Abstract
In 1927, Otto Warburg demonstrated that cancer cells use glucose for growth and division in a manner that is different from normal cells, a phenomenon known nowadays as Warburg Effect. To date, overwhelming evidence indicates that aberration in metabolism plays important roles in cancer progression. More recently, for more than a decade biologists are fascinated by the functions of small RNAs known as microRNAs (miRNAs), which play vital roles in many important biological processes, such as cell proliferation, differentiation, EMT/MET transition, cell signaling, response to infection, induction of pluripotent stem cells and cell metabolism. As discussed in other excellent chapters of this book, the roles of miRNAs in cancer development have been extensively studied. Here in this chapter, we will discuss the significance of miRNAs in regulating cancer cell metabolism. Specifically, we will focus on the roles of miRNAs in mediating metabolism of three major energy substrates including glucose, lipid and glutamine metabolism in cancer development.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The theory put forward by Otto Warburg many years ago is that the faster growth rate of cancer cells than that of their normal counterparts is largely because cancer cells tend to drive glycolysis followed by lactate fermentation in the cytosol as energy source rather than oxidation of pyruvate fed into mitochondria, even in the presence of oxygen. This claim is now known as Warburg Effect (Wang et al. 2009; Warburg 1956). After more than eight decades, discovery and characterization of specific metabolic changes in cancer cells continue to progress rapidly and reprogramming energy metabolism is considered an emerging hallmark of cancers (Hanahan and Weinberg 2011). During cancer development, in order to meet the excessive demand of cellular growth, cancer cells may adapt a serial of strategic alterations regarding energy metabolism. For instance, glucose, as a ubiquitous energy source in most organisms, can supply carbon, oxygen, hydrogen for anabolic processes, and its uptake is frequently enhanced in cancer cells. Glutamine, a non-essential amino acid, serves as carbon source for energy production, conduces to biosynthetic reaction by supplying carbon and nitrogen, regulates redox homeostasis through synthesis of glutathione (Dang 2010). Increased glutaminolysis is emerged as a novel feature of the metabolic profile of cancer cells, along with increased aerobic glycolysis. Another energy resource, fatty acid, which is seldom appreciated before but under the spotlight now, is required for energy storage, membrane synthesis and generation of signaling molecules (Currie et al. 2013).
Today, mutations in oncogenes and tumor suppressors are known to be answer for the malignant transformation. Furthermore, Warburg Effect is generally accepted as the result of these mutations, rather than a cause. Several famous oncogenes and tumor suppressors are conversantly participated in modifying glycolysis, glutaminolysis, lipid metabolism and various biosynthetic pathways. For instance, PI3K pathway/hypoxia/c-Myc promotes glucose uptake (Cairns et al. 2011). AMP-activated protein kinase (AMPK) regulates glucose switch through mTOR pathway (Cairns et al. 2011). Oncogenic Kras enhances glucose import at gene expression and protein levels as well as regulates glutamine metabolism in tissue-specific PDAC model (Ying et al. 2012; Son et al. 2013). Both c-Myc and p53 can manipulate glutaminolysis through glutaminase1 or glutaminase2 (Gao et al. 2009; Hu et al. 2010; Suzuki et al. 2010). Hypoxia can mediate lipogenesis by IDH1 through reductive glutamine metabolism (Metallo et al. 2011).
In the meantime, nearly two decades ago, microRNAs (miRNAs), which are approximately 22 nucleotides long, were first discovered in Caenorhabditis elegans (Lee et al. 1993; Reinhart et al. 2000). They are encoded by miRNA genes which are transcribed by RNA polymerase II or III, then forming the primary miRNA in the nucleus, the primary miRNA is processed by Drosha and DGCR8, resulting in pre-miRNA, then it is exported from nucleus into cytoplasm by exportin 5 and reprocess by Dicer. Dicer is a RNAse III enzyme that processes the pre-miRNA to a 22 nucleotides miRNA. Presumably, miRNAs regulate a third of human genes through base pairing with regions within the 3′ untranslated region of the target mRNA (Cannell et al. 2008). There is abundant evidence on the multitudinous roles of miRNAs in biological processes, covering proliferation, differentiation, aging, apoptosis, metastasis, signal transduction and so on (Babashah and Soleimani 2011).
Evidence is emerging that miRNAs are involved in multiple cancer development process including cancer metabolism. Whether we can cut off the specific energy supply for cancer cell growth by restoring normal aerobic metabolism, and how miRNAs involved in this alteration, are among the fiercely investigated topics for cancer researchers. Here, we will focus on the special roles of miRNAs played on tumor metabolism, including glucose, glutamine and lipid metabolism.
