Abstract
There are currently few antifungals in use which show efficacy against fungal diseases. These antifungals mostly target specific components of fungal plasma membrane or its biosynthetic pathways. However, more recent class of antifungals in use is echinocandins which target the fungal cell wall components. The availability of mostly fungistatic antifungals in clinical use, often led to the development of tolerance to these very drugs by the pathogenic fungal species. Thus, the development of clinical multidrug resistance (MDR) leads to higher tolerance to drugs and its emergence is helped by multiple mechanisms. MDR is indeed a multifactorial phenomenon wherein a resistant organism possesses several mechanisms which contribute to display reduced susceptibility to not only single drug in use but also show collateral resistance to several drugs. Considering the limited availability of antifungals in use and the emergence of MDR in fungal infections, there is a continuous need for the development of novel broad spectrum antifungal drugs with better efficacy. Here, we briefly present an overview of the current understanding of the antifungal drugs in use, their mechanism of action and the emerging possible novel antifungal drugs with great promise.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
14.1 Introduction
Fungal infections have emerged as one of the major causes of human disease, especially in immunocompromised individuals (Shapiro et al. 2011). Fungal infections which are generally superficial can also turn into systemic infections as the disease incidence prolongs (Cannon et al. 2009; Brown et al. 2012). Among the different mycotic infections caused by these opportunistic fungi, candidiasis, an infection caused by Candida, is the most threatening due to severity of the disease and higher worldwide occurrence. Other fungal diseases like cryptococcal meningitis and invasive aspergillosis are also life threatening (López-Martínez 2010). The pathogenicity of fungal infections proceeds in well-organized steps. For example, Candida cell surface adhesion factors first promote its adherence to host surface, followed by an invasion and damage of the host tissues due to release of various virulence factors (Gow and Hube 2012). Eukaryotic fungal pathogens pose an additional therapeutic challenge since they show close evolutionary relationship with the human hosts, thus minimizing the choice of novel drug targets that can be exploited to selectively kill the pathogen (Heitman 2011; Shapiro et al. 2011). Nonetheless, there are many drug categories currently in use against fungal infections which exploit exclusive novel fungal targets. For example, most antifungals are directed against ergosterol which is a typical sterol of fungal cells. The sterol component of cell membranes of fungi is targeted either by blocking the enzymes important for its synthesis or by directly depleting the ergosterol from the plasma membrane (PM) (White et al. 1998). In addition, several other drugs, now in use, also target unique components of cell wall (CW) in fungal cells (White et al. 1998). The fact that the increase in incidence of worldwide fungal infections and emergence of antifungal drug resistance which outcompetes the development of novel antifungal compounds, it becomes important to understand the various facets of infections and to understand the basic mechanisms that govern the development of resistance. This enforces the widening of the hunt for the development of new antifungals targeting novel pathways. This chapter focuses on some of the aspects of antifungals, their mechanisms of action and development of resistance against them.
14.2 Antifungal Drugs
The limited availability of antifungals is a major impediment for the effective treatment of fungal infections (Vandeputte et al. 2012). This is further compounded by the fact that the generation of newer antifungals has lagged behind when compared to the pace of emergence of fungal infections. The components of the fungal CW such as mannans, glucans and chitins; and a few of the enzymes of the ergosterol biosynthetic pathways which are unique to fungal cells are commonly targeted for the development of antifungal agents (St Georgiev 2000; Munro et al. 2001). Among the enzymes of the ergosterol biosynthetic pathway, squalene epoxidase (ERG1), 14α-lanosterol demethylase or CYP51 (ERG11), Δ14–reductase (ERG24) and Δ8-Δ7-isomerase (ERG2) have been the targets of most antifungal agents (Fig. 14.1) (Sanglard et al. 2003). Some of the commonly used antifungal drugs and their mechanisms of action are discussed below:
14.2.1 Azoles
The fungistatic azoles primarily act on ergosterol biosynthesis by targeting 14α-lanosterol demethylase encoded by ERG11 gene resulting in the inhibition of cytochrome P450-dependent conversion of lanosterol to ergosterol (Fig. 14.1). The resulting ergosterol depletion interferes with the bulk functions of ergosterol as a membrane component, but more importantly, severe ergosterol depletion may also interfere with the “sparking” functions of ergosterol, affecting cell growth and proliferation (White et al. 1998; Sanglard et al. 2009; Shapiro et al. 2011). The blocking of 14α-demethylase results in the accumulation of toxic methylated sterols leading to the membrane stress (Shapiro et al. 2011). In case of yeast Cryptococcus neoformans, azoles such as fluconazole (FLC) and itraconazole also result in the accumulation of obtusifolione in the ergosterol biosynthetic pathway, mainly due to the inhibition of NADPH-dependent 3-ketosteroid reductase (ERG 27), catalyzing the last C-4 demethylation step in ergosterol biosynthesis (Vanden Bossche et al. 1993; Ghannoum et al. 1994).
Azoles mainly include two subclasses based on the number of nitrogen atoms in a ring; The first class includes imidazoles which consist of miconazole, oxiconazole, econazole, ketoconazole, tioconazole, and clotrimazole with two nitrogen atoms in an azole ring, while another class includes triazoles such as FLC, posaconazole, itraconazole, terconazole, and voriconazole which contain three nitrogen atoms in a cyclic ring (Fig. 14.2). Imidazoles are mainly used for the mucosal fungal infections while triazoles are administered both for the systemic as well as for the mucosal infections (Sanglard et al. 2009; Vandeputte et al. 2012). Depletion of membrane ergosterol due to the use of azoles are also shown to disrupt vacuolar ATPase functions resulting in an impairment of the vacuolar acidification and ion homeostasis (Zhang et al. 2010). Since azoles are fungistatic, their prolonged use poses greater threat of emergence of drug resistance among the surviving fungal population (Shapiro et al. 2011).
