, complete loss of centromeric DNA causing formation of acentric chromosomes might not be the only pre-mitotic cause of chromosome segregation errors. A more limited loss of centromeric DNA similar to centromere degeneration observed in older cells, which is more difficult to detect by fluorescence-based techniques than complete loss, could lead to the formation of centromeres with suboptimal function. Some of these suboptimal centromeres might be responsible for microtubule-chromosome attachment problems leading to the formation of lagging chromosomes.
Among chromosome-microtubule issues there is merotelic attachment in which spindle fibers from both poles attach to a single kinetochore. This condition is particularly dangerous for chromosome stability as it often causes the occurrence of lagging chromosomes not sensed by the mitotic checkpoint apparatus. Lagging chromosomes can be inherited as extra-chromosomes in one of the daughter cells (Santaguida and Amon 2015). This extra chromosome could end up in micronuclei, in which replication and repair is inefficient due to limited availability of nuclear factors predisposing to catastrophic events such as chromosome pulverization (Crasta et al. 2012). In this case a small change in the centromeric DNA structure leading to weakening of chromosome attachment to spindle might lead to catastrophic consequences.
Merotelic attachment of mitotic origin uncorrected by the classic mitotic checkpoints could also generate forces sufficiently strong to physically shear the centromere leading to chromosome breakage (Guerrero et al. 2010). Loss of centromeric DNA and decreased loop formation could make the centromeric chromatin less resistant to such forces and more prone to break. Incomplete maturation and decreased levels of entanglements due to centromeric DNA abnormalities might also weaken the centromeric spring. Therefore, although it is not clear whether forces generated during mitosis are strong enough to break the normal chromatin or DNA structure present at centromeres (Ganem and Pellman 2012), centromere weakening might lower the tolerance to this type of mechanical stress.
Centromere DNA breakage might be an early event in cellular transformation. Inappropriate repair of centromere breakage in cells combined to inefficient DNA repair might predispose to breakage-fusion-bridge (BFB) cycles generating further chromosome instability (Martinez and van Wely 2011; Forsburg 2013). The BFB cycle has initially been described for telomeric end regions of chromosomes. However, although the telomere fusion hypothesis is believed to be the major engine of BFB it does not explain frequent loss and gains of whole chromosome arms in tumors (Beroukhim et al. 2010; Martinez and van Wely 2011; Forsburg 2013). Furthermore, cytogenetic and microarray analysis of breakage-fusion sites revealed frequent occurrence of tandem fusions of chromosome arms with parallel orientation containing interstitial centromere and telomere sequences (Martinez and van Wely 2011). Such events likely derive from a whole chromosome arm produced by centromere breakage that fused to an unprotected telomere of an intact chromosome, leading to the formation of a dicentric chromosome with two centromere-kinetochores structures. Dicentric chromosomes could undergo further breakage at random places when pulled by the spindle fibers bound to old and new centromeres. These events account for more than 50% of chromosome translocation in some tumors, whereas telomere–telomere fusions with antiparallel orientation produced by telomere erosion constitute only 1% of total translocations when analyzed with classical cytogenetic techniques (Martinez and van Wely 2011). These observations strongly suggest that centromere breakage is a major event in the BFB cycle.
To better understand these processes at molecular and structural level the genomic structure of centromeric DNA should be better characterized and annotated. The identification of unique non-repetitive sequences interspersed in centromeric DNA might be useful to design probes to monitor stability of centromeric DNA during cell cycle in unchallenged and stressful conditions. Such tools could be used to test the occurrence of contraction and expansion cycles at repetitive centromeric DNA during unperturbed and challenged DNA replication. These studies could help understanding why these repetitive DNA sequences are retained despite their problematic maintenance. Understanding how these processes occur at molecular level will therefore be essential to clarify the origin of genome instability predisposing to cancer.