2 MicroRNAs Regulate Glucose Metabolism
2.1 MiRNAs and Glucose Homeostasis
As summarized in Table 4.1, miRNAs are shown to regulate insulin secretion, biosynthesis and sensitivity to maintain the glucose homeostasis. Lin28/let-7 axis is demonstrated as the central regulator of mammalian glucose metabolism in mouse models (Zhu et al. 2011). Let-7 is the well-known tumor suppressor, whose biogenesis blocked by the RNA-binding protein Lin28. In mice, lin28 and let-7 play the opposite role in glucose metabolism through insulin-PI3K-mTOR pathway. Global knockdown of the let-7 family with an antimiR was sufficient to prevent and treat impaired glucose tolerance in mice with diet-induced obesity (Frost and Olson 2011). Let-7 and let-7-targets such as c-Myc and Kras all have significant effects on cancer metabolism, hence, the regulation of glucose homeostasis by lin28/let-7 axis may have profound implications in cancer metabolism (Zhu et al. 2010).
The expression of miR-143 is up-regulated in the liver of genetic and dietary mouse model of obesity (Jordan et al. 2011). Highly expressed miR-143 inhibits insulin-stimulated AKT activation and impairs glucose metabolism by directly targeting oxysterol-binding-protein-related protein (ORP). In the regulation of glucose homeostasis, the pancreatic specific miRNA miR-375 (Kloosterman et al. 2007) has a profound effect. miR-375 as well as miR-124 and let-7b regulate insulin exocytosis in the pancreatic beta cell by suppressing myotrophin (MTPN) which helps secretary granules to exocytotic sites (Gauthier and Wollheim 2006). Recent study showed that miR-375 directly targets 3′-phosphoinositidedependent protein kinase-1(PDPK1), and significantly impairs the activity of PDPK1 (El Ouaamari et al. 2008), a master kinase related to human disease such as diabetes and cancer.
2.2 MiRNAs and Glycolysis
It is well known that the most fundamental metabolism alteration in tumor cells is increased levels of glucose uptake and glycolysis (also known as Warburg Effect). Glucose can’t pass across the hydrophobic cell membrane freely. They need a transport, glucose transporter proteins, also called solute carrier family 2 (GLUT). The level of glucose uptake can be affected by the activity of GLUT in cell membranes. GLUT3 is inhibited by miR-195-5p (Fei et al. 2012), which directly targets GLUT3 3′-untranslated region in human bladder cancer T24 cell lines. Via this molecular mechanism, miR-195-5p suppresses proliferation and promotes apoptosis of cancer cells in vitro. GLUT4, another transporter of GLUT family, can be up-regulated by miR-223 (Lu et al. 2010) and miR-133 (Horie et al. 2009) in rat cardiomyocytes.
MiRNAs regulate glycolysis by directly targeting key metabolic enzymes. Hexokiase 2 (HK2), the first enzyme of glycolysis that phosphorylates glucose to produce glucose 6-phosphate, was identified as a target of miR-143 (Fang et al. 2012). MiR-143 was shown to down-regulate HK2 protein level and inhibit glucose metabolism in lung cancer. This phenomenon was also confirmed in breast cancer. Further it was found that pro-inflammatory cytokines promote glycolysis mediated by miR-155, which up-regulates HK2 in two cascades of miR-155-SOCS1-STAT3-HK2 and miR-155-C/EBPβ-miR-143-HK2 (Jiang et al. 2012). Phosphogulcose isomerase (PGI) is a housekeeping cytosolic enzyme that brings the interconversion of glucose-6-phoshpate and fructose-6-phosphate, playing an important role in glycolysis and gluconeogenesis. A decrease of PGI expression leads to over-expression of miR-200 (Ahmad et al. 2011), which is associated with reversal of mesenchymal-epithelial transition (EMT) phenotype in human breast cancer cells (Davalos et al. 2012). MiR-15a/16-1 cluster, the first tumor suppressor miRNAs described in human malignancies (Calin et al. 2002), are reported to target the enzyme Aldo A which catalyzes a reversible aldol reaction in glycolysis (Calin et al. 2008). MiR-122 is the most abundantly expressed miRNA in the liver, and may be involved in lipid and cholesterol metabolism. While PTP1B is a newly identified direct target of miR-122, it is a negative regulator of insulin signaling cascade (Yang et al. 2012). And miR-122 is also predicted to target aldo A (Chen et al. 2012). Thus, these miRNAs are reported to have roles in cancer glycolysis.