Structure of various azole antifungal compounds. These include imidazoles with two nitrogen atoms in a ring (i) Clotrimazole, (ii) Econazole, (iii) Ketoconazole (iv) Miconazole, (v) Oxiconazole, (vi) Tioconazole or triazoles containing three nitrogen atoms in a ring, (vii) Itraconazole (viii) Fluconazole (ix) Voriconazole (x) Posaconazole
14.2.2 Polyenes
Polyenes are the amphipathic organic natural molecules called macrolides and are generally produced by Streptomyces (Vandeputte et al. 2012). Polyenes directly bind to ergosterol of fungal cell membranes leading to the formation of pores in membrane, resulting in the loss of ionic balance, membrane integrity and cell death (Sanglard et al. 2009) (Fig. 14.1). Polyenes mainly include amphotericin B (AmpB), natamycin and nystatin (Fig. 14.3). AmpB is mostly effective in systemic invasive fungal infections and is used generally against Cryptococcus, Candida and Aspergillus species (Lemke et al. 2005; Sanglard et al. 2009) while nystatin and natamycin are preferred for topical infections due to their low absorption (Vandeputte et al. 2012). Although polyenes are fungicidal in nature and have been in use for a long time but they show many side effects in humans which limits their use. However, lipid formulations of AmpB are less toxic and are relatively better for the treatment of fungal infections (Shapiro et al. 2011).
14.2.3 Pyrimidine Analogs
Pyrimidine analogs which include 5-fluorocytosine (5-FC) and 5-fluorouracil (5-FU) are the synthetic structural analogs of nucleotide cytosine (Fig. 14.3). Pyrimidine analog 5-FC is converted to 5-FU by cytosine deaminase which after conversion to downstream products gets incorporated into DNA and RNA during the synthesis of these biomolecules where it inhibits cellular functioning by blocking protein synthesis or inhibiting DNA replication. These drug analogs show activity against different Candida and Cryptococcus species (Lemke et al. 2005; Sanglard et al. 2009). 5-FC is rapidly absorbed and thus gives good bioavailability, however; it also shows many side effects (Lemke et al. 2005; Vandeputte et al. 2012). 5-FC is comparatively less effective antifungal drug because the fungal cells frequently develop tolerance to it. For this reason, it is generally preferred in combination therapy (Sanglard et al. 2009).
14.2.4 Allylamine, Thiocarbamates and Morpholines
Allylamines and thiocarbamates inhibit the ERG1 gene of ergosterol biosynthesis while morpholines which include fenpropimorph and amorolfine (Fig. 14.3) inhibit the ERG24 and ERG2 genes of ergosterol biosynthesis (Fig. 14.1). Allylamines include terbinafine while thiocarbamates include tolnaftate (Fig. 14.3). All of these drugs are mostly used for the control of dermatophyte fungal infections (Gubbins and Anaissie 2006; Sanglard et al. 2009).
14.2.5 Echinocandins
The lipopeptide echinocandins which include caspofungin, micafungin and anidulafungin (Fig. 14.4) are comparatively recent class of antifungal drugs which target the synthesis of CW components by acting as non-competitive inhibitors of β-1,3 glucan synthase required for β-glucan synthesis (Fig. 14.5) (Perlin 2011; Shapiro et al. 2011). Defects in the synthesis of CW components affect the integrity of fungal cells resulting in CW stress. As a result, echinocandin treated cells become osmotically sensitive, form pseudohyphae, show separation defects, reduced sterol contents and thickened CW. Echinocandins are generally non-toxic to mammalian cells because they act on specific CW synthesis pathway unique to fungal cells (Sanglard et al. 2009; Perlin 2011; Shapiro et al. 2011).
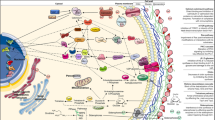
Different mechanisms of multidrug resistance adopted by fungal cells. The commonly observed mechanisms of drug resistance particularly against azoles, polyenes and echinocandins include (1) changes in membrane property/ lipid composition affecting normal drug import (2) over expression of drug efflux proteins leading to rapid drug extrusion (3) an alteration of the drug target (genes encoding ergosterol biosynthetic pathway enzymes or glucan synthases) leading to poor binding of toxic drugs to its target (4) overproduction of genes synthesizing the drug target proteins. (5) Echinocandins block the activity of glucan synthase important for synthesis of CW components affecting CW integrity leading to cell stress
14.2.6 Emerging Novel Antifungals
Keeping in view the facts that there are limited antifungal strategies and paucity of effective drugs, there is continuous hunt for the development of novel and effective antifungals to combat the fungal infections. Many different drug categories which show promising antifungal activity are at various stages of the development and some of them are listed in Table 14.1 and few potential antifungal strategies are discussed in the following section.
14.2.7 Emerging Antifungal Strategies
As mentioned above, the development of novel drug candidates have not kept pace with the frequency of the development of tolerance to available drugs, it is imperative to search for the different strategies to combat fungal infections. For instance, the transcription factor Upc2 which regulates the expression of ERG genes has been exploited as a potential target for the development of antifungal drugs. Many small molecules screened from the commercially available compounds collection library have already been shown to inhibit the azole mediated up regulation of Upc2 and its target genes in S. cerevisiae and C. glabrata. However, the full potential of Upc2 remains to be explored before it could be successfully employed to improve antifungal strategies (Gallo-Ebert et al. 2014). Several new targets which include glucan synthesis, 26S proteasome, cAMP homeostasis, microtubule dynamics, and translational elongation also show great promise as antifungal targets (Roemer and Krysan 2014).
Combination therapy wherein different drugs are given in combination to treat fungal infections is among most favorite strategies. Some of the benefits of combination therapies include broad spectrum of treatment, synergy of effects between different drugs, lower doses of drug usage and lesser chances of the development of drug resistance. Synergistic effect shown by the drugs mainly occurs due to the additive effects on both CW and cell membrane components of fungal cell. For instance, the damaged CW due to one of the antifungal components potentiates effective action of drugs directed against cell membrane components. The compromised CW integrity could also facilitate permeability of drugs across cell membranes to intracellular targets. Combination of azoles and allylamines displays synergistic effects due to the inhibition of the same pathway at different steps (Tobudic et al. 2010a; Rodrigues et al. 2014). Notably, combination therapy requires critical evaluation of the possible antagonistic and agonist property of different drugs when administered in combinations (Lewis and Kontoyiannis 2001). Table 14.2 summarizes some of the drugs which show better efficacy when given in combination.
14.3 Resistance Against Antifungal Drugs
Fungal cells have developed several strategies to deal with the antifungals. They have learnt to modify the antifungal drug targets or most commonly increase the efflux of the incoming drugs. Some of the known mechanisms of MDR are depicted in Fig. 14.5 and are briefly discussed below.