References
Alipour E, Marko JF (2012) Self-organization of domain structures by DNA-loop-extruding enzymes. Nucleic Acids Res 40:11202–11212
Aravamudhan P, Felzer–Kim I, Joglekar AP (2013) The budding yeast point centromere associates with two Cse4 molecules during mitosis. Curr Biol 23:770–774
Aze A, Sannino V, Soffientini P, Bachi A, Costanzo V (2016) Centromeric DNA replication reconstitution reveals DNA loops and ATR checkpoint suppression. Nat Cell Biol 18:684–691
Bakhoum SF, Silkworth WT, Nardi IK, Nicholson JM, Compton DA, Cimini D (2014) The mitotic origin of chromosomal instability. Curr Biol 24:R148–R149
Baumann C, Korner R, Hofmann K, Nigg EA (2007) PICH, a centromere-associated SNF2 family ATPase, is regulated by Plk1 and required for the spindle checkpoint. Cell 128:101–114
Benarroch-Popivker D, Pisano S, Mendez-Bermudez A, Lototska L, Kaur P, Bauwens S, Djerbi N, Latrick CM, Fraisier V, Pei B, Gay A, Jaune E, Foucher K, Cherfils-Vicini J, Aeby E, Miron S, Londono-Vallejo A, Ye J, Le Du MH, Wang H, Gilson E, Giraud-Panis MJ (2016) TRF2-Mediated control of telomere dna topology as a mechanism for chromosome-end protection. Mol Cell 61:274–286
Bergmann JH, Rodriguez MG, Martins NM, Kimura H, Kelly DA, Masumoto H, Larionov V, Jansen LE, Earnshaw WC (2011) Epigenetic engineering shows H3K4me2 is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore. EMBO J 30:328–340
Bermudez I, Garcia-Martinez J, Perez-Ortin JE, Roca J (2010) A method for genome-wide analysis of DNA helical tension by means of psoralen-DNA photobinding. Nucleic Acids Res 38:e182
Bernad R, Sanchez P, Rivera T, Rodriguez-Corsino M, Boyarchuk E, Vassias I, Ray-Gallet D, Arnaoutov A, Dasso M, Almouzni G, Losada A (2011) Xenopus HJURP and condensin II are required for CENP-A assembly. J Cell Biol 192:569–582
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J, Barretina J, Boehm JS, Dobson J, Urashima M, Mc Henry KT, Pinchback RM, Ligon AH, Cho YJ, Haery L, Greulich H, Reich M, Winckler W, Lawrence MS, Weir BA, Tanaka KE, Chiang DY, Bass AJ, Loo A, Hoffman C, Prensner J, Liefeld T, Gao Q, Yecies D, Signoretti S, Maher E, Kaye FJ, Sasaki H, Tepper JE, Fletcher JA, Tabernero J, Baselga J, Tsao MS, Demichelis F, Rubin MA, Janne PA, Daly MJ, Nucera C, Levine RL, Ebert BL, Gabriel S, Rustgi AK, Antonescu CR, Ladanyi M, Letai A, Garraway LA, Loda M, Beer DG, True LD, Okamoto A, Pomeroy SL, Singer S, Golub TR, Lander ES, Getz G, Sellers WR, Meyerson M (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463:899–905
Black BE, Foltz DR, Chakravarthy S, Luger K, Woods VL Jr, Cleveland DW (2004) Structural determinants for generating centromeric chromatin. Nature 430:578–582
Bloom K, Joglekar A (2010) Towards building a chromosome segregation machine. Nature 463:446–456
Bloom KS, Carbon J (1982) Yeast centromere DNA is in a unique and highly ordered structure in chromosomes and small circular minichromosomes. Cell 29:305–317
Bloom KS, Fitzgerald-Hayes M, Carbon J (1983) Structural analysis and sequence organization of yeast centromeres. Cold Spring Harb Symp Quant Biol 47(Pt 2):1175–1185
Bloom K, Amaya E, Yeh E (1984) Centromeric DNA structure in yeast chromatin. In: Borisy GG, Cleveland DW, Murphy DB (Eds.), Molecular biology of the cytoskeleton. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 175–184
