Abstract
Lung cancer is the leading cause of cancer-related deaths in the United States and the 5-year overall survival outlook for a patient has not improved in several decades. Recently, however, molecular and genomic profiling of the lung tumors has revealed recurring somatic mutations. As a result the therapeutic landscape of lung cancer is undergoing a paradigm shift from a purely histology-based understanding of the disease to subtype distinctions based on tumor genetics, which has launched cancer-specific, mechanism-based targeted therapies with clear benefit to patients. While targeted therapy advancements are being made at an ever increasing rate, a new challenge in the form of drug resistance has also emerged. This review summarizes the current literature for these issues.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Lung cancer
- Targeted therapy
- Genomics
- Tyrosine-kinase inhibitors
- Molecular biomarkers
- Fusion genes
- Drug resistance
- EGFR
- ALK
1 Oncogenes, Tumor Suppressors and Targeted Therapeutics
Carcinogenesis and the course of the disease for each patient are influenced by many factors including ancestral genetics or germ-line polymorphisms and behavioral or life-style issues. But ultimately cancer is a disease dictated by somatic mutations . Decades of research has contributed to the understanding that cancer initiation and progression are governed by the activation of cancer driver genes, termed oncogenes, and inactivation of key tumor suppressor genes. The importance of oncogenes is underscored by the progress made in developing molecularly targeted drugs to block the function of oncogenes, often proteins with kinase function such as the epidermal growth factor receptors EGFR and HER2 [1].
There is a fundamental distinction for activating mutations arising in oncogenes compared to other mutations that are termed passenger mutations. If the mutations confer a selective growth advantage to the cancer cells they are considered to be driver oncogene mutations. Molecularly targeted therapy exploits tumor dependence on activation of driver oncogenes. Although tumor suppressors are not as directly amenable to targeted therapy, other therapeutic avenues are being explored. There are occasions where tumor suppressor gene inactivation by mutation results in the activation of kinases downstream the signaling pathway [2]. For example, the inactivation of the tumor suppressor gene PTEN activates the AKT kinase thus giving the hope in targeted therapy [3, 4]. Also, an active line of research explores mechanisms and drugs with potential to re-activate tumor suppressor pathways [5, 6].
Recently, the therapeutic landscape of Non-Small Cell Lung Cancer (NSCLC) underwent a paradigm shift from a purely histology based approach to a treatment of molecular subtypes driven by distinct genetic alterations. The evolution of this new direction started with the discovery that gain-of-function somatic mutations in epidermal growth factor receptor (EGFR) recurring in NSCLC are sensitive to EGFR tyrosine kinase inhibitors like gefitinib [7–9]. This led to the finding of a number of other genes with driver mutations in lung cancer such as HER2, KRAS, BRAF, NRAS, PIK3CA, AKT [10]. In 2007, Soda et al. discovered an EML4-ALK fusion gene, a product of a chromosomal rearrangement and a transforming agent of NSCLC [11]. After this pioneering work, the list of tumorigenic fusion genes in lung cancer is ever increasing. Therefore, this cause and effect based genomic research now links specific oncogenes and recurring mutations to the disease and provides rationale for biomarkers and molecularly targeted treatments that are improving lung cancer patient outcomes [12].
With in-depth mechanistic understanding it is appreciated that there are various ways to aberrantly or constitutively activate a proto-oncogene, but fundamentally and chief among them are genomic aberrations in the form of somatic coding mutations, copy number changes and genomic rearrangements leading to tumorigenic gene fusions. It is often straightforward to appreciate how genome aberrations in the form of coding mutations or somatic copy number alterations (SCNAs) contribute to cancer. DNA coding mutations can activate an oncogene directly or disrupt protein regulatory domains, and disruption of tumor suppressor function occurs when mutations translate to missense or truncated protein sequence. In SCNA , the operative effect of DNA imbalance disrupts the gene expression levels and thus the proper availability of the protein to function normally.
Knowledge of lung cancer molecular biology and mutation drivers has rapidly increased in recent years largely due to advancements made in high-throughput technologies that allow for genomic and transcriptomic-scale analyses . This review captures the present state of molecular genomics research of lung cancer. The diversity of oncogenic somatic mutations in lung cancer subtypes, the heightened status of fusion genes in lung cancer, and how the information is translating to the clinic are major topics.
2 Somatic Copy Number Alterations and Coding Mutation Frequencies According to Lung Cancer Subtype
Lung cancers are a heterogeneous group of tumors that are traditionally categorized by histology. By far the majority of lung cancers are categorized as non-small cell lung cancer (NSCLC) and about 15 % minority are small cell lung cancer. NSCLCs are further subdivided into adenocarcinomas (~45 %), squamous cell lung cancer (~23 %), and large cell lung cancer (~3 %), with other subtypes representing the remaining approximate 28 % [13]. During the last decade there has been a shift in classification of lung cancer based on tumor genetics. This attempt not only provided the actionable targets for the effective therapy but highlighted the importance of reconsidering the tumor reclassification, from histology based to molecular based. For example, in a recent genomic study of lung cancer classification, existence of the large cell lung cancer subtype was brought into question when these specimens were discovered to fit with adenocarcinomas or squamous cell lung cancer [12]. Moreover, the genomics approach used in this study recognized adenocarcinoma and squamous cell lung cancer cases that were not classifiable by histology.
2.1 Lung Adenocarcinoma
Of the three major subtypes of lung cancers, patients with adenocarcinoma benefit the most from molecular genomic based cancer therapeutics today. While 25–30 % of patients receive targeted therapies like gefitinib and erlotinib , another 25–30 % can enroll in clinical trials targeting other known oncogenic drivers [14]. The major oncogenic drivers in lung adenocarcinoma include activating mutations in EGFR, KRAS, BRAF, HER2, MET or translocations of ALK, ROS1 and RET [15, 16]; and all of these targets have drugs that are approved or in clinical trials. Tumor suppressor loss-of-function mutations occurring in lung adenocarcinoma include TP53, CDKN2A, PTEN, STK11, RB1, NF1, KEAP1 and SMARCA4 [16–18]. Targeting these tumor suppressor alterations is therapeutically challenging at the moment [16], but their presence may be highly informative such as in the case of TP53 mutation association with lack of response to EGFR inhibitors and recurrence [12, 19]. The key genes that are targets for treatment or that hold therapeutic potential for adenocarcinoma are discussed in greater detail below; focusing on EGFR as the model gene for lung cancer targeted therapy.
2.1.1 EGFR
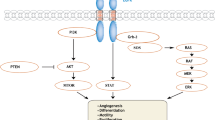
The epidermal growth factor receptor is a transmembrane tyrosine kinase that has an extracellular ligand-binding domain and an intracellular tyrosine kinase domain (Fig. 1a). EGFR belongs to the ErbB/HER family of growth factors and these proteins play a pivotal role in cell proliferation, adhesion, migration and invasion [20]. When the reversible tyrosine kinase inhibitors erlotinib and gefitinib were first used in early clinical trials in an unselected patient cohort, it showed only a modest efficacy (a response rate of about 10 %) over placebo [21]. Then in 2004 an underlying connection between EGFR activating mutations and improved lung cancer response to tyrosine kinase inhibitors laid the groundwork for molecularly informed targeted therapy [7–9]. In lung cancer, the key EGFR mutations occur in exons 18 through 21 and alter the ATP binding pocket of the kinase domain. Most mutations detected are exon 19 deletions of which there are over 20 variants (most common delE746-A750). The next most common are missense mutations in exon 21—the most frequent point mutation is the L858R (Fig. 1b). Biochemical studies later showed how these EGFR mutants preferentially bind the tyrosine kinase inhibitors like erlotinib or gefitinib over ATP [22, 23]. Therefore, these mutations are the cause of ligand-independent activation of the EGFR signaling and confer sensitivity to tyrosine kinase inhibitors. A range of less frequent in-frame insertions and duplication mutations in exon 20 have also been reported [24, 25]. As research continues, less frequent novel EGFR mutations with biologically plausible activating function are likely to be discovered. For example, a recent high-throughput whole genome and exome sequencing study using 183 lung adenocarcinoma and matched normal pairs detected two novel exon 25 and 26 deletions truncating C-terminus of the EGFR [16].