14.3.1 Azole Target Protein (Erg11p) Is Modified in Resistant Isolates of Candida
The modification of the target protein represents one of the commonest mechanisms of MDR where the target protein of azoles, Erg11p, is modified by the chromosomal mutations leading to the replacement of native amino acids. This is evident from the fact that several point mutations in ERG11 gene which encodes Erg11p have been identified in clinical drug resistant isolates of Candida. Interestingly, these mutations appear to be predominantly restricted to certain hot spot regions of Erg11p. The exact placement of all the identified mutations in a 3D model of the protein confirms that these mutations are not randomly distributed but rather are clustered in select hot spot regions (Marichal et al. 1999; Wang et al. 2009). These point mutations either individually or in combination, invariably prevent normal binding of FLC to target protein by reducing the affinity of the drug towards Erg11p (Wang et al. 2009; Morio et al. 2010).
14.3.2 Azole Resistance Leads to an Overexpression of ERG11
Apart from spontaneous point mutations in ERG11 (discussed above), in many FLC resistant clinical isolates, very often an over expression of ERG11 is also observed (Hoot et al. 2011; Flowers et al. 2012; Sasse et al. 2012). The zinc cluster transcription factor Upc2p regulates the expression of ERG11 and other genes involved in ergosterol biosynthesis (White and Silver 2005). Several studies have confirmed that an overexpression of UPC2 increases resistance of Candida cells towards azole drugs, while its disruption results in cells hypersusceptibility to azoles. A comparison of sequence of UPC2 between genetically matched pair azole susceptible and resistant isolates led to the identification of point mutations in the encoded protein. These gain of function (GOF) mutations results in an over expression of ERG11 and hyper-resistance to azoles (Heilmann et al. 2010; Hoot et al. 2011; Flowers et al. 2012).
14.3.3 Azole Resistance and Ergosterol Biosynthetic Pathway
In clinical azole resistance, several mutations have also been detected in other ERG genes. For example, the point mutations in ERG3 also occur which could be present either alone or in combination with ERG11 mutations, resulting in a change in the ratios of various cell sterol biosynthetic intermediates, and increased tolerance to azoles and polyenes. The cytochrome P450 spectral studies performed in a system reconstituted with purified ERG5 (Δ22-desaturase or CYP61) of C. glabrata revealed an interactions between azoles and the heme-protein, implying that ERG5 could also be a target of azoles and may contribute in the development of antifungal resistance (Lamb et al. 1999). Indeed, C. albicans drug resistant clinical isolate with a combination of a single mutation in the ERG5 gene along with a stretch of amino acid duplication in the ERG11 gene has been identified (Martel et al. 2010). This mutant was not only resistant to azoles but also displayed collateral resistance to AmpB due to the depletion of membrane ergosterol (Martel et al. 2010). ERG6 in C. glabrata is involved in azole resistance due to various base pair alterations leading to missense mutations (Vandeputte et al. 2007). Similarly, an erg6 disruptant strain of C. lusitaniae was susceptible to AmpB due to decreased membrane ergosterol levels. Coinciding with this, several clinical isolates of C. lusitaniae show increased expression of ERG6 along with a decrease in ERG3 expression and enhanced resistance to AmpB (Young et al. 2003).
Together, the azole-induced upregulation of ERG11, along with other genes of the ergosterol biosynthetic pathway, suggests the existence of a common mechanism of upregulation in C. albicans (Henry et al. 2000). Transcript profiling of azole treated Candida show that almost 15 % of genes differentially expressed upon drug treatment fall under the category of sterol metabolism in a wild type strain of C. albicans (Liu et al. 2005). Notably, while a global regulation of ERG genes was evident from the transcript profiling, several genes of diverse functions as well as of unknown functions were also differentially regulated by the drug treatment (De Backer et al. 2001; Liu et al. 2005). This reinforces that azole resistance could be the result of many factors which remains to be identified. The dissection of the mechanisms mediating these phenotypes could provide newer insights into the phenomenon of MDR.
14.3.4 Drug Import Impacts Tolerance
The hydrophobic nature of drugs facilitates their easy import by passive diffusion. However, the contribution of drug import in the overall scenario of MDR is not well established. Nonetheless, there are a few instances to suggest that passive diffusion of drugs could be an important determinant of MDR. For example, fluctuations in membrane fluidity are shown to affect passive diffusion leading to an increase in susceptibility to drugs. The erg mutants of C. albicans possess high membrane fluidity, which led to an enhanced diffusion and susceptibility to azoles (Kohli et al. 2002; Prasad et al. 2010). In another study, permeability constrains imposed by Candida cells have been reemphasized in the development of MDR. It is shown that azoles can enter in C. albicans, C. kruesi and C. neoformans cells by diffusion (Mansfield et al. 2010). The kinetics of import in de-energized cells suggests that FLC import proceeds via facilitated diffusion (FD) mediated through a transporter rather than by passive diffusion. Other azoles compete for FLC import, suggesting that all the azoles utilize the same FD mechanism. FLC import was also shown to vary among C. albicans resistant clinical isolates, suggesting that altered FD may be a previously uncharacterized mechanism of resistance to azole drugs (Mansfield et al. 2010). However, the identification of a membrane transporter protein involved in FD of azoles remains elusive (Mansfield et al. 2010). Interestingly, drug inactivation which is a common mechanism in bacteria has not been observed in Candida cells.
14.3.5 Drug Efflux as a Common Strategy of Drug Tolerance
Increased efflux, which results in reduced intracellular accumulation of the incoming drugs, is another prominent mechanism of MDR in fungi (Prasad et al. 1995; Prasad and Kapoor 2005). In C. albicans, for example, this is achieved by increasing the efflux of drugs from cells by overproducing the PM efflux pump proteins. An over expression of genes encoding efflux pump proteins, particularly ABC (ATP Binding Cassette) multidrug transporter proteins Cdr1 and Cdr2 or MFS (Major Facilitator Superfamily) efflux pump protein Mdr1, have been commonly observed in azole resistant clinical isolates of C. albicans (White T et al. 2002; Karababa et al. 2004; Kusch et al. 2004; Prasad and Kapoor 2005). Invariably, MDR Candida cells, which show enhanced expression of efflux pump encoding genes, also show simultaneous increase in the efflux of drugs, thus implying a causal relationship between efflux pump encoding gene expression levels and intracellular concentration of the drug (Cannon et al. 2009). A brief description of these transporters is included, however, for more details, the reader is recommended to see the accompanying chapter “Efflux Pump Proteins of Candida in Clinical Drug Resistance”.