Bouck D, Bloom K (2005) The role of centromere-binding factor 3 (CBF3) in spindle stability, cytokinesis, and kinetochore attachment. Biochem Cell Biol 83:696–702
Branzei D, Foiani M (2010) Maintaining genome stability at the replication fork. Nat Rev Mol Cell Biol 11:208–219
Burrell RA, McClelland SE, Endesfelder D, Groth P, Weller MC, Shaikh N, Domingo E, Kanu N, Dewhurst SM, Gronroos E, Chew SK, Rowan AJ, Schenk A, Sheffer M, Howell M, Kschischo M, Behrens A, Helleday T, Bartek J, Tomlinson IP, Swanton C (2013) Replication stress links structural and numerical cancer chromosomal instability. Nature 494:492–496
Camahort R, Shivaraju M, Mattingly M, Li B, Nakanishi S, Zhu D, Shilatifard A, Workman JL, Gerton JL (2009) Cse4 is part of an octameric nucleosome in budding yeast. Mol Cell 35:794–805
Chambers AL, Ormerod G, Durley SC, Sing TL, Brown GW, Kent NA, Downs JA (2012) The INO80 chromatin remodeling complex prevents polyploidy and maintains normal chromatin structure at centromeres. Genes Dev 26:2590–2603
Chen Y, Baker RE, Keith KC, Harris K, Stoler S, Fitzgerald-Hayes M (2000) The N terminus of the centromere H3-like protein Cse4p performs an essential function distinct from that of the histone fold domain. Mol Cell Biol 20:7037–7048
Cho US, Harrison SC (2012) Ndc10 is a platform for inner kinetochore assembly in budding yeast. Nat Struct Mol Biol 19:48–55
Cohen S, Segal D (2009) Extrachromosomal circular DNA in eukaryotes: possible involvement in the plasticity of tandem repeats. Cytogenet Genome Res 124:327–338
Cohen S, Yacobi K, Segal D (2003) Extrachromosomal circular DNA of tandemly repeated genomic sequences in Drosophila. Genome Res 13:1133–1145
Cole HA, Howard BH, Clark DJ (2011) The centromeric nucleosome of budding yeast is perfectly positioned and covers the entire centromere. Proc Natl Acad Sci U S A 108:12687–12692
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, Nezi L, Protopopov A, Chowdhury D, Pellman D (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482:53–58
Crosetto N, Mitra A, Silva MJ, Bienko M, Dojer N, Wang Q, Karaca E, Chiarle R, Skrzypczak M, Ginalski K, Pasero P, Rowicka M, Dikic I (2013) Nucleotide-resolution DNA double-strand break mapping by next-generation sequencing. Nat Methods 10:361–365
Dai C, Li W, Tjong H, Hao S, Zhou Y, Li Q, Chen L, Zhu B, Alber F, Zhou XJ (2016) Mining 3D genome structure populations identifies major factors governing the stability of regulatory communities. Nat Commun 7:11549
Dalal Y, Wang H, Lindsay S, Henikoff S (2007) Tetrameric structure of centromeric nucleosomes in interphase Drosophila cells. PLoS Biol 5:e218
De Cian A, Praly E, Ding F, Singh V, Lavelle C, Le Cam E, Croquette V, Pietrement O, Bensimon D (2012) ATP-independent cooperative binding of yeast Isw1a to bare and nucleosomal DNA. PLoS ONE 7:e31845
Dechassa ML, Wyns K, Li M, Hall MA, Wang MD, Luger K (2011) Structure and Scm3-mediated assembly of budding yeast centromeric nucleosomes. Nat Commun 2:313
Diaz-Ingelmo O, Martinez-Garcia B, Segura J, Valdes A, Roca J (2015) DNA topology and global architecture of point centromeres. Cell Rep 13:667–677
Duan Z, Andronescu M, Schutz K, McIlwain S, Kim YJ, Lee C, Shendure J, Fields S, Blau CA, Noble WS (2010) A three-dimensional model of the yeast genome. Nature 465:363–367