2.3 MiRNAs and Glucose Stress
The change of miRNA expression is also a cause of abnormal glucose condition and subsequent cellular functions. In mesenchymal stem cells, hyperglycemic condition lead to a decrease of miR-32-5p, which promotes cell cycle by targeting tensin homologs deleted on chromosome 10 (PTEN) (Zhu et al. 2013). MiR-30a-5p over-expression induce beta cell dysfunction in vitro, which can be induced by high glucose at the concentration of 33.3 mmol/l (Kim et al. 2013). Another glucose induced miRNA, miR-29c, is known as a tumor suppressor which induces cell apoptosis, increases extracellular matrix protein accumulation and activates Rho kinase in mouse diabetic nephropathy model (Long et al. 2011). In glioma cells, a single miRNA was shown to adapt the cells to different glucose conditions. Glucose induced miR-451 promotes cell growth. While decreased miR-451 expression in low glucose slows proliferation, it enhances migration and survival (Godlewski et al. 2010). Deprivation of glucose leads to a change of miRNAs such as miR-466h-5p (Druz et al. 2012). Cultivation of mouse cell line with glucose-free medium causes the oxidative stress. The inhibition of histone deacetylases (HDACs) activity by this oxidative stress increases acetylation in miR-466h-5p promoter region, which lead to the activation of this miRNA.
3 MicroRNAs Modulate Lipid Metabolism
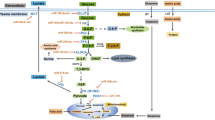
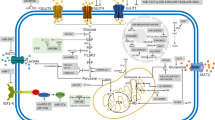
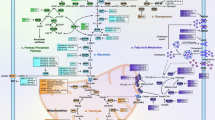
Alteration and its impact of lipid metabolism in cancer cells have long been ignored compared to the intensive interest in two other star nutrients, glucose and glutamine. This sounds a bit strange considering the pivotal roles of fatty acid in energy storage, cell proliferation and generation of membrane signal. However, last few years have witnessed a rapid development of the field and now a growing body of research is focused on lipid metabolism. Herein, we will summarize the role of several miRNAs that are involved in regulation of lipid homeostasis (Fig. 4.1).
One of the most notable miRNAs involved in lipid metabolism is liver-specific miR-122, which accounts for approximately 70 % of total miRNAs in the liver. In addition to its known roles in regulating HCV/HBV (hepatitis C/B virus) replication and expression, hepatocyte differentiation and glucose metabolism (Fukuhara et al. 2012; Qiu et al. 2010; Hu et al. 2012), miR-122 is the first miRNA to be linked recently to fat and cholesterol metabolism control. Two studies demonstrated the fundament roles of miR-122 in regulating lipid metabolism by 2′-O-methoxyethyl (2′-MOE) phosphorothioate-modified antisense oligonucleotide (ASO) in vivo (Esau et al. 2006; Fabani and Gait 2007). More recently, two independent groups discovered the tumor suppressor role of miR-122 in mouse liver using genetic deletion technology (Hsu et al. 2012; Tsai et al. 2012). Hsu et al. utilized mice which is germline knockout (KO) or liver-specific knockout (LKO) of miR-122 to describe the characteristics of fatty acid metabolism in vivo. They found that in LKO mice, the hepatic lipid level, especially triglyceride (TG) accumulation, was increased by up-regulation of Agpat1 and Mogat1 which are responsible for TG biosynthesis. But the secretion of TG into serum is reduced, thus resulting in hepatic microsteatosis in early adult life of LKO mice. Moreover, many genes involved in development, cellular proliferation, death and cancer, such as, c-Myc, c-Jun, CCND1, Igf2, Epcam, Rhoa, are aberrantly expressed in addition to the variation of lipid metabolism genes. In KO mice, increased monocytes and neutrophils are recruited to livers, leading to inflammation, the major factor attributed to malignant transformation of hepatocellular carcinoma. Production of pro-tumorigenic cytokines including IL-6 and TNF-α are also increased in LKO and KO mice which possess higher HCC incidence along with aging (Hsu et al. 2012). Tsai et al. also described the similar tumor suppressor role of miR-122 using genetic deletion in mice. They demonstrated that serum HDL and VLDL were significantly reduced in the miR-122−/− mice, which, on the contrary, causes liver steatosis. In addition, miR-122 can control hepatic fibrogenesis through one target named Krüppel-like factor 6 (KLF6) that is expressed mainly in hepatic carcinoma, whereas its expression is much lower in the normal hepatocytes of WT livers (Tsai et al. 2012). As a whole, these studies represented endeavors to decipher and manipulate the vital roles of miR-122 in regulating liver homeostasis using miR-122 antisense oligonucleotides or using an loss-of-function model in vivo (Wen and Friedman 2012).