Oncogenic EGFR, targeted therapy and drug resistance. (a) The EGFR proto-oncogene encodes a tansmembrane protein (EGFR) containing an extracellular ligand-binding domain and an intracellular component with a catalytic tyrosine kinase domain. Under normal physiology, binding of a ligand (e.g. EGF) causes homodimerization of EGFR or heterodimerization with other ERBB family members to activate kinase function and induce phosphorylation. (b) With the transversion (T > G) point mutation at nucleotide position 2573, EGFR becomes oncogenic and this genetic change substitutes an arginine (R) for leucine (L) at codon 858 in exon 21. This L858R amino acid change leads to ligand-independent, constitutive activation of EGFR signaling. While this alteration disrupts the autoinhibitory interactions it also sensitizes the protein to inhibition by tyrosine kinase inhibitors like erlotinib and gefitinib. (c) More than half of the patients acquire resistance to reversible tyrosine kinase inhibitors erlotinib or gefitinib through a second mutation at T790M. This threonine to methionine amino acid change markedly decreased drug binding affinity. Afatinib is an irreversible ERBB family blocker shown to inhibit the effects of T790M mutation
Having understood the association between gain-of-function mutations of EGFR and sensitivity to EGFR tyrosine kinase inhibitors, studies demonstrated the superiority of EGFR tyrosine kinase inhibitors over chemotherapy in terms of progression-free survival, response and quality of life [26, 27]. Currently, gefitinib has been approved in Europe to treat NSCLC harboring EGFR mutations. Erlotinib was approved by the United States Food and Drug Administration (FDA) for the first line treatment of NSCLC with detected sensitizing mutations.
Although patients with EGFR mutations respond to tyrosine kinase inhibitor drugs initially, all eventually develop resistance due to the secondary mutations or other mechanisms. The most common secondary point mutation is the EGFR T790M activating mutation in exon 20 (Fig. 1c). This amino acid substitution introduces a bulky methionine at the wild-type threonine [28]. Presumably this gatekeeper mutation alters the ATP binding pocket of EGFR to reduce inhibitor binding capacity and increase affinity for ATP [23]. The second-generation irreversible EGFR tyrosine kinase inhibitor, afatinib was recently given FDA approval as a first-line therapy. When gefitinib, erlotinib or afatinib are administered as the first-line therapy for the patients with sensitization EGFR mutations, 60–80 % of the patients responded with median progression-free survival of 9–12 months and median survival in excess of 2 years [27].
Finally, it is of note that EGFR alterations are primarily in adenocarcinoma subtype and present in approximately 10 % of patients of European or African descent [29–31] though there is some dispute in the literature [32–34], while 40 % Asian patients harbor an EGFR mutation [35–37]. The majority of them are never smoker, younger, female patients [24, 36–38]. EGFR mutations are very rare in histologically pure squamous cell lung cancer [39, 40].
2.1.2 KRAS
KRAS belongs to the RAS family of proto-oncogenes and it plays a central role in downstream signal transduction induced by an array of growth factor receptors including EGFR [41]. The KRAS encoded G-protein acts as an on or off switch depending on whether the binding partner is GTP (guanosine triphosphate) or GDP (guanosine diphosphate). Mutated KRAS codes for a protein lacking GTPase activity; thus, binding of GTP locks in constitutive activation of downstream RAF/MEK/ERK and PI3K/AKT/mTOR signaling pathways [42, 43]. The most common activating mutations for KRAS include those in codon 12 and less frequently in codons 13 and 61. KRAS mutation is the most frequent oncogenic alteration in lung adenocarcinoma representing between 25 and 40 % of cases [38, 44, 45]. In general KRAS mutations do not co-occur with EGFR mutations hence it can be used as a potential negative predictive marker for the efficacy of EGFR tyrosine kinase inhibitors [24, 38]. Moreover, if KRAS is mutated it is logical that such tumors are resistant to EGFR tyrosine kinase inhibitors since KRAS acts on molecules downstream in the EGFR signaling pathway [46]. MEK1/MEK2 inhibitor selumetinib (in combination with docetaxel) was recently used in a randomized phase II study using 87 patients with advanced NSCLC having KRAS mutations [47]. In this study the combination arm, selumetinib plus docetaxel compared to placebo plus docetaxel, showed superior overall survival, though results did not reach statistical significance. Therefore, a phase III trial with a larger group of patients is needed to confirm the above results [27]. Ongoing clinical trials to inhibit KRAS mutations by targeting downstream pathways in NSCLC are studying the effects of a variety of drugs and targets, including the MEK inhibitors trametinib; tivantinib with erlotinib; or the hsp90 inhibitor IPI504 plus the mTOR inhibitor everolimus [27].
2.1.3 BRAF
The proto-oncogene BRAF encodes a serine/threonine protein kinase. This is the downstream effector protein of KRAS that activates the MAPK pathway regulating cell proliferation and survival [48]. BRAF mutations are very common in melanomas (approximately 66 % [48]) and they represent about 3 % of NSCLC [49]. Of all BRAF mutations in lung adenocarcinoma the V600E codon mutation accounts for 50 % [50]. V600E is within exon 15 and is an activating point mutation resulting in increased kinase activity, while most other BRAF codon mutations identified in lung adenocarcinoma, including G469A in exon 11 and D594G in exon 15, show low or intermediate kinase activity [48–50]. A recent case report showed the clinical benefit of the drug vemurafenib in treating a NSCLC patient with a tumor V600E mutation [51]. The on-going clinical trials targeting either BRAF or its downstream effectors are studying outcomes for BRAF inhibitor dabrafenib on NSCLC patients with BRAF V600E mutation, the MEK inhibitor, trametinib for patients with non-V600E mutations and the drug dasatinib for the NSCLC patients with uncharacterized BRAF mutations [27].
2.1.4 HER2
Like EGFR, HER2 is also a member of the ErbB family of epidermal growth factor receptor tyrosine kinases. HER2 is activated in 25–30 % of breast cancers due to focal amplification of the chromosome region 17q12 comprising the HER2 gene. The contribution of HER2 amplification in lung adenocarcinoma has been estimated to be 35 % based on immunohistochemistry studies [52]. Although not found in breast cancer, HER2 is also observed to be activated in approximately 2 % of lung adenocarcinomas due to an in-frame insertion [53]. These activating mutations occur in exon 20 as in-frame insertions of 3 to 12 base pairs [54]. A clinical trial investigating outcomes of monoclonal antibody trastuzumab targeting HER2 overexpression in NSCLC showed no benefit alone [55] or in combination with chemotherapy [56]. However, individual clinical case reports support the potential for patients with HER2 amplification in lung cancer [57]. Moreover, studies of HER2 binding tyrosine kinase inhibitors including afatinib [58], dacomitinib and neratinib [59] have yielded promising preliminary results against HER2 mutants in NSCLC.
2.1.5 MET
The proto-oncogene MET codes for the transmembrane receptor tyrosine kinase also known as hepatocyte growth factor receptor. The binding of hepatocyte growth factor (HGF ligand) to the MET receptor activates the downstream RAS/RAF/MEK/MAPK; PI3K/AKT and c-SRC kinase pathways [60]. Mutations in MET are rare and it is most often gene copy number increase that leads to overexpression of the MET protein [61, 62]. A key observation for this mutation is that the amplification of MET gene is associated with developing secondary resistance to EGFR tyrosine kinase inhibitors. Evidence suggests that 5 % of the patients with EGFR mutations who initially responded to gefitinib or erlotinib acquire resistance due to MET amplification [63–65]. Here, the increased MET kinase activity drives the PI3K/AKT pathway bypassing the EGFR-directed tyrosine kinase inhibition [65]. The findings indicate the importance of blocking both EGFR and MET as a means of treating patients with acquired resistance. A recent randomized double blind phase II study investigating the effect of the MET receptor-targeted monoclonal antibody onartuzumab plus erlotinib compared to placebo plus erlotinib showed significant improvements in clinical outcomes with respect to progression-free survival and overall survival [66]. Moreover, this study illustrated the importance of parallel diagnostic testing after seeing worse outcomes with MET amplification negative patients. Therefore, the MET immunohistochemisty assay developed in the phase II study was incorporated as a diagnostic test for use of onartuzumab in the randomized phase III trial investigating the effect of onartuzumab and erlotinib [67]. A number of MET inhibitors and neutralizing antibodies are drugs presently in development. Some of the examples are MET inhibitor cabozantinib [68], MET tyrosine kinase inhibitor crizotinib [69], and hepatocyte growth factor neutralizing antibody rilotumumab [70, 71]. It has been noted that MET amplifications and KRAS mutations are mutually exclusive, meaning they are not co-expressed in lung cancer specimens [72].