Durand-Dubief M, Will WR, Petrini E, Theodorou D, Harris RR, Crawford MR, Paszkiewicz K, Krueger F, Correra RM, Vetter AT, Miller JR, Kent NA, Varga-Weisz P (2012) SWI/SNF-like chromatin remodeling factor Fun30 supports point centromere function in S. cerevisiae. PLoS Genet 8:e1002974
Earnshaw WC, Allshire RC, Black BE, Bloom K, Brinkley BR, Brown W, Cheeseman IM, Choo KH, Copenhaver GP, Deluca JG, Desai A, Diekmann S, Erhardt S, Fitzgerald-Hayes M, Foltz D, Fukagawa T, Gassmann R, Gerlich DW, Glover DM, Gorbsky GJ, Harrison SC, Heun P, Hirota T, Jansen LE, Karpen G, Kops GJ, Lampson MA, Lens SM, Losada A, Luger K, Maiato H, Maddox PS, Margolis RL, Masumoto H, McAinsh AD, Mellone BG, Meraldi P, Musacchio A, Oegema K, O’Neill RJ, Salmon ED, Scott KC, Straight AF, Stukenberg PT, Sullivan BA, Sullivan KF, Sunkel CE, Swedlow JR, Walczak CE, Warburton PE, Westermann S, Willard HF, Wordeman L, Yanagida M, Yen TJ, Yoda K, Cleveland DW (2013) Esperanto for histones: CENP-A, not CenH3, is the centromeric histone H3 variant. Chromosome Res 21:101–106
Edwards NS, Murray AW (2005) Identification of xenopus CENP-A and an associated centromeric DNA repeat. Mol Biol Cell 16:1800–1810
Erliandri I, Fu H, Nakano M, Kim JH, Miga KH, Liskovykh M, Earnshaw WC, Masumoto H, Kouprina N, Aladjem MI, Larionov V (2014) Replication of alpha-satellite DNA arrays in endogenous human centromeric regions and in human artificial chromosome. Nucleic Acids Res 42:11502–11516
Errico A, Aze A, Costanzo V (2014) Mta2 promotes Tipin-dependent maintenance of replication fork integrity. Cell Cycle 13:2120–2128
Espelin CW, Kaplan KB, Sorger PK (1997) Probing the architecture of a simple kinetochore using DNA-protein crosslinking. J Cell Biol 139:1383–1396
Espelin CW, Simons KT, Harrison SC, Sorger PK (2003) Binding of the essential Saccharomyces cerevisiae kinetochore protein Ndc10p to CDEII. Mol Biol Cell 14:4557–4568
Fachinetti D, Han JS, McMahon MA, Ly P, Abdullah A, Wong AJ, Cleveland DW (2015) DNA sequence-specific binding of CENP-B enhances the fidelity of human centromere function. Dev Cell 33:314–327
Fernius J, Hardwick KG (2007) Bub1 kinase targets Sgo1 to ensure efficient chromosome biorientation in budding yeast mitosis. PLoS Genet 3:e213
Forsburg SL (2013) The CINs of the centromere. Biochem Soc Trans 41:1706–1711
Fudenberg G, Imakaev M, Lu C, Goloborodko A, Abdennur N, Mirny LA (2016) Formation of chromosomal domains by loop extrusion. Cell Rep 15:2038–2049
Funk M, Hegemann JH, Philippsen P (1989) Chromatin digestion with restriction endonucleases reveals 150–160 bp of protected DNA in the centromere of chromosome XIV in Saccharomyces cerevisiae. Mol Gen Genet 219:153–160
Furuyama T, Henikoff S (2009) Centromeric nucleosomes induce positive DNA supercoils. Cell 138:104–113
Ganem NJ, Pellman D (2012) Linking abnormal mitosis to the acquisition of DNA damage. J Cell Biol 199:871–881
Gkikopoulos T, Singh V, Tsui K, Awad S, Renshaw MJ, Scholfield P, Barton GJ, Nislow C, Tanaka TU, Owen-Hughes T (2011) The SWI/SNF complex acts to constrain distribution of the centromeric histone variant Cse4. EMBO J 30:1919–1927
Goloborodko A, Imakaev MV, Marko JF, Mirny L (2016a) Compaction and segregation of sister chromatids via active loop extrusion. Elife 5:e14864