SREBP, abbreviation of sterol regulatory element binding transcription factor, includes two protein named SREBP1 and SREBP2 whose vital function is played in sterol biosynthesis and cholesterol homeostasis by regulating transcription of sterol-regulated genes. Known miRNAs that are related with SREBP include miR-33a/b. MiR-33a is encoded in intron 16 of SREBP2 gene on chromosome 22, whereas miR-33b is encoded in intron 17 of SREBP1 gene on chromosome 17, respectively (Najafi-Shoushtari et al. 2010). The two mature miRNAs only differ in two nucleotides, but can regulate plentiful overlapping target genes, such as ATP-binding cassette transporter sub-family A member 1 (ABCA1), which is a cholesterol transporter and can mediate cholesterol efflux from within the cell to lipid-free apolipoprotein A1 (APOA-I). When ABCA1 is decreased because of elevated miR-33, then high density lipoprotein (HDL) cholesterol levels formation is destroyed, so the incidence of cardiometabolic diseases increased (Najafi-Shoushtari et al. 2010; Horie et al. 2010; Rayner et al. 2010). In mouse macrophages, miR-33 also targets ATP-binding cassette, sub-family G (white), member 1 (ABCG1), reducing cholesterol efflux to nascent HDL (Rayner et al. 2010; Marquart et al. 2010). Meantime, miR-33a and miR-33b also contribute to the regulation of several proteins involved in fatty acid β-oxidation, modulating the expression of carnitine O-octanoyltransferase (CROT), carnitine palmitoyltransferase 1A (CPT1A), and hydroxyacyl-CoA dehydrogenase-3-ketoacyl-CoA thiolase-enoyl-CoA hydratase (trifunctional protein) β-subunit (HADHB). CROT, CPT1A, and HADHB regulate the transport and degradation of fatty acids in the mitochondria (Rayner et al. 2011; Davalos et al. 2011).
Nevertheless, Tae-Il Jeon and colleagues claimed that additional miRNAs may be more robustly activated by SREBPs than miR-33 to regulate intracellular cholesterol, due to the fact that intracellular cholesterol levels are tightly controlled (Jeon et al. 2013). They performed a genome-wide analysis searching for miRNAs that are differentially expressed in livers of mice through controlled dietary, normal diet supplemented with excess cholesterol versus another diet with a combination of lovastatin plus ezetimibe (LE), drugs inhibit both endogenous cholesterol synthesis and dietary absorption of cholesterol. Intriguing finding is that miR-96/182/183 cluster comes to light. They are expressed from a unique primary transcript and the promoter for this locus is a direct target regulated by SREBP2 in mice. In this paper, miR-96 and miR-182 negatively regulate the expression of INSIG-2 and FBXW7 by targeting their 3′UTR, respectively. INSIG-2 and FBXW7 are two proteins that weaken the maturation of SREBP2, especially nuclear SREBP2 which plays the role of a transcription factor. Further analyses revealed that, when mice are fed with diet with cholesterol, normal, LE ingredient, miR-182 presents the gradient increase tendency because of the different density of cholesterol. Inversely, FBXW7 will decrease and result in accumulation of nuclear SREBP2 in LE group. This phenomenon is also conserved in human cells. So it demonstrates the coordinate and reciprocal regulation of nuclear SREBP2 with miR-182 and FBXW7. Also, these miRNAs can modulate synthesis of lipid through regulate nuclear SREBP.