14.3.5.1 ABC Transporters
The inventory of ABC transporters of C. albicans revealed that there are twenty-eight putative ABC superfamily members, including twelve half transporters that largely remain uncharacterized (Gaur et al. 2005). ABC transport proteins are classified into nine families (A to I) according to the nomenclature adopted by the Human Genome Organization (HUGO) (Dean et al. 2001; Verrier et al. 2008). Of these, yeast proteins belonging to ABCB (MDR) (Thornewell et al. 1997; Sanguinetti et al. 2006; Lamping et al. 2010), ABCC (MRP) (Decottignies et al. 1998; Pagant et al. 2010) and ABCG (PDR) (Golin et al. 2007; Prasad and Goffeau 2012) transporters are most often associated with the antifungal resistance.
Full ABC proteins are made up of two (or three) transmembrane domains (TMDs) and two cytoplasmic nucleotide-binding domains (NBDs). NBDs are the nucleotide binding sites, which bind and hydrolyze ATP required to power the efflux of substrates bound within TMDs drug binding sites. Each TMD is usually comprised of six transmembrane segments (TMS), which generally are continuous alpha helices arranged to form drug binding sites (Prasad and Goffeau 2012).
The PDR protein subfamily of C. albicans comprises seven full-size members: Cdr1p, Cdr2p, Cdr3p, Cdr4p, Cdr11p, CaSnq2p and Ca4531. The C. albicans Cdr1p and Cdr2p proteins are active multidrug transporters, while Cdr3p and Cdr4p do not efflux drugs and play no apparent role in the development of antifungal resistance (Prasad and Goffeau 2012). Other transporters in related fungi, including CgCDR1 (Sanglard et al. 1999), CgCDR2 (PDH1) (Miyazaki et al. 1998) and SNQ2 (Torelli et al. 2008) in C. glabrata, ABC1 in C. krusei (Katiyar and Edlind 2001) and AFR1 in C. neoformans (Sanguinetti et al. 2006), are multidrug transporters and play a role in the development of MDR in these pathogenic species.
14.3.5.2 MFS Transporters
MFS transporters are the second major superfamily of transporters (Saier et al. 1999). A phylogenetic analysis identified 95 putative MFS transporters in C. albicans (Gaur et al. 2008). Most MFS transporters consist of two domains of six-TMSs within a single polypeptide chain with few exceptions (Stephanie et al. 1998). On the basis of hydropathy and phylogenetic analysis, the drug efflux MFS proteins can be divided into two distinct types; Drug: H+ Antiporter-1 (DHA1), consisting of 12 TMSs and Drug: H+ Antiporter-2 (DHA2) that contains 14 TMSs. MDR1 of DHA1 subfamily is a major multidrug transporter of C. albicans. Homologues of CaMDR1 have been identified from C. dubliniensis and C. glabrata, which are designated as CdMDR1 and CgMDR1, respectively (Moran et al. 1998; Sanglard et al. 1999). It appears that an increased expression of CdMDR1 is one of the main mechanisms of FLC resistance in clinical isolates of C. dubliniensis (Moran et al. 1998). Since CgMDR1 confers specific resistance to FLC, its constitutive expression in C. glabrata may be responsible for the intrinsically low susceptibility of this yeast species to triazoles (Sanglard et al. 1999).
Among all the MFS proteins, only one member, MDR1, has been implicated clinically to be involved in azole resistance in S. cerevisiae. FLU1, a close homologue of MDR1 has also been implicated in FLC resistance in S. cerevisiae. However, an over expression of FLU1 has not been detected in FLC resistant clinical isolates of C. albicans. None of the other 95 members of this superfamily are implicated in MDR (Gaur et al. 2008).
As an important MDR gene of the MFS family, MDR1 of C. albicans has been extensively studied for its role in drug resistance. The functional evaluation of critical amino acid residues of the Mdr1 protein revealed that the residues of TMS5 which harbor antiporter motifs are potentially significant for their functionality and contribute to drug:H+ transport. Independent of the substrate specificity of the antiporter, the antiporter motif in the predicted TMS5 is well conserved in all of the functionally related subgroups in bacteria and plants (Pasrija et al. 2007).
14.3.6 Echinocandin Resistance
Echinocandins inhibit the synthesis of β-1,3-glucans which is one of the major component of fungal CW (Fig. 14.5). Mutation in the FKS genes encoding echinocandin drug target glucan synthase enzyme results in its decreased sensitivity towards drug and development of resistance (Fig. 14.5) (Perlin 2007). Point mutations in FKS genes are the only known mechanism by which fungi develop resistance to echinocandin antifungal drugs (Park et al. 2005; Balashov et al. 2006). Drug resistant mutations developed in FKS genes generally fall in two “hot spots” regions of FKS1, essential for enzyme activity (Perlin 2007). Garcia-Effron et al. has also reported that mutations in Fks1 protein (Fks1p) lower the activity of β-glucan synthase without altering its affinity for the drug as is also the case with azole drug target Erg11p (Garcia-Effron et al. 2009). Fks1p mutations in “hot-spot” regions have been characterized in C. albicans as well as in non-albicans spp. (Perlin 2011). A paralog of FKS1 in C. glabrata, FKS2, is also responsible for echinocandin resistance (Perlin 2007).
14.4 Concluding Remarks
The available arsenals of antifungals targeting mostly sterols or its synthesis machinery or CW components of fungal cells are reasonably successful in combating fungal infections. However, the fungistatic nature of many of these antifungals limits their success. The synergy among different drugs is being also projected as an alternate strategy. The limited availability of antifungals and emergence of clinical drug resistance necessitate search of newer compounds and new targets. The current research does promise for novel targets and better drugs.