Goloborodko A, Marko JF, Mirny LA (2016b) Chromosome compaction by active loop extrusion. Biophys J 110:2162–2168
Granick S, Rubinstein M (2004) Polymers: a multitude of macromolecules. Nat Mater 3:586–587
Guerrero AA, Gamero MC, Trachana V, Futterer A, Pacios-Bras C, Diaz-Concha NP, Cigudosa JC, Martinez AC, van Wely KH (2010) Centromere-localized breaks indicate the generation of DNA damage by the mitotic spindle. Proc Nat Acad Sci U S A 107:4159–4164
Guse A, Carroll CW, Moree B, Fuller CJ, Straight AF (2011) In vitro centromere and kinetochore assembly on defined chromatin templates. Nature 477:354–358
Haase J, Mishra PK, Stephens A, Haggerty R, Quammen C, Taylor RM 2nd, Yeh E, Basrai MA, Bloom K (2013) A 3D map of the yeast kinetochore reveals the presence of core and accessory centromere-specific histone. Curr Biol 23:1939–1944
Harris B, Bose T, Lee KK, Wang F, Lu S, Ross RT, Zhang Y, French SL, Beyer AL, Slaughter BD, Unruh JR, Gerton JL (2014) Cohesion promotes nucleolar structure and function. Mol Biol Cell 25:337–346
Hasson D, Panchenko T, Salimian KJ, Salman MU, Sekulic N, Alonso A, Warburton PE, Black BE (2013) The octamer is the major form of CENP-A nucleosomes at human centromeres. Nat Struct Mol Biol 20:687–695
He H, Zhang S, Wang D, Hochwagen A, Li F (2016) Condensin promotes position effects within tandem dna repeats via the rits complex. Cell reports 14:1018–1024
Heidinger-Pauli JM, Mert O, Davenport C, Guacci V, Koshland D (2010) Systematic reduction of cohesin differentially affects chromosome segregation, condensation, and DNA repair. Curr Biol 20:957–963
Henikoff S, Ramachandran S, Krassovsky K, Bryson TD, Codomo CA, Brogaard K, Widom J, Wang JP, Henikoff JG (2014) The budding yeast centromere DNA element II wraps a stable Cse4 hemisome in either orientation in vivo. Elife 3:e01861
Heus JJ, Bloom KS, Zonneveld BJ, Steensma HY, Van den Berg JA (1993a) Chromatin structures of Kluyveromyces lactis centromeres in K. lactis and Saccharomyces cerevisiae. Chromosoma 102:660–667
Heus JJ, Zonneveld BJ, de Steensma HY, van den Berg JA (1993b) The consensus sequence of Kluyveromyces lactis centromeres shows homology to functional centromeric DNA from Saccharomyces cerevisiae. Mol Gen Genet 236:355–362
Hirano T (2012) Condensins: universal organizers of chromosomes with diverse functions. Genes Dev 26:1659–1678
Hsieh TH, Weiner A, Lajoie B, Dekker J, Friedman N, Rando OJ (2015) Mapping nucleosome resolution chromosome folding in yeast by Micro-C. Cell 162:108–119
Hu B, Petela N, Kurze A, Chan KL, Chapard C, Nasmyth K (2015) Biological chromodynamics: a general method for measuring protein occupancy across the genome by calibrating ChIP-seq. Nucleic Acids Res 43:e132
Indjeian VB, Murray AW (2007) Budding yeast mitotic chromosomes have an intrinsic bias to biorient on the spindle. Curr Biol 17:1837–1846
Jayaram M, Ma CH, Kachroo AH, Rowley PA, Guga P, Fan HF, Voziyanov Y (2015) An overview of tyrosine site-specific recombination: from an Flp perspective. Microbiol Spectr 3
Jonstrup AT, Thomsen T, Wang Y, Knudsen BR, Koch J, Andersen AH (2008) Hairpin structures formed by alpha satellite DNA of human centromeres are cleaved by human topoisomerase IIalpha. Nucleic Acid Res 36:6165–6174