Peroxisome-proliferator-activated receptors (PPARs) are nuclear hormone receptors that exert a transcriptional activity regulating genes involved in cell proliferation, cell differentiation, energy homeostasis, diabetes, obesity, atherosclerosis, and other basic metabolically processes (Peyrou et al. 2012). Three major isoforms are known: PPAR-alpha, PPAR-beta/delta, PPAR-gamma. At least five different miRNAs have been reported to regulate PPARs expression in liver cells directly or indirectly. Such as miR-10b, miR-21/miR-122 can directly regulate PPARα/PPARβ(δ) respectively. MiR-27b/miR-132 modulate PPARα/PPARγ indirectly (Zheng et al. 2010; Kida et al. 2011; Gatfield et al. 2009; Mann et al. 2010). The performance of relationship between these miRNAs and PPARs is embodied in steatosis, cholesterol and lipid metabolism. Recently, Alessio Papi et al. found that, when mammospheres derived from breast cancer stem cell are exposed with breast tumor fibroblasts supernatant, enhanced autocrine tumor necrosis factor-a (TNFα) lead to functional interplay between peroxisome proliferator activated receptor-α and hypoxia inducible factor-1α (PPARα/HIF1α), two nuclear transcriptional factors. But high PPARγ expression will antagonize the PPARα/HIF1α interplay. Further, they demonstrated that siHIF1α and siPPARα can decrease miR-130b expression, and pre-miR-130b in turn facilitates PPARα expression and decrease PPARγ. Another miRNA, miR17-5p, down-regulates PPARα, whereas it increases PPARγ expression. Through this regulatory network, they suggest that the antagonist interplay between PPARα and PPARγ is mediated by miR130b and miR17-5p. Next, they pinpoint that apolipoprotein E (ApoE) is over-expressed in mammospheres and PPARα over-expression can induce ApoE expression at mRNA and protein levels. Hence, PPARα/HIF1α interplay can regulate lipid homeostasis through control of ApoE expression (Zissel et al. 2013).
4 MicroRNAs Regulate Glutamine Metabolism
Since most cancer cells rely on aerobic glycolysis for proliferation and metabolism, fewer metabolites produced from glycolysis enter the tricarboxylic acid (TCA) cycle for energy supply. Thus, other alternative metabolites such as glutamine will play such a vital role as feeding the TCA cycle and redox homeostasis, not only as a nitrogen donor (Table 4.1). The c-Myc transcription factor, which is well known to regulate cell proliferation and glucose metabolism, has been validated to stimulate glutamine catabolism by repressing miR-23a/b that target glutaminase 1 (GLS1), resulting in an increased uptake and catabolism of glutamine (Gao et al. 2009). GLS1 catalyzes the conversion of glutamine to glutamate, the latter is further catabolized to α-ketoglutarate, which serves as a TCA substrate. Moreover, glutamate can also be converted to proline in metabolic process. C-Myc not only increases GLS, but also induces proline biosynthesis from glutamine. In 2012, Liu et al. found that c-Myc robustly suppresses the expression of POX/PRODH protein, the first enzyme in proline catabolism, primarily through increasing miR-23b*, which is processed from the same transcript as miR-23b (Liu et al. 2012).
In recent years, energy metabolism has been reported to play a major role in somatic reprogramming. TDH-mediated threonine catabolism could stimulate reprogramming process in mouse embryonic fibroblast (Han et al. 2013). In Drosophila, bioinformatics screen reveals that miR-277 acts as a metabolic switch in modulating amino acid catabolism. Hence, the regulation of amino acid metabolism by miRNAs is emerging to exert significant impact in cancer biology and beyond (Stark et al. 2003).
5 Summary and Future Perspectives
As we have discussed in this chapter, a plethora of studies now focus on the diverse roles of miRNAs played on tumor metabolism including glucose, glutamine and lipid metabolism with an aim to understand and combat cancers. Given that cancer cells adapt specific strategies for energy metabolism, one might expect to explore the regulatory mechanisms of cancer specific metabolism by miRNAs for cancer therapy. Indeed, restoration of miR-122 constitutes a novel approach that could be beneficial for therapy of chronic liver diseases and HCCs that express low levels of miR-122. Moreover, high miR-122 expression abolishes hepatic insulin resistance, resulting in lower incidence of diabetes. Nevertheless, the actual roles and regulatory networks of miRNAs in the real biological process of development and diseases are far more complicated than previously thought. In cancer metabolism, substantial cross-talks exist among the functions of miRNAs, oncogenes, nutrients enzymes and metabolites, dissection of which will greatly enhance our understanding of the complex process underlying human malignancies as well as provide insight for cancer therapy.