References
Balashov SV, Park S, Perlin DS (2006) Assessing resistance to the echinocandin antifungal drug caspofungin in Candida albicans by profiling mutations in FKS1. Antimicrob Agents Chemother 50:2058–2063
Bills GF, Platas G, Overy DP, Collado J, Fillola A, Jiménez MR et al (2009) Discovery of the parnafungins, antifungal metabolites that inhibit mRNA polyadenylation, from the Fusarium larvarum complex and other Hypocrealean fungi. Mycologia 101:449–472
Bink A, Pellens K, Cammue BPA, Thevissen K (2011) Anti-Biofilm Strategies: how to Eradicate Candida Biofilms? Open Mycol J 5:29–38
Brown GD, Denning DW, Gow NA, Levitz SM, Netea MG, White TC (2012) Hidden killers: human fungal infections. Sci Transl Med 4:165rv13
Calabrese EC, Castellano S, Santoriello M, Sgherri C, Quartacci MF, Calucci L et al (2013) Antifungal activity of azole compounds CPA18 and CPA109 against azole-susceptible and -resistant strains of Candida albicans. J Antimicrob Chemother 68:1111–1119
Cannon RD, Lamping E, Holmes AR, Niimi K, Baret PV, Keniya MV et al (2009) Efflux-mediated antifungal drug resistance. Clin Microbiol Rev 22:291–321
Capobianco JO, Zakula D, Coen ML, Goldman RC (1993) Anti-Candida activity of cispentacin: the active transport by amino acid permeases and possible mechanisms of action. Biochem Biophys Res Commun 190:1037–1044
De Backer MD, Ilyina T, Ma XJ, Vandoninck S, Luyten WH, Vanden Bossche H (2001) Genomic profiling of the response of Candida albicans to itraconazole treatment using a DNA microarray. Antimicrob Agents Chemother 45:1660–1670
Dean M, Andrey R, Rando A (2001) The human ATP-binding cassette (ABC) transporter superfamily. Genome Res 11:1156–1166
Decottignies A, Grant AM, Nichols JW, De Wet H, McIntosh DB, Goffeau A (1998) ATPase and multidrug transport activities of the overexpressed yeast ABC protein Yor1p. J Biol Chem 273:12612–12622
Dhamgaye S, Devaux F, Manoharlal R, Vandeputte P, Shah AH, Singh A et al (2012) In vitro effect of malachite green on Candida albicans involves multiple pathways and transcriptional regulators UPC2 and STP2. Antimicrob Agents Chemother 56:495–506
Dhamgaye S, Devaux F, Vandeputte P, Khandelwal NK, Sanglard D, Mukhopadhyay G et al (2014) Molecular mechanisms of action of herbal antifungal alkaloid berberine, in Candida albicans. PLoS One 9:e104554
Domínguez JM, Martín JJ (1998) Identification of elongation factor 2 as the essential protein targeted by sordarins in Candida albicans. Antimicrob Agents Chemother 42:2279–2283
Domínguez JM, Kelly VA, Kinsman OS, Marriott MS, Gómez de las Heras F, Martín JJ (1998) Sordarins: a new class of antifungals with selective inhibition of the protein synthesis elongation cycle in yeasts. Antimicrob Agents Chemother 42:2274–2278
Espinel-Ingroff A (2009) Novel antifungal agents, targets or therapeutic strategies for the treatment of invasive fungal diseases: a review of the literature (2005–2009). Rev Iberoam Micol 26:15–22
Espinel-Ingroff A, Canton E, Martin-Mazuelos E, Pemán J (2009) Pharmacotherapy of Candida Infections with Echinocandins. Clin Med Ther 1:889–897
Flowers SA, Barker KS, Berkow EL, Toner G, Chadwick SG, Gygax SE et al (2012) Gain-of function mutations in UPC2 are a frequent cause of ERG11 upregulation in azole-resistant clinical isolates of Candida albicans. Eukaryot Cell 11:1289–1299
Fostel JM, Lartey PA (2000) Emerging novel antifungal agents. Drug Discov Today 5:25–32
Gallo-Ebert C, Donigan M, Stroke IL, Swanson RN, Manners MT, Francisco J et al (2014) Novel antifungal drug discovery based on targeting pathways regulating the fungus-conserved Upc2 transcription factor. Antimicrob Agents Chemother 58:258–266
Garbati MA, Alasmari FA, Al-Tannir MA, Tleyjeh IM (2012) The role of combination antifungal therapy in the treatment of invasive aspergillosis: a systematic review. Int J Infect Dis 16:e76–e81
Garcia-Effron G, Park S, Perlin DS (2009) Correlating echinocandin MIC and kinetic inhibition of fks1 mutant glucan synthases for Candida albicans: implications for interpretive breakpoints. Antimicrob Agents Chemother 53:112–122
Gaur M, Choudhury D, Prasad R (2005) Complete inventory of ABC proteins in human pathogenic yeast, Candida albicans. J Mol Microbiol Biotechnol 9:3–15
Gaur M, Puri N, Manoharlal R, Rai V, Mukhopadhayay G, Choudhury D et al (2008) MFS transportome of the human pathogenic yeast Candida albicans. BMC Genomics 9:579
Ghannoum MA, Spellberg BJ, Ibrahim AS, Ritchie JA, Currie B, Spitzer ED et al (1994) Sterol composition of Cryptococcus neoformans in the presence and absence of fluconazole. Antimicrob Agents Chemother 38:2029–2033
Golin J, Ambudkar SV, May L (2007) The yeast Pdr5p multidrug transporter: how does it recognize so many substrates? Biochem Biophys Res Commun 356:1–5
Gow NA, Hube B (2012) Importance of the Candida albicans cell wall during commensalism and infection. Curr Opin Microbiol 15:406–412
Gubbins PO, Anaissie E (2006) Overview of antifungal agents. Pharmacy practice news special edition 59–64
Gunawardana G, Rasmussen RR, Scherr M, Frost D, Brandt KD, Choi W et al (1997) Corynecandin: a novel antifungal glycolipid from Coryneum modonium. J Antibiot (Tokyo) 50:884–886
Harris GH, Shafiee A, Cabello MA, Curotto JE, Genilloud O, Göklen KE et al (1998) Inhibition of fungal sphingolipid biosynthesis by rustmicin, galbonolide B and their new 21-hydroxy analogs. J Antibiot (Tokyo) 51:837–844
Heilmann CJ, Schneider S, Barker KS, Rogers PD, Morschhäuser J (2010) An A643T mutation in the transcription factor Upc2p causes constitutive ERG11 upregulation and increased fluconazole resistance in Candida albicans. Antimicrob Agents Chemother 54:353–359
Heitman J (2011) Microbial pathogens in the fungal kingdom. Fungal Biol Rev 25:48–60
Henry KW, Nickels JT, Edlind TD (2000) Upregulation of ERG genes in Candida species by azoles and other sterol biosynthesis inhibitors. Antimicrob Agents Chemother 44:2693–2700