Keith KC, Baker RE, Chen Y, Harris K, Stoler S, Fitzgerald-Hayes M (1999) Analysis of primary structural determinants that distinguish the centromere-specific function of histone variant Cse4p from histone H3. Mol Cell Biol 19:6130–6139
Koch J (2000) Neocentromeres and alpha satellite: a proposed structural code for functional human centromere DNA. Hum Mol Genet 9:149–154
Kouprina N, Earnshaw WC, Masumoto H, Larionov V (2013) A new generation of human artificial chromosomes for functional genomics and gene therapy. Cell Mol Life Sci 70:1135–1148
Krassovsky K, Henikoff JG, Henikoff S (2012) Tripartite organization of centromeric chromatin in budding yeast. Proc Natl Acad Sci U S A 109:243–248
Laha S, Das SP, Hajra S, Sanyal K, Sinha P (2011) Functional characterization of the Saccharomyces cerevisiae protein Chl1 reveals the role of sister chromatid cohesion in the maintenance of spindle length during S-phase arrest. BMC Genet 12:83
Lawrimore J, Bloom KS, Salmon ED (2011) Point centromeres contain more than a single centromere-specific Cse4 (CENP-A) nucleosome. J Cell Biol 195:573–582
Lawrimore J, Vasquez PA, Falvo MR, Taylor RM 2nd, Vicci L, Yeh E, Forest MG, Bloom K (2015) DNA loops generate intracentromere tension in mitosis. J Cell Biol 210:553–564
Lawrimore J, Aicher JK, Hahn P, Fulp A, Kompa B, Vicci L, Falvo M, Taylor RM 2nd, Bloom K (2016) ChromoShake: a chromosome dynamics simulator reveals that chromatin loops stiffen centromeric chromatin. Mol Biol Cell 27:153–166
Lezzi M, Gilbert LI (1970) Differential effects of K+ and Na+ on specific bands of isolated polytene chromosomes of Chironomus tentans. J Cell Sci 6:615–627
Liu Y, Nielsen CF, Yao Q, Hickson ID (2014) The origins and processing of ultra fine anaphase DNA bridges. Curr Opin Genet Dev 26:1–5
Logsdon GA, Barrey EJ, Bassett EA, DeNizio JE, Guo LY, Panchenko T, Dawicki-McKenna JM, Heun P, Black BE (2015) Both tails and the centromere targeting domain of CENP-A are required for centromere establishment. J Cell Biol 208:521–531
Magidson V, He J, Ault JG, O’Connell CB, Yang N, Tikhonenko I, McEwen BF, Sui H, Khodjakov A (2016) Unattached kinetochores rather than intrakinetochore tension arrest mitosis in taxol-treated cells. J Cell Biol 212:307–319
Marshall OJ, Chueh AC, Wong LH, Choo KH (2008) Neocentromeres: new insights into centromere structure, disease development, and karyotype evolution. Am J Hum Genet 82:261–282
Martinez AC, van Wely KH (2011) Centromere fission, not telomere erosion, triggers chromosomal instability in human carcinomas. Carcinogenesis 32:796–803
McFarlane RJ, Humphrey TC (2010) A role for recombination in centromere function. Trends Genet 26:209–213
Mizuguchi G, Xiao H, Wisniewski J, Smith MM, Wu C (2007) Nonhistone Scm3 and histones CenH3-H4 assemble the core of centromere-specific nucleosomes. Cell 129:1153–1164
Moreau JL, Lee M, Mahachi N, Vary J, Mellor J, Tsukiyama T, Goding CR (2003) Regulated displacement of TBP from the PHO8 promoter in vivo requires Cbf1 and the Isw1 chromatin remodeling complex. Mol Cell 11:1609–1620
Mythreye K, Bloom KS (2003) Differential kinetochore protein requirements for establishment versus propagation of centromere activity in Saccharomyces cerevisiae. J Cell Biol 160:833–843