2.2 Squamous Cell Lung Cancer
Of the major subtypes of lung cancers, squamous cell lung cancer shows the strongest association with cigarette smoking [73]. Furthermore unlike lung adenocarcinoma, presently there are no targeted therapies used in treatment of squamous cell lung cancer patients. Past trials to treat squamous cell lung cancer with chemotherapy and EGFR tyrosine kinase inhibitors showed the ineffectiveness of such treatments [74, 75]. This puts increased emphasis on the need for genomic analyses to find potential oncogenes that may present druggable targets for this cancer subtype. The earliest genomic aberrations found in squamous cell lung cancer included allelic loses at chromosome 3p (3p21, 3p22–24, 3p25), 8p21–23, 9p21 [76]; followed by loses at 17p13 comprising the TP53 tumor suppressor gene and 13q14 containing tumor suppressor RB1 [77]. Using whole-exome sequencing to identify new somatic mutations in this lung cancer subtype, Zheng et al. reported TP53, EP300, LPHN2, C10orf137, MYH2, TGM2 and MS4A3 as mutated genes with oncogenic potential [78]. Comprehensive analyses by The Cancer Genome Atlas (TCGA) shed more light on squamous cell lung cancer in 2012. The project used 178 histopathologically reviewed samples to detect on average 323 SCNAs, 360 exonic mutations and 165 genomic rearrangements per tumor [3]. The study identified statistically significant, recurring mutations in 11 genes, including TP53 mutations in nearly all the specimens. The mutation frequencies of the genes in TCGA data were compatible with the study carried out by Paik et al. that examined specimens from 52 patients [79]. In this study 60 % of the patients harbored functionally relevant mutations in druggable oncogene targets including FGFR1, DDR2, PIK3CA in addition to tumor suppressor PTEN. Research has continued and the evolving knowledge on the specifics of oncogenic drivers of squamous cell lung cancer is further discussed below. Moreover, results from clinical studies are necessary to appreciate if these findings will translate to improve the overall survival of squamous cell lung cancer patients.
2.2.1 Somatic Copy Number Alterations in Squamous Cell Lung Cancer
FGFR1 (Fibroblast growth factor receptor 1 ) is a transmembrane tyrosine kinase and is one of the promising drug targets in squamous cell lung cancer. The amplification of the chromosome region 8p12 was detected in 2010 and focal amplification of FGFR1 was validated in 15 of 155 squamous cell tumors [80]. The amplification was confirmed in an independent cohort of squamous cell lung cancer samples with 22 % cases being positive by fluorescence in situ hybridization (FISH) analysis [80]. According to TCGA analysis amplification of FGFR1 is observed in 7 % of squamous cell lung cancer [3]. Clinical trials employing small molecule inhibitors that block FGFR1 are on-going, these include molecules specific to FGFR1 kinase, multi-kinase inhibitors and pan-FGFR inhibitors [80–82]. FGFR1 amplification and MET amplification frequency (reported at about 6 % in lung squamous cell lung cancer) are both considered to be more prevalent in lung squamous cell lung cancer than in adenocarcinoma [83].
SOX2 is a transcription factor that regulates pluripotency of embryonic stem cells as well as morphogenesis of trachea-bronchial epithelia [73]. This lineage-survival oncogene was discovered using comparative genomic hybridization with probes targeting the 3q26 region [84]. About 60–80 % of squamous cell lung cancers show amplifications in this region of chromosome 3 and approximately 20 % harbor a focal amplification that includes the SOX2 gene [85, 86]. According to the TCGA study, SOX2 was amplified in 21 % of the samples analyzed [3]. Although it was demonstrated that the inhibition of SOX2 suppresses cancer cell growth, research also suggests that SOX2 amplification is not sufficient for carcinogenesis in the absence of other oncogenic mutations [84].
At a lower frequency than those estimated above, PDGFRA (platelet-derived growth factor receptor) tyrosine kinase, located in chromosomal region 4q12, is shown to be amplified in 4–8 % of squamous cell lung cancers [3, 87]. There are a number of multi-targeted tyrosine kinase inhibitors against PDGFRA that are in clinical development at this time; including sunitinib, pazopanib, cediranib and nintedanib [88]. HER2 amplifications are also observed in about 4 % of squamous cell lung cancers [89]; evaluation of HER2-directed therapy needs to be done.
2.2.2 Somatic Coding Mutations in Squamous Cell Lung Cancer
Well-documented oncogene mutations recurring at significant frequency in squamous cell lung cancer include the AKT1 codon E17K somatic mutation, which causes constitutive activation of the kinase [90]. Malanga et al. found this mutation in a subset of squamous cell lung cancer (2/36 lung squamous cell lung cancer and 0/53 lung adenocarcinoma) [91]. AKT kinase inhibitors such as MK2206 and GDC-0068 are in clinical trials [92]. BRAF mutations are present in about 4 % of squamous cell lung cancers [50]. A clinical trial is underway to test BRAF-specific kinase inhibitor GSK2118436 on patients with squamous cell lung cancer with BRAF mutations [82]; and other existing data point to MEK inhibition as potentially effective target for non-V600E BRAF mutations in this lung cancer subcategory [93]. DDR2 (Discoidin domain receptor 2) tyrosine kinase is described as an oncogene that promotes cell proliferation and cell survival [94], and mutations in the DDR2 gene render cells sensitive to the small molecule kinase inhibitor dasatinib [95]. A clinical trial is underway to find out the efficacy of dasatinib on the squamous cell lung cancer with activating DDR2 mutations, which are observed at a rate of close to 4 %.
PIK3CA is one of the most common sequence mutated oncogenes in cancer and it is reported to present more frequently in squamous cell lung cancer than in lung adenocarcinoma [96]. In accordance with previous studies missense mutations at codon positions 545 and 1047 were found in 48 % of the samples in the TCGA study [3, 97]. PIK3CA encodes the catalytic subunit of the PI3K lipid kinases and a number of clinical trials are presently underway to examine the impact of targeted therapies and combination PI3K inhibitors and chemotherapy in lung cancer [98]. The PI3K inhibitors in clinical development include XL-147, XL-765, BEZ235, BKM120, GDC-0941, early evidence indicates the response rate to these single agents are low [10, 98, 99].
Other genes reported to show recurring mutations in squamous cell lung cancer include the MLL2 gene encoding a histone methyltranferase that plays a key role in epigenetic programming and embryonic development. The therapeutic strategies to target epigenetic pathways; for example histone methyltransferase inhibitors are also emerging and mutation activated MLL2 holds promise as a novel target [100, 101]. PTEN is a tumor suppressor gene often sequence mutated and inactivated in many types of cancer. The mutation frequency of PTEN reported at 15 % in squamous cell lung cancer is higher than compared to lung adenocarcinoma [3, 102]. Also, how loss of function mutations in the HLA_A class I Major Histocompatibility (MHC I) gene may help cancer cells avoid immune responses as has been proposed and raises the promise of immunotherapy [103, 104].
2.3 Small Cell Lung Cancer
Small cell lung cancer is the third most frequent subtype of lung cancer diagnosis representing around 200,000 cases worldwide annually. According to overall survival rates, patients with small cell lung cancer by far face the lowest probability of survival [105]. The 5-year overall survival outlook for these patients is about 5 % and this has not improved for the last four decades [106]. Efforts to study somatic mutations in small cell lung cancer, which is rarely treated by surgery, trail behind other histologic subtypes due to lack of specimens. However, very recent studies present the first results of comprehensive profiling of small cell lung cancer specimens. Rudin et al. characterized 80 small cell lung cancer specimens including cancer-derived cell lines and 36 primary tumors and paired normal tissue [107] A key finding was a significant SOX2 amplification frequency ~27 % and the demonstration of decreased proliferation in a small cell lung cancer cell model using shRNA knockdown of SOX2 [107]. Peifer et al., by accessing small cell lung tumor specimens from a global genome research consortium, were able to sequence 29 exomes, 2 genomes and 15 transcriptomes [108]. Their SCNA algorithm identified almost universal deletions at chromosome 3p and 13q (affecting RB1), 17p (containing TP53) and frequent gains of 3q and 5p as well as for the FGFR1 gene.
Iwakawa et al. used genome-wide copy number analysis and whole-transcriptome sequencing to study the genome-wide amplifications and translocations in small cell lung cancer [109]. Their copy number analysis found 34 genes to be frequently amplified in small cell lung cancer. Among them three MYC family genes MYCL1 (1p34.2), MYCN (2p24.3) and MYC (8q24.21) were frequently amplified in concordance with the previous small scale studies using [110–112]. This is an important finding in small cell lung cancer as inhibitors against MYC family protein products are gaining research traction [113–115]. In addition, the study identified the chromosomal region 9p24.1 as demonstrating mutual exclusivity with MYC amplifications. Furthermore, mRNA expression of the gene KIAA1432 (from the 9p24.1 region) was strongly correlated with the KIAA1432 amplification suggesting a novel cancer gene activated in small cell lung cancer. Compared to prevalence of kinase gene mutations in lung adenocarcinoma, targeting molecular markers of the small cell lung cancer (e.g. SOX2) may be therapeutically challenging. However, extensive basic and clinical research on the genomic aberrations of small cell lung cancer will enable efforts to understand and develop treatment options for this exceptionally aggressive disease. Since lack of small cell lung cancer patient specimens is a major problem, Sos et al. screened 267 compounds across 44 cell lines of this lung cancer subcategory to establish a genomic characterization framework [115]. By comparing SCNAs identified in 60 patient-derived small cell lung cancer cell lines with results from 63 primary tumor specimens described above, the authors demonstrated the comparable genomic landscape of small cell lung cancer between the two sample types. Then they showed the effectiveness of the Aurora kinase inhibitors against small cell lung cancer cell lines harboring MYC amplification.