Herreros E, Martinez CM, Almela MJ, Marriott MS, De Las Heras FG, Gargallo-Viola D (1998) Sordarins: in vitro activities of new antifungal derivatives against pathogenic yeasts, Pneumocystis carinii, and filamentous fungi. Antimicrob Agents Chemother 42:2863–2869
Hodgetts S, Nooney L, Al-Akeel R, Curry A, Awad S, Matthews R et al (2008) Efungumab and caspofungin: pre-clinical data supporting synergy. J Antimicrob Chemother 61:1132–1139
Hoot SJ, Smith AR, Brown RP, White TC (2011) An A643V amino acid substitution in Upc2p contributes to azole resistance in well-characterized clinical isolates of Candida albicans. Antimicrob Agents Chemother 55:940–942
Iwamoto T, Tsujii E, Ezaki M, Fujie A, Hashimoto S, Okuhara M et al (1990) FR109615, a new antifungal antibiotic from Streptomyces setonii. Taxonomy, fermentation, isolation, physico-chemical properties and biological activity. J Antibiot (Tokyo) 43:1–7
Jiang B, Xu D, Allocco J, Parish C, Davison J, Veillette K et al (2008) PAP inhibitor with in vivo efficacy identified by Candida albicans genetic profiling of natural products. Chem Biol 15:363–374
Jiménez-Ortigosa C, Paderu P, Motyl MR, Perlin DS (2014) Enfumafungin derivative MK-3118 shows increased in vitro potency against clinical echinocandin-resistant Candida Species and Aspergillus species isolates. Antimicrob Agents Chemother 58:1248–1251
Kakeya H, Miyazaki Y, Senda H, Kobayashi T, Seki M, Izumikawa K et al (2008a) Efficacy of SPK-843, a novel polyene antifungal, in comparison with amphotericin B, liposomal amphotericin B, and micafungin against murine pulmonary aspergillosis. Antimicrob Agents Chemother 52:1868–1870
Kakeya H, Miyazaki Y, Senda H, Kobayashi T, Seki M, Izumikawa K et al (2008b) Efficacy of SPK-843, a novel polyene antifungal, in a murine model of systemic cryptococcosis. Antimicrob Agents Chemother 52:1871–1872
Kaneto R, Chiba H, Agematu H, Shibamoto N, Yoshioka T, Nishida H et al (1993) Mer-WF3010, a new member of the papulacandin family. I. Fermentation, isolation and characterization. J Antibiot (Tokyo) 46:247–250
Kantarcioglu AS, Yucel A, Vidotto V (2003) In vitro activity of a new polyene SPK-843 against Candida spp, Cryptococcus neoformans and Aspergillus spp. clinical isolates. J Chemother 15:296–298
Karababa M, Coste AT, Rognon B, Bille J, Sanglard D (2004) Comparison of gene expression profiles of Candida albicans azole-resistant clinical isolates and laboratory strains exposed to drugs inducing multidrug transporters. Antimicrob Agents Chemother 48:3064–3079
Katiyar SK, Edlind TD (2001) Identification and expression of multidrug resistance related ABC transporter genes in Candida krusei. Med Mycol 39:109–116
Kinsman OS, Chalk PA, Jackson HC, Middleton RF, Shuttleworth A, Rudd BA et al (1998) Isolation and characterisation of an antifungal antibiotic (GR135402) with protein synthesis inhibition. J Antibiot (Tokyo) 51:41–49
Kitamura A, Someya K, Hata M, Nakajima R, Takemura M (2009) Discovery of a small-molecule inhibitor of {beta}-1,6-glucan synthesis. Antimicrob Agents Chemother 53:670–677
Kohli A, Smirti, Mukhopadhyay K, Rattan A, Prasad R (2002) In vitro low-level resistance to azole in Candida albicans is associated with changes in membrane fluidity and asymmetry. Antimicrob Agents Chemother 46:1046–1052
Konishi M, Nishio M, Saitoh K, Miyaki T, Oki T, Kawaguchi H (1989) Cispentacin, a new antifungal antibiotic. I. Production, isolation, physico-chemical properties and structure. J Antibiot (Tokyo) 42:1749–1755
Kusch H, Biswas K, Schwanfelder S, Engelmann S, Rogers PD, Hecker M et al (2004) A proteomic approach to understanding the development of multidrug-resistant Candida albicans strains. Mol Genet Genomics 271:554–565
Lamb DC, Maspahy S, Kelly DE, Manning NJ, Geber A, Bennett JE et al (1999) Purification, reconstitution, and inhibition of cytochrome P-450 sterol delta22-desaturase from the pathogenic fungus Candida glabrata. Antimicrob Agents Chemother 43:1725–1728
Lamping E, Baret PV, Holmes AR, Monk BC, Goffeau A, Cannon RD (2010) Fungal PDR transporters: phylogeny, topology, motifs and function. Fungal Genet Biol 47:127–142
Lemke A, Kiderlen AF, Kayser O (2005) Amphotericin B. Appl Microbiol Biotechnol 68:151–162
Lewis RE, Kontoyiannis DP (2001) Rationale for combination antifungal therapy. Pharmacotherapy 21:149S–164S
Liu TT, Lee REB, Barker KS, Lee RE, Wei L, Homayouni R et al (2005) Genome-wide expression profiling of the response to azole, polyene, echinocandin, and pyrimidine antifungal agents in Candida albicans. Antimicrob Agents Chemother 49:2226–2236
López-Martínez R (2010) Candidosis, a new challenge. Clin Dermatol 28:178–184
Mandala SM, Thornton RA, Rosenbach M, Milligan J, Garcia-Calvo M, Bull HG et al (1997) Khafrefungin, a novel inhibitor of sphingolipid synthesis. J Biol Chem 272:32709–32714
Mandala SM, Thornton RA, Milligan J, Rosenbach M, Garcia-Calvo M, Bull HG et al (1998) Rustmicin, a potent antifungal agent, inhibits sphingolipid synthesis at inositol phosphoceramide synthase. J Biol Chem 273:14942–14949
Mansfield BE, Oltean HN, Oliver BG, Hoot SJ, Leyde SE, Hedstrom L et al (2010) Azole drugs are imported by facilitated diffusion in Candida albicans and other pathogenic fungi. PLoS Pathog 6(9):e1001126
Marichal P, Koymans L, Willemsens S, Bellens D, Verhasselt P, Luyten W et al (1999) Contribution of mutations in the cytochrome P450 14alpha-demethylase (Erg11p, Cyp51p) to azole resistance in Candida albicans. Microbiology 145:2701–2713
Martel CM, Parker JE, Bader O, Weig M, Gross U, Warrilow AG et al (2010) A clinical isolate of Candida albicans with mutations in ERG11 (encoding sterol 14alpha-demethylase) and ERG5 (encoding C22 desaturase) is cross resistant to azoles and amphotericin B. Antimicrob Agents Chemother 54:3578–3583