Nakagome Y, Abe T, Misawa S, Takeshita T, Iinuma K (1984) The “loss” of centromeres from chromosomes of aged women. Am J Hum Genet 36:398–404
Nakano M, Cardinale S, Noskov VN, Gassmann R, Vagnarelli P, Kandels-Lewis S, Larionov V, Earnshaw WC, Masumoto H (2008) Inactivation of a human kinetochore by specific targeting of chromatin modifiers. Dev Cell 14:507–522
Okumura Y, Ito K (2001) The polyrotaxane gel: a topological gel by figure-of-eight cross-links. Adv Mater 13:485–487
Panyukov S, Zhulina EB, Sheiko SS, Randall GC, Brock J, Rubinstein M (2009a) Tension amplification in molecular brushes in solutions and on substrates (dagger). J Phys Chem B 113:3750
Panyukov SV, Sheiko SS, Rubinstein M (2009b) Amplification of tension in branched macromolecules. Phys Rev Lett 102:148301
Paques F, Haber JE (1999) Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol and Mol Biol Rev 63:349–404
Paques F, Leung WY, Haber JE (1998) Expansions and contractions in a tandem repeat induced by double-strand break repair. Mol Cell Biol 18:2045–2054
Pearson CG, Maddox PS, Zarzar TR, Salmon ED, Bloom K (2003) Yeast kinetochores do not stabilize Stu2p-dependent spindle microtubule dynamics. Mol Biol Cell 14:4181–4195
Perriches T, Singleton MR (2012) Structure of yeast kinetochore Ndc10 DNA-binding domain reveals unexpected evolutionary relationship to tyrosine recombinases. J Biol Chem 287:5173–5179
Pietrasanta LI, Thrower D, Hsieh W, Rao S, Stemmann O, Lechner J, Carbon J, Hansma H (1999) Probing the Saccharomyces cerevisiae centromeric DNA (CEN DNA)-binding factor 3 (CBF3) kinetochore complex by using atomic force microscopy. Proc Natl Acad Sci U S A 96:3757–3762
Russell ID, Grancell AS, Sorger PK (1999) The unstable F-box protein p58-Ctf13 forms the structural core of the CBF3 kinetochore complex. J Cell Biol 145:933–950
Saksouk N, Barth TK, Ziegler-Birling C, Olova N, Nowak A, Rey E, Mateos-Langerak J, Urbach S, Reik W, Torres-Padilla ME, Imhof A, Dejardin J, Simboeck E (2014) Redundant mechanisms to form silent chromatin at pericentromeric regions rely on BEND3 and DNA methylation. Mol Cell 56:580–594
Samoshkin A, Arnaoutov A, Jansen LE, Ouspenski I, Dye L, Karpova T, McNally J, Dasso M, Cleveland DW, Strunnikov A (2009) Human condensin function is essential for centromeric chromatin assembly and proper sister kinetochore orientation. PLoS ONE 4:e6831
Santaguida S, Amon A (2015) Short- and long-term effects of chromosome mis-segregation and aneuploidy. Nat Rev Mol Cell Biol 16:473–485
Saunders M, Fitzgerald-Hayes M, Bloom K (1988) Chromatin structure of altered yeast centromeres. Proc Natl Acad Sci U S A 85:175–179
Shivaraju M, Unruh JR, Slaughter BD, Mattingly M, Berman J, Gerton JL (2012) Cell-cycle-coupled structural oscillation of centromeric nucleosomes in yeast. Cell 150:304–316
Simi S, Simili M, Bonatti S, Campagna M, Abbondandolo A (1998) Fragile sites at the centromere of Chinese hamster chromosomes: a possible mechanism of chromosome loss. Mutat Res 397:239–246
Snider CE, Stephens AD, Kirkland JG, Hamdani O, Kamakaka RT, Bloom K (2014) Dyskerin, tRNA genes, and condensin tether pericentric chromatin to the spindle axis in mitosis. J Cell Biol 207
Stephens AD, Haase J, Vicci L, Taylor RM 2nd, Bloom K (2011) Cohesin, condensin, and the intramolecular centromere loop together generate the mitotic chromatin spring. J Cell Biol 193:1167–1180