3 Genomic Translocations and Expressed Fusion Genes
Compared to point mutations in oncogenes, a genomic translocation that gives rise to an oncogenic fusion gene can have more deleterious effects on protein function and on downstream cellular pathways (Fig. 2). Yet gene fusions are proving to be excellent cancer-specific drug targets and oncogenic tyrosine kinase gene fusions are the best examples. In 2007, Soda et al. discovered the first druggable EML4-ALK fusion protein—an oncokinase—in NSCLC [11]. The marked response of patients with ALK positive NSCLC to the small-molecule tyrosine kinase inhibitor crizotinib [116, 117] catalyzed the field to search for expression of other novel oncogenic fusion genes. Application of high-throughput RNA sequencing analysis has greatly contributed to the identification of additional fusion genes in lung cancer involving kinases: ROS1[118], RET [119], FGFR1/2/3 [120, 121], NTRK1[122], ERBB4 and BRAF [123], and AXL and PDGFRA [124]. Also, fusion genes involving the EGFR ligand NRG1 (CD74-NRG1, SLC3A2-NRG1) have been reported [123]. The particular importance of ALK, ROS1 and RET fusion genes in lung cancer is expanded on below.
CD74-ROS1 translocation and expressed fusion genes. CD74 and ROS1 genomic rearrangements (double stranded DNA) results in the mRNA expression of two different fusion variants. Left, depiction of CD74 exon 6 (red) fusion with either ROS1 exon 34 (light blue) or exon 35 (dark blue). Right, the predicted protein configuration of the two spliced forms and their plasma membrane orientation are depicted. Of the two variants only the major spliced form CD74-ROS1 exon 34, which shows an additional transmembrane domain (light blue) that positions the ROS1 tyrosine kinase domain intracellularly, is considered to be oncogenic. The original patient with lung cancer expressing this mutation initially responded to crizotinib, later the drug resistance was developed due to the amino acid substitution G2032R
3.1 ALK
The inversion on chromosome 2p leads to the formation of the most commonly expressed ALK fusion, EML4-ALK. As the genomic inversion does not occur at the same location all the time, it results in expression of a number of EML4-ALK variants [11]. In all the variants, the intracellular tyrosine kinase domain of ALK starting at exon 20 is present while the EML4 truncates at different points. The two most common variants E13:A20 (33 %) and E6a/b:A20 (29 %), which are also referred to as variant 1 and 3a/b respectively, represent approximately 60 % of detected EML4-ALK variants [125]. The NSCLC cell lines H3122 and DFC1031 contain the E13:A20 variant while H2228 harbors the E6a/b:A20 [126]. In NSCLC other ALK fusion partners have also been discovered, including TFG [118], KIF5B [127], HIP1 [128], KLC1 [129], TPR [130]. Each of these fusion partners mediates the ligand independent dimerization of ALK to constitutively activate ALK kinase function. The prevalence of the ALK rearrangements occurs in 3 to 7 % of unselected patients with NSCLC [11, 126]. This amounts to an estimated 65,000 new patients each year with ALK rearrangements [131] a number that is in the range of annual total number of Chronic Myeloid Leukemia cases [132, 133]. Like EGFR mutations, ALK rearrangements tend to occur in younger age patients with adenocarcinoma histology and never or light smoking history [117, 134]. Also, ALK rearrangements are the second genetic biomarker related to FDA-approved targeted therapy for NSCLC. Small molecule tyrosine kinase inhibitor crizotinib (originally developed for MET) was approved in 2011 along with the break apart FISH as the diagnostic test to detect the ALK positive advanced NSCLC patients [27, 125, 135]. In a recent phase 1 trial enrolling patients with ALK rearrangement positive lung cancer, the higher potency tyrosine kinase inhibitor ceritinib inhibited the resistance developed by the crizotinib treatment exemplifying the power of mechanism based rational drug design [136]. Mechanistically, the benefit of ceritinib over crizotinib is that it is uniquely effective at inhibiting secondary ALK mutation L1196M. For the first time in the history of targeted therapy, ceritinib received FDA approval just after the phase I clinical trial [137].
3.2 ROS1
The analysis of 41 cell lines and 150 NSCLC tumors led Rikova et al. to characterize the first ROS1 rearrangement in NSCLC [118]. In one of the cell lines (HCC78) the authors identified the ROS1-SLC34A2 fusion and one of the tumor samples harbored the CD74-ROS1 fusion. Follow-up studies discovered a number of ROS1 fusion gene partners: TPM3 [138], SDC4 [138, 139], EZR [140], LRIG3 [138], FIG [141], KDELR2 [142], CCDC6 [124]. ROS1 is located on human chromosome 6 and with the exception of FIG and EZR all other fusion partners are coming from different chromosomes [143]. In all the different fusion proteins, the ROS1 tyrosine kinase domain remains intact [138]. For ROS1 fusion genes the mechanism remains unknown [119], but the likely oncogenic consequence is constitutive activation of ROS1 tyrosine kinase function. Furthermore, the expression of ROS1 fusion genes both in vitro and in vivo leads to oncogenic transformation [138]. Emerging data indicates that, ROS1 fusion genes may preferentially activate downstream PI3K/AKT/mTOR and MAPK/ERK pathways [144].
3.3 RET
In 2011, researchers discovered the first RET gene fusion partnered with the gene KIF5B in NSCLC [145]. In 2012, three studies each added more variants to the list of expressed KIF5B-RET fusion genes [138, 146, 147]. Although KIF5B is the most common fusion partner of RET, other partners have also been reported such as CCDC6, TRIM33 and NCOA4 [148, 149]. The RET tyrosine kinase domain is conserved in all the fusions. In contrast to ROS1, RET fusion partners like ALK fusion partners contain a coiled-coil domain. Positioned at the 5′ end of the fusion gene this domain promotes ligand independent dimerization and hence constitutive activation of RET kinase function.
Although the prevalence of ROS1 and RET fusion genes are about 1–2 % in an unselected population of NSCLC [138, 150], there is great interest for these two fusions as novel targets due to three main reasons. First, ROS1 and RET fusions tend to occur without the presence of other driver mutations and this knowledge of mutual exclusivity can be used to strategize screening and detection [147]. Second, NSCLC patients harboring ROS1 or RET fusions show unique clinicopathologic features [138, 150] (e.g. relatively younger age, never smoker with adenocarcinoma histology) facilitating clinical enrollments [119]. Third, there are already inhibitor drugs targeting ROS1 and RET in clinical trials [148; 150]. It took only 4 years from the first identification of an ALK fusion gene in NSCLC for the FDA to conditionally approve an ALK-targeted tyrosine kinase inhibitor [135]; and in less than 6 months of publication on RET fusion genes, Drilon et al. initiated a clinical trial with cabozantinib [148]. Again underscoring how the transition from genomic research to molecularly-defined therapy in lung cancer can advance at an incredibly rapid rate.
With high-throughput sequencing of greater numbers of lung cancer transcriptomes across all histological subtypes, additional oncogenic variants of fusion genes may be discovered. However, it is important that complementary work be done to establish or refute if any one specific fusion gene event is tumorigenic and clinically actionable. For example, although the ROS1 gene fusions KDELR2-ROS1 and CCDC6-ROS1 have been discovered in NSCLC, their tumorigenic potential has not been established [151]. In another example, a genomic translocation suggested to give rise to expression of a CCDC6-RET fusion gene has been detected in two forms: CCDC6 exon 1 fused to RET exon 12 (C1; R12) and CCDC6 intron 1 fused to RET exon 11 (C1; R11). However, only CCDC6-RET (C1; R12) is expressed and contributes to malignancy while CCDC6-RET (C1; R11) represents a benign breakpoint in the genome, therefore it is of no obvious clinical importance [152].
4 Challenges and Conclusions
The hallmarks of a cancer cell, distinct from normal cell biology, include the capacity for unlimited and unmitigated proliferation; resistance to anti-proliferative and apoptotic cues; and the ability to survive and proliferate in stressful conditions [103]. Underlying these malignant phenotypes is aberrant molecular biology in the form of deregulated signaling pathways or functional networks of genes that are ultimately governed by a mutated genome [153]. Much progress has been made to develop anti-cancer drugs that target the protein products of well-studied, recurrently mutated oncogenes. And to date the greatest clinical successes for molecularly targeted treatments in lung cancer have come from efforts to target EGFR and ALK kinases. Certainly more are on the horizon that will increasingly define and include all lung cancer subtypes, as stories of rapid discovery and drug development are unfolding in the literature.
Despite targeted treatment advances and marked improvements in patient outcomes over traditional chemotherapies, targeted therapies often fail for patients due to de novo or acquired drug resistance. A few examples of de novo resistance mechanisms in lung cancer stem from the observation that nearly 30 % of patients with tumors positive for EGFR mutations show no initial response [154–158]. EGFR mutations carrying exon 20 insertions are not sensitive to EGFR-tyrosine kinase inhibitor drugs. Unlike other EGFR-activating mutations, the exon 20 insertion D770_N771insNPG promotes EGFR function without increasing affinity for EGFR tyrosine kinase inhibitors [159, 160]. In another example, the EGFR T790M mutation, which confers EGFR-targeted drug resistance when it arises in a tumor, also exists as a heterozygous germ-line variant in 0.5 % of lung adenocarcinoma patients [161, 162].