Miceli MH, Bernardo SM, Lee SA (2009) In vitro analyses of the combination of high-dose doxycycline and antifungal agents against Candida albicans biofilms. Int J Antimicrob Agents 34:326–332
Mitsuyama J, Nomura N, Hashimoto K, Yamada E, Nishikawa H, Kaeriyama M et al (2008) In vitro and in vivo antifungal activities of T-2307, a novel arylamidine. Antimicrob Agents Chemother 52:1318–1324
Miyazaki H, Miyazaki Y, Geber A, Parkinson T, Hitchcock C et al (1998) Fluconazole resistance associated with drug efflux and increased transcription of a drug transporter gene, PDH1, in Candida glabrata. Antimicrob Agents Chemother 42:1695–1701
Moran GP, Sanglard D, Donnelly SM, Shanley DB, Sullivan DJ, Coleman DC (1998) Identification and expression of multidrug transporters responsible for fluconazole resistance in Candida dubliniensis. Antimicrob Agents Chemother 42:1819–1830
Morio F, Loge C, Besse B, Hennequin C, Le Pape P (2010) Screening for amino acid substitutions in the Candida albicans Erg11 protein of azole-susceptible and azole-resistant clinical isolates: new substitutions and a review of the literature. Diagn Microbiol Infect Dis 66:373–384
Munro CA, Winter K, Buchan A, Henry K, Becker JM, Brown AJ et al (2001) Chs1 of Candida albicans is an essential chitin synthase required for synthesis of the septum and for cell integrity. Mol Microbiol 39:1414–1426
Nagiec MM, Nagiec EE, Baltisberger JA, Wells GB, Lester RL, Dickson RC (1997) Sphingolipid synthesis as a target for antifungal drugs. Complementation of the inositol phosphorylceramide synthase defect in a mutant strain of Saccharomyces cerevisiae by the AUR1 gene. J Biol Chem 272:9809–9817
Nishi I, Sunada A, Toyokawa M, Asari S, Iwatani Y (2009) In vitro antifungal combination effects of micafungin with fluconazole, voriconazole, amphotericin B, and flucytosine against clinical isolates of Candida species. J Infect Chemother 15:1–5
Ogawa A, Hashida-Okado T, Endo M, Yoshioka H, Tsuruo T, Takesako K et al (1998) Role of ABC transporters in aureobasidin A resistance. Antimicrob Agents Chemother 42:755–761
Okada H, Kamiya S, Shiina Y, Suwa H, Nagashima M, Nakajima S et al (1998) BE-31405, a new antifungal antibiotic produced by Penicillium minioluteum. I. Description of producing organism, fermentation, isolation, physico-chemical and biological properties. J Antibiot (Tokyo) 51:1081–1086
Olson JA, Adler-Moore JP, Smith PJ, Proffitt RT (2005) Treatment of Candida glabrata infection in immunosuppressed mice by using a combination of liposomal amphotericin B with caspofungin or micafungin. Antimicrob Agents Chemother 49:4895–4902
Pagant S, Halliday JJ, Kougentakis C, Miller EA (2010) Intragenic suppressing mutations correct the folding and intracellular traffic of misfolded mutants of Yor1p, a eukaryotic drug transporter. J Biol Chem 285:36304–36314
Pai MP, Samples ML, Mercier RC, Spilde MN (2008) Activities and ultrastructural effects of antifungal combinations against simulated Candida endocardial vegetations. Antimicrob Agents Chemother 52:2367–2376
Parish CA, Smith SK, Calati K, Zink D, Wilson K, Roemer T et al (2008) Isolation and structure elucidation of parnafungins, antifungal natural products that inhibit mRNA polyadenylation. J Am Chem Soc 130:7060–7066
Park S, Kelly R, Kahn JN, Robles J, Hsu MJ, Register E et al (2005) Specific substitutions in the echinocandin target Fks1p account for reduced susceptibility of rare laboratory and clinical Candida sp. isolates. Antimicrob Agents Chemother 49:3264–3273
Pasrija R, Banerjee D, Prasad R (2007) Structure and function analysis of CaMdr1p, a major facilitator superfamily antifungal efflux transporter protein of Candida albicans: identification of amino acid residues critical for drug/H+ transport. Eukaryot Cell 6:443–453
Perlin DS (2007) Resistance to echinocandin-class antifungal drugs. Drug Resist Updat 10:121–130
Perlin DS (2011) Current perspectives on echinocandin class drugs. Future Microbiol 6:441–457
Petraitis V, Petraitiene R, Kelaher AM, Sarafandi AA, Sein T, Mickiene D et al (2004) Efficacy of PLD-118, a novel inhibitor of candida isoleucyl-tRNA synthetase, against experimental oropharyngeal and esophageal candidiasis caused by fluconazole-resistant C. albicans. Antimicrob Agents Chemother 48:3959–3967
Prasad R, Goffeau A (2012) Yeast ATP-binding cassette transporters conferring multidrug resistance. Annu Rev Microbiol 66:39–63
Prasad R, Kapoor K (2005) Multidrug resistance in yeast Candida. Int Rev Cytol 242:215–248
Prasad R, DeWergifosse P, Goffeau A, Balzi E (1995) Molecular cloning and characterization of a novel gene of Candida albicans, CDR1, conferring multiple resistance to drugs and antifungals. Curr Genet 27:320–329
Prasad T, Hameed S, Manoharlal R, Biswas S, Mukhopadhyay CK, Goswami SK et al (2010) Morphogenic regulator EFG1 affects the drug susceptibilities of pathogenic Candida albicans. FEMS Yeast Res 10:587–596
Rodrigues ME, Silva S, Azeredo J, Henriques M (2014) Novel strategies to fight Candida species infection. Crit Rev Microbiol 10:1–13
Roemer T, Krysan DJ (2014) Antifungal drug development: challenges, unmet clinical needs, and new approaches. Cold Spring Harb Perspect Med 4 pii:a019703
Saier MH Jr, Beatty JT, Goffeau A, Harley KT, Heijne WHM, Huang SC et al (1999) The major facilitator superfamily. J Mol Microbiol Biotechnol 1:257–279
Sanglard D, Ischer F, Calabrese D, Majcherczyk PA, Bille J (1999) The ATP binding cassette transporter gene CgCDR1 from Candida glabrata is involved in the resistance of clinical isolates to azole antifungal agents. Antimicrob Agents Chemother 43:2753–2765
Sanglard D, Ischer F, Parkinson T, Falconer D, Bille J (2003) Candida albicans mutations in the ergosterol biosynthetic pathway and resistance to several antifungal agents. Antimicrob Agents Chemother 47:2404–2412