References
Ahmad A, Aboukameel A, Kong D, Wang Z, Sethi S, Chen W et al (2011) Phosphoglucose isomerase/autocrine motility factor mediates epithelial-mesenchymal transition regulated by miR-200 in breast cancer cells. Cancer Res 71(9):3400–3409
Babashah S, Soleimani M (2011) The oncogenic and tumour suppressive roles of microRNAs in cancer and apoptosis. Eur J Cancer 47(8):1127–1137
Cairns RA, Harris IS, Mak TW (2011) Regulation of cancer cell metabolism. Nat Rev Cancer 11(2):85–95
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al (2002) Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 99(24):15524–15529
Calin GA, Cimmino A, Fabbri M, Ferracin M, Wojcik SE, Shimizu M et al (2008) MiR-15a and miR-16-1 cluster functions in human leukemia. Proc Natl Acad Sci U S A 105(13):5166–5171
Cannell IG, Kong YW, Bushell M (2008) How do microRNAs regulate gene expression? Biochem Soc Trans 36(6):1224
Chen B, Li H, Zeng X, Yang P, Liu X, Zhao X et al (2012) Roles of microRNA on cancer cell metabolism. J Transl Med 10:228
Currie E, Schulze A, Zechner R, Walther TC, Farese RV (2013) Cellular fatty acid metabolism and cancer. Cell Metab 18(2):153–161
Dang CV (2010) Glutaminolysis: supplying carbon or nitrogen, or both for cancer cells? Cell Cycle 9(19):3884–3886
Davalos A, Goedeke L, Smibert P, Ramirez CM, Warrier NP, Andreo U et al (2011) miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proc Natl Acad Sci U S A 108(22):9232–9237
Davalos V, Moutinho C, Villanueva A, Boque R, Silva P, Carneiro F et al (2012) Dynamic epigenetic regulation of the microRNA-200 family mediates epithelial and mesenchymal transitions in human tumorigenesis. Oncogene 31(16):2062–2074
Druz A, Betenbaugh M, Shiloach J (2012) Glucose depletion activates mmu-miR-466h-5p expression through oxidative stress and inhibition of histone deacetylation. Nucleic Acids Res 40(15):7291–7302
El Ouaamari A, Baroukh N, Martens GA, Lebrun P, Pipeleers D, van Obberghen E (2008) miR-375 targets 3′-phosphoinositide-dependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic cells. Diabetes 57(10):2708–2717
Esau C, Davis S, Murray SF, Yu XX, Pandey SK, Pear M et al (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3(2):87–98
Fabani MM, Gait MJ (2007) miR-122 targeting with LNA/2′-O-methyl oligonucleotide mixmers, peptide nucleic acids (PNA), and PNA-peptide conjugates. RNA 14(2):336–346
Fang R, Xiao T, Fang Z, Sun Y, Li F, Gao Y et al (2012) MicroRNA-143 (miR-143) regulates cancer glycolysis via targeting hexokinase 2 gene. J Biol Chem 287(27):23227–23235
Fei X, Qi M, Wu B, Song Y, Wang Y, Li T (2012) MicroRNA-195-5p suppresses glucose uptake and proliferation of human bladder cancer T24 cells by regulating GLUT3 expression. FEBS Lett 586(4):392–397
Frost RJA, Olson EN (2011) Control of glucose homeostasis and insulin sensitivity by the Let-7 family of microRNAs. Proc Natl Acad Sci 108(52):21075–21080
Fukuhara T, Kambara H, Shiokawa M, Ono C, Katoh H, Morita E et al (2012) Expression of microRNA miR-122 facilitates an efficient replication in nonhepatic cells upon infection with hepatitis C virus. J Virol 86(15):7918–7933
Gao P, Tchernyshyov I, Chang T-C, Lee Y-S, Kita K, Ochi T et al (2009) c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458(7239):762–765
Gatfield D, Le Martelot G, Vejnar CE, Gerlach D, Schaad O, Fleury-Olela F et al (2009) Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev 23(11):1313–1326
Gauthier BR, Wollheim CB (2006) MicroRNAs: ‘ribo-regulators’ of glucose homeostasis. Nat Med 12(1):36–38
Godlewski J, Nowicki MO, Bronisz A, Nuovo G, Palatini J, De Lay M et al (2010) MicroRNA-451 regulates LKB1/AMPK signaling and allows adaptation to metabolic stress in glioma cells. Mol Cell 37(5):620–632
Han C, Gu H, Wang J, Lu W, Mei Y, Wu M (2013) Regulation of L-threonine dehydrogenase in somatic cell reprogramming. Stem Cells 31(5):953–965