The most frequent mechanism of acquired resistance is the gain of second-site EGFR mutations, which is estimated to occur in more than 50 % of the patients; among them the T790M mutation contributes more than 90 % [163]. In EML4-ALK fusion-gene positive patients, the gatekeeper mutation L1196M, analogous to EGFR T790M, requires the contribution from additional mutations within the ALK sequence and the net effect allows it to block crizotinib from its binding site [164, 165]. More recently, a second-site mutation was discovered within the ROS1 fusion gene CD74-ROS1; it was causally linked to acquired resistance to crizotinib [166] (Fig. 2). The single G2032R amino acid change provides sufficient steric bulk to block inhibitor drug binding.
To better appreciate how acquired resistance arises, bear in mind that targeted therapies can promote minority populations of tumor cells harboring another driver oncogene, or cause reversible growth inhibition or autophagy allowing subpopulations of cancer cells the opportunity to evolve mechanisms for drug resistance leading to recurrence. Moreover, current targeted therapies inhibit the oncogene directly, and by default the proto-oncogene, thereby causing dose-limiting side effects. To overcome drug resistance , an array of new drugs including second and third generation EGFR and other tyrosine kinase inhibitors are being utilized and developed, as single and combination agents. The recent success of ceritinib in overcoming crizotinib drug resistance in ALK rearranged NSCLC is a milestone example [136].
A major challenge for research efforts to catalog the driver mutations in lung cancer is the high mutation frequency in lung cancer compared to other cancers. For example squamous cell lung carcinoma shows a median mutation frequency of 8.15 per megabase (Mb) while that of AML is only 0.28 mutations /Mb [167]. This poses the difficulty of detecting oncogenic drivers among the vast majority of passive mutations. Even the most comprehensive sequencing endeavors like the study of 183 lung adenocarcinomas raises gaps in our understanding [16]. In this study 15 % of the patients did not show a single mutation in known oncogenes or genes with known cancer function [103]. A recent saturation analysis across 21 tumor types estimated the requirement of 600–5000 samples per lung tumor type to achieve near-saturation [168]. The number of lung cancer samples necessary to detect a mutation at 3 % frequency extrapolates to about 2000 samples.
To conclude, the end-goal of research is transfer of the accumulated knowledge and evolution of knowledge of tumor biology to the clinic; here genomic technologies and cancer type-specific, single-pass comprehensive mutation panels are poised to transform clinical testing. The many complexities accompanying this paradigm shift should not be underestimated and difficulties remain for even the most forward thinking institutes, but they are foreseeably overcome by expert collaborative teams made up of health care professionals; basic and translational scientists; and regulatory agencies.
References
Sharma SV, Settleman J (2007) Oncogene addiction: setting the stage for molecularly targeted cancer therapy. Genes Dev 21:3214–3231
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr et al (2013) Cancer genome landscapes. Science 339:1546–1558
(2012) Comprehensive genomic characterization of squamous cell lung cancers. Nature 489:519–525
Vivanco I, Sawyers CL (2002) The phosphatidylinositol 3-kinase AKT pathway in human cancer. Nat Rev Cancer 2:489–501
Beltran AS, Blancafort P (2011) Reactivation of MASPIN in non-small cell lung carcinoma (NSCLC) cells by artificial transcription factors (ATFs). Epigenetics 6:224–235
Lee SH, Park BJ (2011) p53 activation by blocking Snail: a novel pharmacological strategy for cancer. Curr Pharm Des 17:610–617
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H et al (2004) EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304:1497–1500
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA et al (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350:2129–2139
Pao W, Miller V, Zakowski M, Doherty J, Politi K et al (2004) EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci U S A 101:13306–13311
Pao W, Girard N (2011) New driver mutations in non-small-cell lung cancer. Lancet Oncol 12:175–180
Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y et al (2007) Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 448:561–566
(2013) A genomics-based classification of human lung tumors. Sci Transl Med 5:209ra153
Howlander N, Noone AM, Krapcho M, Neyman N, Aminou R, Waldron W, Altekruse SF, Kosary CL, Rulhl J, Tatalovich Z, Cho H, Mariotto A, Eisner MP, Lewis DR, Chen HS, Feuer EJ, Cronin KA, Edwards BK (2011) SEER cancer statistics review, 1975-2008. National Cancer Institute, Bethesda
Pao W, Iafrate AJ, Su Z (2011) Genetically informed lung cancer medicine. J Pathol 223:230–240
Pao W, Hutchinson KE (2012) Chipping away at the lung cancer genome. Nat Med 18:349–351
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ et al (2012) Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150:1107–1120
Kan Z, Jaiswal BS, Stinson J, Janakiraman V, Bhatt D et al (2010) Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466:869–873
Sanchez-Cespedes M, Parrella P, Esteller M, Nomoto S, Trink B et al (2002) Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res 62:3659–3662
Huang S, Benavente S, Armstrong EA, Li C, Wheeler DL et al (2011) p53 modulates acquired resistance to EGFR inhibitors and radiation. Cancer Res 71:7071–7079
Prenzel N, Fischer OM, Streit S, Hart S, Ullrich A (2001) The epidermal growth factor receptor family as a central element for cellular signal transduction and diversification. Endocr Relat Cancer 8:11–31
Shepherd FA, Rodrigues Pereira J, Ciuleanu T, Tan EH, Hirsh V et al (2005) Erlotinib in previously treated non-small-cell lung cancer. N Engl J Med 353:123–132
Yun CH, Boggon TJ, Li Y, Woo MS, Greulich H et al (2007) Structures of lung cancer-derived EGFR mutants and inhibitor complexes: mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 11:217–227
Yun CH, Mengwasser KE, Toms AV, Woo MS, Greulich H et al (2008) The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc Natl Acad Sci U S A 105:2070–2075
Tam IY, Chung LP, Suen WS, Wang E, Wong MC et al (2006) Distinct epidermal growth factor receptor and KRAS mutation patterns in non-small cell lung cancer patients with different tobacco exposure and clinicopathologic features. Clin Cancer Res 12:1647–1653
Yamamoto H, Toyooka S, Mitsudomi T (2009) Impact of EGFR mutation analysis in non-small cell lung cancer. Lung Cancer 63:315–321
Hirsch FR, Janne PA, Eberhardt WE, Cappuzzo F, Thatcher N et al (2013) Epidermal growth factor receptor inhibition in lung cancer: status 2012. J Thorac Oncol 8:373–384
Cardarella S, Johnson BE (2013) The impact of genomic changes on treatment of lung cancer. Am J Respir Crit Care Med 188:770–775
Balak MN, Gong Y, Riely GJ, Somwar R, Li AR et al (2006) Novel D761Y and common secondary T790M mutations in epidermal growth factor receptor-mutant lung adenocarcinomas with acquired resistance to kinase inhibitors. Clin Cancer Res 12:6494–6501
Shigematsu H, Lin L, Takahashi T, Nomura M, Suzuki M et al (2005) Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J Natl Cancer Inst 97:339–346
Eberhard DA, Giaccone G, Johnson BE (2008) Biomarkers of response to epidermal growth factor receptor inhibitors in Non-Small-Cell Lung Cancer Working Group: standardization for use in the clinical trial setting. J Clin Oncol 26:983–994
Russell PA, Barnett SA, Walkiewicz M, Wainer Z, Conron M et al (2013) Correlation of mutation status and survival with predominant histologic subtype according to the new IASLC/ATS/ERS lung adenocarcinoma classification in stage III (N2) patients. J Thorac Oncol 8:461–468
Bauml J, Mick R, Zhang Y, Watt CD, Vachani A et al (2013) Frequency of EGFR and KRAS mutations in patients with non small cell lung cancer by racial background: do disparities exist? Lung Cancer 81:347–353