Sanglard D, Coste A, Ferrari S (2009) Antifungal drug resistance mechanisms in fungal pathogens from the perspective of transcriptional gene regulation. FEMS Yeast Res 9:1029–1050
Sanguinetti M, Posteraro B, La Sorda M, Torelli R, Fiori B, Santangelo R et al (2006) Role of AFR1, an ABC transporter-encoding gene, in the in vivo response to fluconazole and virulence of Cryptococcus neoformans. Infect Immun 74:1352–1359
Sasse C, Schillig R, Reimund A, Merk J, Morschhäuser J (2012) Inducible and constitutive activation of two polymorphic promoter alleles of the Candida albicans multidrug efflux pump MDR1. Antimicrob Agents Chemother 56:4490–4494
Serena C, Fernández-Torres B, Pastor FJ, Trilles L, Lazéra Mdos S, Nolard N et al (2005) In vitro interactions of micafungin with other antifungal drugs against clinical isolates of four species of Cryptococcus. Antimicrob Agents Chemother 49:2994–2996
Shapiro RS, Robbins N, Cowen LE (2011) Regulatory circuitry governing fungal development, drug resistance, and disease. Microbiol Mol Biol Rev 75:213–267
Sharma M, Manoharlal R, Shukla S, Puri N, Prasad T, Ambudkar SV et al (2009) Curcumin modulates efflux mediated by yeast ABC multidrug transporters and is synergistic with antifungals. Antimicrob Agents Chemother 53:3256–3265
Sharma M, Manoharlal R, Puri N, Prasad R (2010) Antifungal curcumin induces reactive oxygen species and triggers an early apoptosis but prevents hyphae development by targeting the global repressor TUP1 in Candida albicans. Biosci Rep 30:391–404
Shi W, Chen Z, Chen X, Cao L, Liu P, Sun S (2010) The combination of minocycline and fluconazole causes synergistic growth inhibition against Candida albicans: an in vitro interaction of antifungal and antibacterial agents. FEMS Yeast Res 10:885–893
Shibata T, Takahashi T, Yamada E, Kimura A, Nishikawa H, Hayakawa H et al (2012) T-2307 causes collapse of mitochondrial membrane potential in yeast. Antimicrob Agents Chemother 56:5892–5897
St Georgiev V (2000) Membrane transporters and antifungal drug resistance. Curr Drug Targets 1:261–284
Stephanie SP, Paulsen IT, Saier MH Jr (1998) Major facilitator superfamily. Microbiol Mol Biol Rev 62:1–34
Thornewell SJ, Peery RB, Skatrud PL (1997) Cloning and characterization of CneMDR1: a Cryptococcus neoformans gene encoding a protein related to multidrug resistance proteins. Gene 201:21–29
Tobudic S, Kratzer C, Lassnigg A, Graninger W, Presterl E (2010a) In vitro activity of antifungal combinations against Candida albicans biofilms. J Antimicrob Chemother 65:271–274
Tobudic S, Lassnigg A, Kratzer C, Graninger W, Presterl E (2010b) Antifungal activity of amphotericin B, caspofungin and posaconazole on Candida albicans biofilms in intermediate and mature development phases. Mycoses 53:208–214
Torelli R, Posteraro B, Ferrari S, La Sorda M, Fadda G, Sanglard D et al (2008) The ATP-binding cassette transporter-encoding gene CgSNQ2 is contributing to the CgPDR1-dependent azole resistance of Candida glabrata. Mol Microbiol 68:186–201
Uppuluri P, Nett J, Heitman J, Andes D (2008) Synergistic effect of calcineurin inhibitors and fluconazole against Candida albicans biofilms. Antimicrob Agents Chemother 52:1127–1132
Vanden Bossche H, Marichal P, Le Jeune L, Coene MC, Gorrens J, Cools W (1993) Effects of itraconazole on cytochrome P-450-dependent sterol 14 alpha-demethylation and reduction of 3-ketosteroids in Cryptococcus neoformans. Antimicrob Agents Chemother 37:2101–2105
Vandeputte P, Tronchin G, Berge`s T, Hennequin C, Chabasse D, Bouchara JP (2007) Reduced susceptibility to polyenes associated with a missense mutation in the ERG6 gene in a clinical isolate of Candida glabrata with pseudohyphal growth. Antimicrob Agents Chemother 51:982–990
Vandeputte P, Ferrari S, Coste AT (2012) Antifungal resistance and new strategies to control fungal infections. Int J Microbiol 2012:713687
Verrier PJ, Bird D, Burla B, Dassa E, Forestier C, Geisler M et al (2008) Plant ABC proteins—a unified nomenclature and updated inventory. Trends Plant Sci 13:151–159
Wang H, Kong F, Sorrell TC, Wang B, McNicholas P, Pantarat N et al (2009) Rapid detection of ERG11 gene mutations in clinical Candida albicans isolates with reduced susceptibility to fluconazole by rolling circle amplification and DNA sequencing. BMC Microbiol 14:167
White TC, Silver PM (2005) Regulation of sterol metabolism in Candida albicans by the UPC2 gene. Biochem Soc Trans 33:1215–1218
White TC, Marr KA, Bowden RA (1998) Clinical, cellular, and molecular factors that contribute to antifungal drug resistance. Clin Microbiol Rev 11:382–402
White TC, Holleman S, Dy F, Mirels LF, Stevens DA (2002) Resistance mechanisms in clinical isolates of Candida albicans. Antimicrob Agents Chemother 46:1704–1713
Wiederhold NP, Najvar LK, Fothergill AW, Bocanegra R, Olivo M, McCarthy DI et al (2015) The Novel Arylamidine T-2307 Maintains In Vitro and In Vivo Activity against Echinocandin-Resistant Candida albicans. Antimicrob Agents Chemother 59:1341–1343
Young LY, Hull CM, Heitman J (2003) Disruption of ergosterol biosynthesis confers resistance to amphotericin B in Candida lusitaniae. Antimicrob Agents Chemother 47:2717–2724
Zhang YQ, Gamarra S, Garcia-Effron G, Park S, Perlin DS, Rao R (2010) Requirement for ergosterol in V-ATPase function underlies antifungal activity of azole drugs. PLoS Pathog 6:e1000939
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Prasad, R., Shah, A.H., Rawal, M.K. (2016). Antifungals: Mechanism of Action and Drug Resistance. In: Ramos, J., Sychrová, H., Kschischo, M. (eds) Yeast Membrane Transport. Advances in Experimental Medicine and Biology, vol 892. Springer, Cham. https://doi.org/10.1007/978-3-319-25304-6_14
Download citation
DOI: https://doi.org/10.1007/978-3-319-25304-6_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-25302-2
Online ISBN: 978-3-319-25304-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)