Stephens AD, Haggerty RA, Vasquez PA, Vicci L, Snider CE, Shi F, Quammen C, Mullins C, Haase J, Taylor RM 2nd, Verdaasdonk JS, Falvo MR, Jin Y, Forest MG, Bloom K (2013a) Pericentric chromatin loops function as a nonlinear spring in mitotic force balance. J Cell Biol 200:757–772
Stephens AD, Snider CE, Haase J, Haggerty RA, Vasquez PA, Forest MG, Bloom K (2013b) Individual pericentromeres display coordinated motion and stretching in the yeast spindle. J Cell Biol 203:407–416
Stigler J, Camdere GO, Koshland DE, Greene EC (2016) Single-molecule imaging reveals a collapsed conformational state for dna-bound cohesin. Cell Rep 15:988–998
Sun M, Nishino T, Marko JF (2013) The SMC1-SMC3 cohesin heterodimer structures DNA through supercoiling-dependent loop formation. Nucleic Acids Res 41:6149–6160
Tsabar M, Haase J, Harrison B, Snider CE, Eldridge B, Kaminsky L, Hine RM, Haber JE, Bloom K (2016) A cohesin-based partitioning mechanism revealed upon transcriptional inactivation of centromere. PLoS Genet 12:e1006021
Tsouroula K, Furst A, Rogier M, Heyer V, Maglott-Roth A, Ferrand A, Reina-San-Martin B, Soutoglou E (2016) Temporal and spatial uncoupling of dna double strand break repair pathways within mammalian heterochromatin. Mol Cell 63:293–305
Vasquez PA, Hult C, Adalsteinsson D, Lawrimore J, Forest MG, Bloom K (2016) Entropy gives rise to topologically associating domains. Nucleic Acids Res 44:5540–5549
Verdaasdonk JS, Gardner R, Stephens AD, Yeh E, Bloom K (2012) Tension-dependent nucleosome remodeling at the pericentromere in yeast. Mol Biol Cell 23:2560–2570
Wang X, Le TB, Lajoie BR, Dekker J, Laub MT, Rudner DZ (2015) Condensin promotes the juxtaposition of DNA flanking its loading site in Bacillus subtilis. Genes Dev 29:1661–1675
Weir JR, Faesen AC, Klare K, Petrovic A, Basilico F, Fischbock J, Pentakota S, Keller J, Pesenti ME, Pan D, Vogt D, Wohlgemuth S, Herzog F, Musacchio A (2016) Insights from biochemical reconstitution into the architecture of human kinetochores. Nature 537:249–253
Wisniewski J, Hajj B, Chen J, Mizuguchi G, Xiao H, Wei D, Dahan M, Wu C (2014) Imaging the fate of histone Cse4 reveals de novo replacement in S phase and subsequent stable residence at centromeres. Elife 3:e02203
Xiao H, Mizuguchi G, Wisniewski J, Huang Y, Wei D, Wu C (2011) Nonhistone Scm3 binds to AT-rich DNA to organize atypical centromeric nucleosome of budding yeast. Mol Cell 43:369–380
Yeh E, Haase J, Paliulis LV, Joglekar A, Bond L, Bouck D, Salmon ED, Bloom KS (2008) Pericentric chromatin is organized into an intramolecular loop in mitosis. Curr Biol 18:81–90
Zebarjadian Y, King T, Fournier MJ, Clarke L, Carbon J (1999) Point mutations in yeast CBF5 can abolish in vivo pseudouridylation of rRNA. Mol Cell Biol 19:7461–7472
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Bloom, K., Costanzo, V. (2017). Centromere Structure and Function. In: Black, B. (eds) Centromeres and Kinetochores. Progress in Molecular and Subcellular Biology, vol 56. Springer, Cham. https://doi.org/10.1007/978-3-319-58592-5_21
Download citation
DOI: https://doi.org/10.1007/978-3-319-58592-5_21
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-58591-8
Online ISBN: 978-3-319-58592-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)