Yang SH, Mechanic LE, Yang P, Landi MT, Bowman ED et al (2005) Mutations in the tyrosine kinase domain of the epidermal growth factor receptor in non-small cell lung cancer. Clin Cancer Res 11:2106–2110
Reinersman JM, Johnson ML, Riely GJ, Chitale DA, Nicastri AD et al (2011) Frequency of EGFR and KRAS mutations in lung adenocarcinomas in African Americans. J Thorac Oncol 6:28–31
Sos ML, Koker M, Weir BA, Heynck S, Rabinovsky R et al (2009) PTEN loss contributes to erlotinib resistance in EGFR-mutant lung cancer by activation of Akt and EGFR. Cancer Res 69:3256–3261
Kosaka T, Yatabe Y, Endoh H, Kuwano H, Takahashi T et al (2004) Mutations of the epidermal growth factor receptor gene in lung cancer: biological and clinical implications. Cancer Res 64:8919–8923
Tokumo M, Toyooka S, Kiura K, Shigematsu H, Tomii K et al (2005) The relationship between epidermal growth factor receptor mutations and clinicopathologic features in non-small cell lung cancers. Clin Cancer Res 11:1167–1173
Ding L, Getz G, Wheeler DA, Mardis ER, McLellan MD et al (2008) Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455:1069–1075
Rekhtman N, Paik PK, Arcila ME, Tafe LJ, Oxnard GR et al (2012) Clarifying the spectrum of driver oncogene mutations in biomarker-verified squamous carcinoma of lung: lack of EGFR/KRAS and presence of PIK3CA/AKT1 mutations. Clin Cancer Res 18:1167–1176
Ohtsuka K, Ohnishi H, Fujiwara M, Kishino T, Matsushima S et al (2007) Abnormalities of epidermal growth factor receptor in lung squamous-cell carcinomas, adenosquamous carcinomas, and large-cell carcinomas: tyrosine kinase domain mutations are not rare in tumors with an adenocarcinoma component. Cancer 109:741–750
Karnoub AE, Weinberg RA (2008) Ras oncogenes: split personalities. Nat Rev Mol Cell Biol 9:517–531
Downward J (2003) Targeting RAS signalling pathways in cancer therapy. Nat Rev Cancer 3:11–22
Rodenhuis S, Slebos RJ (1992) Clinical significance of ras oncogene activation in human lung cancer. Cancer Res 52:2665s–2669s
Sequist LV, Heist RS, Shaw AT, Fidias P, Rosovsky R et al (2011) Implementing multiplexed genotyping of non-small-cell lung cancers into routine clinical practice. Ann Oncol 22:2616–2624
Riely GJ, Kris MG, Rosenbaum D, Marks J, Li A et al (2008) Frequency and distinctive spectrum of KRAS mutations in never smokers with lung adenocarcinoma. Clin Cancer Res 14:5731–5734
Linardou H, Dahabreh IJ, Kanaloupiti D, Siannis F, Bafaloukos D et al (2008) Assessment of somatic k-RAS mutations as a mechanism associated with resistance to EGFR-targeted agents: a systematic review and meta-analysis of studies in advanced non-small-cell lung cancer and metastatic colorectal cancer. Lancet Oncol 9:962–972
Janne PA, Shaw AT, Pereira JR, Jeannin G, Vansteenkiste J et al (2013) Selumetinib plus docetaxel for KRAS-mutant advanced non-small-cell lung cancer: a randomised, multicentre, placebo-controlled, phase 2 study. Lancet Oncol 14:38–47
Davies H, Bignell GR, Cox C, Stephens P, Edkins S et al (2002) Mutations of the BRAF gene in human cancer. Nature 417:949–954
Brose MS, Volpe P, Feldman M, Kumar M, Rishi I et al (2002) BRAF and RAS mutations in human lung cancer and melanoma. Cancer Res 62:6997–7000
Paik PK, Arcila ME, Fara M, Sima CS, Miller VA et al (2011) Clinical characteristics of patients with lung adenocarcinomas harboring BRAF mutations. J Clin Oncol 29:2046–2051
Gautschi O, Pauli C, Strobel K, Hirschmann A, Printzen G et al (2012) A patient with BRAF V600E lung adenocarcinoma responding to vemurafenib. J Thorac Oncol 7:e23–e24
Hirsch FR, Varella-Garcia M, Franklin WA, Veve R, Chen L et al (2002) Evaluation of HER-2/neu gene amplification and protein expression in non-small cell lung carcinomas. Br J Cancer 86:1449–1456
Shigematsu H, Takahashi T, Nomura M, Majmudar K, Suzuki M et al (2005) Somatic mutations of the HER2 kinase domain in lung adenocarcinomas. Cancer Res 65:1642–1646
Stephens P, Hunter C, Bignell G, Edkins S, Davies H et al (2004) Lung cancer: intragenic ERBB2 kinase mutations in tumours. Nature 431:525–526
Clamon G, Herndon J, Kern J, Govindan R, Garst J et al (2005) Lack of trastuzumab activity in nonsmall cell lung carcinoma with overexpression of erb-B2: 39810: a phase II trial of Cancer and Leukemia Group B. Cancer 103:1670–1675
Gatzemeier U, Groth G, Butts C, Van Zandwijk N, Shepherd F et al (2004) Randomized phase II trial of gemcitabine-cisplatin with or without trastuzumab in HER2-positive non-small-cell lung cancer. Ann Oncol 15:19–27
Kelly RJ, Carter CA, Giaccone G (2012) HER2 mutations in non-small-cell lung cancer can be continually targeted. J Clin Oncol 30:3318–3319
De Greve J, Teugels E, Geers C, Decoster L, Galdermans D et al (2012) Clinical activity of afatinib (BIBW 2992) in patients with lung adenocarcinoma with mutations in the kinase domain of HER2/neu. Lung Cancer 76:123–127
Gandhi L, Camidge DR, Ribeiro de Oliveira M, Bonomi P, Gandara D et al (2011) Phase I study of Navitoclax (ABT-263), a novel Bcl-2 family inhibitor, in patients with small-cell lung cancer and other solid tumors. J Clin Oncol 29:909–916
Sadiq AA, Salgia R (2013) MET as a possible target for non-small-cell lung cancer. J Clin Oncol 31:1089–1096
Cappuzzo F, Marchetti A, Skokan M, Rossi E, Gajapathy S et al (2009) Increased MET gene copy number negatively affects survival of surgically resected non-small-cell lung cancer patients. J Clin Oncol 27:1667–1674
Toschi L, Cappuzzo F (2010) Clinical implications of MET gene copy number in lung cancer. Future Oncol 6:239–247
Sequist LV, Waltman BA, Dias-Santagata D, Digumarthy S, Turke AB et al (2011) Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med 3:75ra26
Yu HA, Arcila ME, Rekhtman N, Sima CS, Zakowski MF et al (2013) Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin Cancer Res 19:2240–2247
Engelman JA, Zejnullahu K, Mitsudomi T, Song Y, Hyland C et al (2007) MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 316:1039–1043
Spigel DR, Ervin TJ, Ramlau RA, Daniel DB, Goldschmidt JH Jr et al (2013) Randomized phase II trial of Onartuzumab in combination with erlotinib in patients with advanced non-small-cell lung cancer. J Clin Oncol 31:4105–4114
Spigel DR, Edelman MJ, Mok T, O’Byrne K, Paz-Ares L et al (2012) Treatment Rationale Study Design for the MetLung Trial: a randomized, double-blind phase III study of onartuzumab (MetMAb) in combination with erlotinib versus erlotinib alone in patients who have received standard chemotherapy for stage IIIB or IV met-positive non-small-cell lung cancer. Clin Lung Cancer 13:500–504
Yakes FM, Chen J, Tan J, Yamaguchi K, Shi Y et al (2011) Cabozantinib (XL184), a novel MET and VEGFR2 inhibitor, simultaneously suppresses metastasis, angiogenesis, and tumor growth. Mol Cancer Ther 10:2298–2308
Cui JJ, Tran-Dube M, Shen H, Nambu M, Kung PP et al (2011) Structure based drug design of crizotinib (PF-02341066), a potent and selective dual inhibitor of mesenchymal-epithelial transition factor (c-MET) kinase and anaplastic lymphoma kinase (ALK). J Med Chem 54:6342–6363
Burgess TL, Sun J, Meyer S, Tsuruda TS, Elliott G et al (2010) Biochemical characterization of AMG 102: a neutralizing, fully human monoclonal antibody to human and nonhuman primate hepatocyte growth factor. Mol Cancer Ther 9:400–409
Gordon MS, Sweeney CS, Mendelson DS, Eckhardt SG, Anderson A et al (2010) Safety, pharmacokinetics, and pharmacodynamics of AMG 102, a fully human hepatocyte growth factor-neutralizing monoclonal antibody, in a first-in-human study of patients with advanced solid tumors. Clin Cancer Res 16:699–710
Beau-Faller M, Ruppert AM, Voegeli AC, Neuville A, Meyer N et al (2008) MET gene copy number in non-small cell lung cancer: molecular analysis in a targeted tyrosine kinase inhibitor naive cohort. J Thorac Oncol 3:331–339
Drilon A, Rekhtman N, Ladanyi M, Paik P (2012) Squamous-cell carcinomas of the lung: emerging biology, controversies, and the promise of targeted therapy. Lancet Oncol 13:e418–e426