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674
Horie T, Ono K, Nishi H, Iwanaga Y, Nagao K, Kinoshita M et al (2009) MicroRNA-133 regulates the expression of GLUT4 by targeting KLF15 and is involved in metabolic control in cardiac myocytes. Biochem Biophys Res Commun 389(2):315–320
Horie T, Ono K, Horiguchi M, Nishi H, Nakamura T, Nagao K et al (2010) MicroRNA-33 encoded by an intron of sterol regulatory element-binding protein 2 (Srebp2) regulates HDL in vivo. Proc Natl Acad Sci 107(40):17321–17326
Hsu S-h, Wang B, Kota J, Yu J, Costinean S, Kutay H et al (2012) Essential metabolic, anti-inflammatory, and anti-tumorigenic functions of miR-122 in liver. J Clin Invest 122(8):2871–2883
Hu W, Zhang C, Wu R, Sun Y, Levine A, Feng Z (2010) Glutaminase 2, a novel p53 target gene regulating energy metabolism and antioxidant function. Proc Natl Acad Sci 107(16):7455–7460
Hu J, Xu Y, Hao J, Wang S, Li C, Meng S (2012) MiR-122 in hepatic function and liver diseases. Protein Cell 3(5):364–371
Jeon T-I, Esquejo RM, Roqueta-Rivera M, Phelan PE, Moon Y-A, Govindarajan SS et al (2013) An SREBP-responsive microRNA operon contributes to a regulatory loop for intracellular lipid homeostasis. Cell Metab 18(1):51–61
Jiang S, Zhang L-F, Zhang H-W, Hu S, Lu M-H, Liang S et al (2012) A novel miR-155/miR-143 cascade controls glycolysis by regulating hexokinase 2 in breast cancer cells. EMBO J 31(8):1985–1998
Jordan SD, Krüger M, Willmes DM, Redemann N, Wunderlich FT, Brönneke HS et al (2011) Obesity-induced overexpression of miRNA-143 inhibits insulin-stimulated AKT activation and impairs glucose metabolism. Nat Cell Biol 13(4):434–446
Kida K, Nakajima M, Mohri T, Oda Y, Takagi S, Fukami T et al (2011) PPARα is regulated by miR-21 and miR-27b in human liver. Pharm Res 28(10):2467–2476
Kim JW, You YH, Jung S, Suh-Kim H, Lee IK, Cho JH et al (2013) miRNA-30a-5p-mediated silencing of Beta2/NeuroD expression is an important initial event of glucotoxicity-induced beta cell dysfunction in rodent models. Diabetologia 56(4):847–855
Kloosterman WP, Lagendijk AK, Ketting RF, Moulton JD, Plasterk RH (2007) Targeted inhibition of miRNA maturation with morpholinos reveals a role for miR-375 in pancreatic islet development. PLoS Biol 5(8):e203
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854
Liu W, Le A, Hancock C, Lane AN, Dang CV, Fan TW et al (2012) Reprogramming of proline and glutamine metabolism contributes to the proliferative and metabolic responses regulated by oncogenic transcription factor c-MYC. Proc Natl Acad Sci U S A 109(23):8983–8988
Long J, Wang Y, Wang W, Chang BH, Danesh FR (2011) MicroRNA-29c is a signature microRNA under high glucose conditions that targets Sprouty homolog 1, and its in vivo knockdown prevents progression of diabetic nephropathy. J Biol Chem 286(13):11837–11848
Lu H, Buchan RJ, Cook SA (2010) MicroRNA-223 regulates Glut4 expression and cardiomyocyte glucose metabolism. Cardiovasc Res 86(3):410–420
Mann J, Chu DCK, Maxwell A, Oakley F, Zhu NL, Tsukamoto H et al (2010) MeCP2 controls an epigenetic pathway that promotes myofibroblast transdifferentiation and fibrosis. Gastroenterology 138(2):705–714.e704
Marquart TJ, Allen RM, Ory DS, Baldan A (2010) miR-33 links SREBP-2 induction to repression of sterol transporters. Proc Natl Acad Sci U S A 107(27):12228–12232
Metallo CM, Gameiro PA, Bell EL, Mattaini KR, Yang J, Hiller K et al (2011) Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 481(7381):380–384
Najafi-Shoushtari SH, Kristo F, Li Y, Shioda T, Cohen DE, Gerszten RE et al (2010) MicroRNA-33 and the SREBP host genes cooperate to control cholesterol homeostasis. Science 328(5985):1566–1569
Peyrou M, Ramadori P, Bourgoin L, Foti M (2012) PPARs in liver diseases and cancer: epigenetic regulation by microRNAs. PPAR Res 2012:1–16