Johnson DH, Fehrenbacher L, Novotny WF, Herbst RS, Nemunaitis JJ et al (2004) Randomized phase II trial comparing bevacizumab plus carboplatin and paclitaxel with carboplatin and paclitaxel alone in previously untreated locally advanced or metastatic non-small-cell lung cancer. J Clin Oncol 22:2184–2191
Scagliotti GV, Parikh P, von Pawel J, Biesma B, Vansteenkiste J et al (2008) Phase III study comparing cisplatin plus gemcitabine with cisplatin plus pemetrexed in chemotherapy-naive patients with advanced-stage non-small-cell lung cancer. J Clin Oncol 26:3543–3551
Wistuba II, Behrens C, Virmani AK, Mele G, Milchgrub S et al (2000) High resolution chromosome 3p allelotyping of human lung cancer and preneoplastic/preinvasive bronchial epithelium reveals multiple, discontinuous sites of 3p allele loss and three regions of frequent breakpoints. Cancer Res 60:1949–1960
Wistuba II, Behrens C, Milchgrub S, Bryant D, Hung J et al (1999) Sequential molecular abnormalities are involved in the multistage development of squamous cell lung carcinoma. Oncogene 18:643–650
Zheng CX, Gu ZH, Han B, Zhang RX, Pan CM et al (2013) Whole-exome sequencing to identify novel somatic mutations in squamous cell lung cancers. Int J Oncol 43:755–764
Roth BJ, Krilov L, Adams S, Aghajanian CA, Bach P et al (2013) Clinical cancer advances 2012: annual report on progress against cancer from the American Society of Clinical Oncology. J Clin Oncol 31:131–161
Weiss J, Sos ML, Seidel D, Peifer M, Zander T et al (2010) Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci Transl Med 2:62ra93
Perez-Moreno P, Brambilla E, Thomas R, Soria JC (2012) Squamous cell carcinoma of the lung: molecular subtypes and therapeutic opportunities. Clin Cancer Res 18:2443–2451
Kim CH (2013) Druggable targets of squamous cell lung cancer. Tuberc Respir Dis (Seoul) 75:231–235
Go H, Jeon YK, Park HJ, Sung SW, Seo JW et al (2010) High MET gene copy number leads to shorter survival in patients with non-small cell lung cancer. J Thorac Oncol 5:305–313
Bass AJ, Watanabe H, Mermel CH, Yu S, Perner S et al (2009) SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet 41:1238–1242
Hussenet T, du Manoir S (2010) SOX2 in squamous cell carcinoma: amplifying a pleiotropic oncogene along carcinogenesis. Cell Cycle 9:1480–1486
Hussenet T, Dali S, Exinger J, Monga B, Jost B et al (2010) SOX2 is an oncogene activated by recurrent 3q26.3 amplifications in human lung squamous cell carcinomas. PLoS One 5, e8960
Ramos AH, Dutt A, Mermel C, Perner S, Cho J et al (2009) Amplification of chromosomal segment 4q12 in non-small cell lung cancer. Cancer Biol Ther 8:2042–2050
Rooney M, Devarakonda S, Govindan R (2013) Genomics of squamous cell lung cancer. Oncologist 18:707–716
Grob TJ, Kannengiesser I, Tsourlakis MC, Atanackovic D, Koenig AM et al (2012) Heterogeneity of ERBB2 amplification in adenocarcinoma, squamous cell carcinoma and large cell undifferentiated carcinoma of the lung. Mod Pathol 25:1566–1573
Carpten JD, Faber AL, Horn C, Donoho GP, Briggs SL et al (2007) A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 448:439–444
Malanga D, Scrima M, De Marco C, Fabiani F, De Rosa N et al (2008) Activating E17K mutation in the gene encoding the protein kinase AKT1 in a subset of squamous cell carcinoma of the lung. Cell Cycle 7:665–669
Heist RS, Sequist LV, Engelman JA (2012) Genetic changes in squamous cell lung cancer: a review. J Thorac Oncol 7:924–933
Solit DB, Garraway LA, Pratilas CA, Sawai A, Getz G et al (2006) BRAF mutation predicts sensitivity to MEK inhibition. Nature 439:358–362
Day E, Waters B, Spiegel K, Alnadaf T, Manley PW et al (2008) Inhibition of collagen-induced discoidin domain receptor 1 and 2 activation by imatinib, nilotinib and dasatinib. Eur J Pharmacol 599:44–53
Hammerman PS, Sos ML, Ramos AH, Xu C, Dutt A et al (2011) Mutations in the DDR2 kinase gene identify a novel therapeutic target in squamous cell lung cancer. Cancer Discov 1:78–89
Yamamoto H, Shigematsu H, Nomura M, Lockwood WW, Sato M et al (2008) PIK3CA mutations and copy number gains in human lung cancers. Cancer Res 68:6913–6921
Samuels Y, Diaz LA Jr, Schmidt-Kittler O, Cummins JM, Delong L et al (2005) Mutant PIK3CA promotes cell growth and invasion of human cancer cells. Cancer Cell 7:561–573
Shames DS, Wistuba II (2014) The evolving genomic classification of lung cancer. J Pathol 232:121–133
Thomas A, Rajan A, Lopez-Chavez A, Wang Y, Giaccone G (2013) From targets to targeted therapies and molecular profiling in non-small cell lung carcinoma. Ann Oncol 24:577–585
Popovic R, Licht JD (2012) Emerging epigenetic targets and therapies in cancer medicine. Cancer Discov 2:405–413
Zagni C, Chiacchio U, Rescifina A (2013) Histone methyltransferase inhibitors: novel epigenetic agents for cancer treatment. Curr Med Chem 20:167–185
Jin G, Kim MJ, Jeon HS, Choi JE, Kim DS et al (2010) PTEN mutations and relationship to EGFR, ERBB2, KRAS, and TP53 mutations in non-small cell lung cancers. Lung Cancer 69:279–283
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674
Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL et al (2012) Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med 366:2455–2465
van Meerbeeck JP, Fennell DA, De Ruysscher DK (2011) Small-cell lung cancer. Lancet 378:1741–1755
Chute JP, Chen T, Feigal E, Simon R, Johnson BE (1999) Twenty years of phase III trials for patients with extensive-stage small-cell lung cancer: perceptible progress. J Clin Oncol 17:1794–1801
Rudin CM, Durinck S, Stawiski EW, Poirier JT, Modrusan Z et al (2012) Comprehensive genomic analysis identifies SOX2 as a frequently amplified gene in small-cell lung cancer. Nat Genet 44:1111–1116
Peifer M, Fernandez-Cuesta L, Sos ML, George J, Seidel D et al (2012) Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nat Genet 44:1104–1110
Iwakawa R, Takenaka M, Kohno T, Shimada Y, Totoki Y et al (2013) Genome-wide identification of genes with amplification and/or fusion in small cell lung cancer. Genes Chromosomes Cancer 52:802–816
Kim YH, Girard L, Giacomini CP, Wang P, Hernandez-Boussard T et al (2006) Combined microarray analysis of small cell lung cancer reveals altered apoptotic balance and distinct expression signatures of MYC family gene amplification. Oncogene 25:130–138
Wistuba II, Gazdar AF, Minna JD (2001) Molecular genetics of small cell lung carcinoma. Semin Oncol 28:3–13
Larsen JE, Minna JD (2011) Molecular biology of lung cancer: clinical implications. Clin Chest Med 32:703–740
Soucek L, Whitfield J, Martins CP, Finch AJ, Murphy DJ et al (2008) Modelling Myc inhibition as a cancer therapy. Nature 455:679–683
Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J et al (2011) BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146:904–917
Sos ML, Dietlein F, Peifer M, Schottle J, Balke-Want H et al (2012) A framework for identification of actionable cancer genome dependencies in small cell lung cancer. Proc Natl Acad Sci U S A 109:17034–17039
Kwak EL, Bang YJ, Camidge DR, Shaw AT, Solomon B et al (2010) Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med 363:1693–1703
Shaw AT, Yeap BY, Mino-Kenudson M, Digumarthy SR, Costa DB et al (2009) Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J Clin Oncol 27:4247–4253
Rikova K, Guo A, Zeng Q, Possemato A, Yu J et al (2007) Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131:1190–1203
Gainor JF, Shaw AT (2013) Novel targets in non-small cell lung cancer: ROS1 and RET fusions. Oncologist 18:865–875
Wu YM, Su F, Kalyana-Sundaram S, Khazanov N, Ateeq B et al (2013) Identification of targetable FGFR gene fusions in diverse cancers. Cancer Discov 3:636–647
Shaw AT, Hsu PP, Awad MM, Engelman JA (2013) Tyrosine kinase gene rearrangements in epithelial malignancies. Nat Rev Cancer 13:772–787
Vaishnavi A, Capelletti M, Le AT, Kako S, Butaney M et al (2013) Oncogenic and drug-sensitive NTRK1 rearrangements in lung cancer. Nat Med 19:1469–1472
Nakaoku T, Tsuta K, Ichikawa H, Shiraishi K, Sakamoto H et al (2014) Druggable oncogene fusions in invasive mucinous lung adenocarcinoma. Clin Cancer Res 20:3087–3093
Seo JS, Ju YS, Lee WC, Shin JY, Lee JK et al (2012) The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res 22:2109–2119
Sasaki T, Rodig SJ, Chirieac LR, Janne PA (2010) The biology and treatment of EML4-ALK non-small cell lung cancer. Eur J Cancer 46:1773–1780
Koivunen JP, Mermel C, Zejnullahu K, Murphy C, Lifshits E et al (2008) EML4-ALK fusion gene and efficacy of an ALK kinase inhibitor in lung cancer. Clin Cancer Res 14:4275–4283
Takeuchi K, Choi YL, Togashi Y, Soda M, Hatano S et al (2009) KIF5B-ALK, a novel fusion oncokinase identified by an immunohistochemistry-based diagnostic system for ALK-positive lung cancer. Clin Cancer Res 15:3143–3149
Fang DD, Zhang B, Gu Q, Lira M, Xu Q et al (2014) HIP1-ALK, a novel ALK fusion variant that responds to crizotinib. J Thorac Oncol 9:285–294
Togashi Y, Soda M, Sakata S, Sugawara E, Hatano S et al (2012) KLC1-ALK: a novel fusion in lung cancer identified using a formalin-fixed paraffin-embedded tissue only. PLoS One 7, e31323
Choi YL, Lira ME, Hong M, Kim RN, Choi SJ et al (2014) A novel fusion of TPR and ALK in lung adenocarcinoma. J Thorac Oncol 9:563–566
Bray F, Ren JS, Masuyer E, Ferlay J (2013) Global estimates of cancer prevalence for 27 sites in the adult population in 2008. Int J Cancer 132:1133–1145
Ferlay J, Shin HR, Bray F, Forman D, Mathers C et al (2010) Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 127:2893–2917
Jabbour E, Cortes JE, Ghanem H, O’Brien S, Kantarjian HM (2008) Targeted therapy in chronic myeloid leukemia. Expert Rev Anticancer Ther 8:99–110
Wong DW, Leung EL, So KK, Tam IY, Sihoe AD et al (2009) The EML4-ALK fusion gene is involved in various histologic types of lung cancers from nonsmokers with wild-type EGFR and KRAS. Cancer 115:1723–1733
Ou SH (2011) Crizotinib: a novel and first-in-class multitargeted tyrosine kinase inhibitor for the treatment of anaplastic lymphoma kinase rearranged non-small cell lung cancer and beyond. Drug Des Devel Ther 5:471–485
Shaw AT, Kim DW, Mehra R, Tan DS, Felip E et al (2014) Ceritinib in ALK-rearranged non-small-cell lung cancer. N Engl J Med 370:1189–1197
Chabner BA (2014) Approval after phase I: ceritinib runs the three-minute mile. Oncologist 19:577–578
Takeuchi K, Soda M, Togashi Y, Suzuki R, Sakata S et al (2012) RET, ROS1 and ALK fusions in lung cancer. Nat Med 18:378–381
Davies KD, Le AT, Theodoro MF, Skokan MC, Aisner DL et al (2012) Identifying and targeting ROS1 gene fusions in non-small cell lung cancer. Clin Cancer Res 18:4570–4579
Yoshida A, Kohno T, Tsuta K, Wakai S, Arai Y et al (2013) ROS1-rearranged lung cancer: a clinicopathologic and molecular study of 15 surgical cases. Am J Surg Pathol 37:554–562
Rimkunas VM, Crosby KE, Li D, Hu Y, Kelly ME et al (2012) Analysis of receptor tyrosine kinase ROS1-positive tumors in non-small cell lung cancer: identification of a FIG-ROS1 fusion. Clin Cancer Res 18:4449–4457
Govindan R, Ding L, Griffith M, Subramanian J, Dees ND et al (2012) Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150:1121–1134
Suehara Y, Arcila M, Wang L, Hasanovic A, Ang D et al (2012) Identification of KIF5B-RET and GOPC-ROS1 fusions in lung adenocarcinomas through a comprehensive mRNA-based screen for tyrosine kinase fusions. Clin Cancer Res 18:6599–6608
Acquaviva J, Wong R, Charest A (2009) The multifaceted roles of the receptor tyrosine kinase ROS in development and cancer. Biochim Biophys Acta 1795:37–52
Ju YS, Lee WC, Shin JY, Lee S, Bleazard T et al (2012) A transforming KIF5B and RET gene fusion in lung adenocarcinoma revealed from whole-genome and transcriptome sequencing. Genome Res 22:436–445
Kohno T, Ichikawa H, Totoki Y, Yasuda K, Hiramoto M et al (2012) KIF5B-RET fusions in lung adenocarcinoma. Nat Med 18:375–377
Lipson D, Capelletti M, Yelensky R, Otto G, Parker A et al (2012) Identification of new ALK and RET gene fusions from colorectal and lung cancer biopsies. Nat Med 18:382–384
Drilon A, Wang L, Hasanovic A, Suehara Y, Lipson D et al (2013) Response to Cabozantinib in patients with RET fusion-positive lung adenocarcinomas. Cancer Discov 3:630–635
Chao BH, Briesewitz R, Villalona-Calero MA (2012) RET fusion genes in non-small-cell lung cancer. J Clin Oncol 30:4439–4441
Bergethon K, Shaw AT, Ou SH, Katayama R, Lovly CM et al (2012) ROS1 rearrangements define a unique molecular class of lung cancers. J Clin Oncol 30:863–870
Davies KD, Doebele RC (2013) Molecular pathways: ROS1 fusion proteins in cancer. Clin Cancer Res 19:4040–4045
Suzuki M, Makinoshima H, Matsumoto S, Suzuki A, Mimaki S et al (2013) Identification of a lung adenocarcinoma cell line with CCDC6-RET fusion gene and the effect of RET inhibitors in vitro and in vivo. Cancer Sci 104:896–903
Luo J, Solimini NL, Elledge SJ (2009) Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 136:823–837
Rosell R, Carcereny E, Gervais R, Vergnenegre A, Massuti B et al (2012) Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 13:239–246
Zhou C, Wu YL, Chen G, Feng J, Liu XQ et al (2011) Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol 12:735–742
Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S et al (2010) Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med 362:2380–2388
Mitsudomi T, Morita S, Yatabe Y, Negoro S, Okamoto I et al (2010) Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): an open label, randomised phase 3 trial. Lancet Oncol 11:121–128
Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT et al (2009) Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med 361:947–957
Yasuda H, Kobayashi S, Costa DB (2012) EGFR exon 20 insertion mutations in non-small-cell lung cancer: preclinical data and clinical implications. Lancet Oncol 13:e23–e31
Yasuda H, Park E, Yun CH, Sng NJ, Lucena-Araujo AR et al (2013) Structural, biochemical, and clinical characterization of epidermal growth factor receptor (EGFR) exon 20 insertion mutations in lung cancer. Sci Transl Med 5:216ra177
Girard N, Lou E, Azzoli CG, Reddy R, Robson M et al (2010) Analysis of genetic variants in never-smokers with lung cancer facilitated by an Internet-based blood collection protocol: a preliminary report. Clin Cancer Res 16:755–763
Ohashi K, Maruvka YE, Michor F, Pao W (2013) Epidermal growth factor receptor tyrosine kinase inhibitor-resistant disease. J Clin Oncol 31:1070–1080
Arcila ME, Oxnard GR, Nafa K, Riely GJ, Solomon SB et al (2011) Rebiopsy of lung cancer patients with acquired resistance to EGFR inhibitors and enhanced detection of the T790M mutation using a locked nucleic acid-based assay. Clin Cancer Res 17:1169–1180
Heuckmann JM, Holzel M, Sos ML, Heynck S, Balke-Want H et al (2011) ALK mutations conferring differential resistance to structurally diverse ALK inhibitors. Clin Cancer Res 17:7394–7401
Sasaki T, Okuda K, Zheng W, Butrynski J, Capelletti M et al (2010) The neuroblastoma-associated F1174L ALK mutation causes resistance to an ALK kinase inhibitor in ALK-translocated cancers. Cancer Res 70:10038–10043
Awad MM, Katayama R, McTigue M, Liu W, Deng YL et al (2013) Acquired resistance to crizotinib from a mutation in CD74-ROS1. N Engl J Med 368:2395–2401
Kandoth C, McLellan MD, Vandin F, Ye K, Niu B et al (2013) Mutational landscape and significance across 12 major cancer types. Nature 502:333–339
Lawrence MS, Stojanov P, Mermel CH, Robinson JT, Garraway LA et al (2014) Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505:495–501
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Wijesinghe, P., Bollig-Fischer, A. (2016). Lung Cancer Genomics in the Era of Accelerated Targeted Drug Development. In: Ahmad, A., Gadgeel, S. (eds) Lung Cancer and Personalized Medicine: Novel Therapies and Clinical Management. Advances in Experimental Medicine and Biology, vol 890. Springer, Cham. https://doi.org/10.1007/978-3-319-24932-2_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-24932-2_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-24931-5
Online ISBN: 978-3-319-24932-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)