Qiu L, Fan H, Jin W, Zhao B, Wang Y, Ju Y et al (2010) miR-122-induced down-regulation of HO-1 negatively affects miR-122-mediated suppression of HBV. Biochem Biophys Res Commun 398(4):771–777
Rayner KJ, Suarez Y, Davalos A, Parathath S, Fitzgerald ML, Tamehiro N et al (2010) MiR-33 contributes to the regulation of cholesterol homeostasis. Science 328(5985):1570–1573
Rayner KJ, Sheedy FJ, Esau CC, Hussain FN, Temel RE, Parathath S et al (2011) Antagonism of miR-33 in mice promotes reverse cholesterol transport and regression of atherosclerosis. J Clin Invest 121(7):2921–2931
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE et al (2000) The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403(6772):901–906
Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M et al (2013) Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496(7443):101–105
Stark A, Brennecke J, Russell RB, Cohen SM (2003) Identification of drosophila microRNA targets. PLoS Biol 1(3):e60
Suzuki S, Tanaka T, Poyurovsky MV, Nagano H, Mayama T, Ohkubo S et al (2010) Phosphate-activated glutaminase (GLS2), a p53-inducible regulator of glutamine metabolism and reactive oxygen species. Proc Natl Acad Sci 107(16):7461–7466
Tsai WC, Hsu SD, Hsu CS, Lai TC, Chen SJ, Shen R et al (2012) MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Invest 122(8):2884–2897
Wang L, Tang H, Thayanithy V, Subramanian S, Oberg AL, Cunningham JM et al (2009) Gene networks and microRNAs implicated in aggressive prostate cancer. Cancer Res 69(24):9490–9497
Warburg O (1956) On the origin of cancer cells. Science 123(3191):309–314
Wen J, Friedman JR (2012) miR-122 regulates hepatic lipid metabolism and tumor suppression. J Clin Invest 122(8):2773–2776
Yang YM, Seo SY, Kim TH, Kim SG (2012) Decrease of microRNA-122 causes hepatic insulin resistance by inducing protein tyrosine phosphatase 1B, which is reversed by licorice flavonoid. Hepatology 56(6):2209–2220
Ying H, Kimmelman AC, Lyssiotis CA, Hua S, Chu GC, Fletcher-Sananikone E et al (2012) Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149(3):656–670
Zheng L, Lv G-c, Sheng J, Yang Y-d (2010) Effect of miRNA-10b in regulating cellular steatosis level by targeting PPAR-α expression, a novel mechanism for the pathogenesis of NAFLD. J Gastroenterol Hepatol 25(1):156–163
Zhu H, Shah S, Shyh-Chang N, Shinoda G, Einhorn WS, Viswanathan SR et al (2010) Lin28a transgenic mice manifest size and puberty phenotypes identified in human genetic association studies. Nat Genet 42(7):626–630
Zhu H, Shyh-Chang N, Segrè AV, Shinoda G, Shah SP, Einhorn WS et al (2011) The Lin28/let-7 axis regulates glucose metabolism. Cell 147(1):81–94
Zhu G, Chai J, Ma L, Duan H, Zhang H (2013) Downregulated microRNA-32 expression induced by high glucose inhibits cell cycle progression via PTEN upregulation and Akt inactivation in bone marrow-derived mesenchymal stem cells. Biochem Biophys Res Commun 433(4):526–531
Zissel G, Papi A, Storci G, Guarnieri T, De Carolis S, Bertoni S et al (2013) Peroxisome proliferator activated receptor-α/hypoxia inducible factor-1α interplay sustains carbonic anhydrase IX and apoliprotein E expression in breast cancer stem cells. PLoS One 8(1):e54968
Acknowledgments
Our work is supported in part by National Basic Key Research Program of China (2014CB910600, 2011CBA01103 and 2012CB910104), National Nature Science Foundation of China (31171385 and 31071257). H.Z is supported by Chinese Government “1000 Youth Talent Program”.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Sun, L., He, X., Cao, Y., Gao, P., Zhang, H. (2014). MicroRNAs and Energy Metabolism in Cancer Cells. In: Babashah, S. (eds) MicroRNAs: Key Regulators of Oncogenesis. Springer, Cham. https://doi.org/10.1007/978-3-319-03725-7_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-03725-7_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-03724-0
Online ISBN: 978-3-319-03725-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)