Abstract
Given the phylogenetic relationships of several extant species, the reconstruction of their ancestral genomes at the gene and chromosome level is made difficult by the cycles of whole genome doubling followed by fractionation in plant lineages. Fractionation scrambles the gene adjacencies that enable existing reconstruction methods. We propose an alternative approach that postpones the selection of gene adjacencies for reconstructing small ancestral segments and instead accumulates a very large number of syntenically validated candidate adjacencies to produce long ancestral contigs through maximum weight matching. Likewise, we do not construct chromosomes by successively piecing together contigs into larger segments, but instead count all contig co-occurrences on the input genomes and cluster these, so that chromosomal assemblies of contigs all emerge naturally ordered at each ancestral node of the phylogeny. These strategies result in substantially more complete reconstructions than existing methods. We deploy a number of quality measures: contig lengths, continuity of contig structure on successive ancestors, coverage of the reconstruction on the input genomes, and rearrangement implications of the chromosomal structures obtained. The reconstructed ancestors can be functionally annotated and are visualized by painting the ancestral projections on the descendant genomes, and by highlighting syntenic ancestor-descendant relationships. We apply our methods to genomes drawn from a broad range of monocot orders, confirming the tetraploidization event “tau” in the stem lineage between the alismatids and the lilioids.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Genome reconstruction
- Gene order
- Polyploidization
- Fractionation
- Monocots
- Generalized adjacencies
- Multiple orthology
- Safe phylogeny
- Maximum weight matching
- Co-occurrence matrix
- Complete-link clustering
- Linear ordering problem
1 Introduction
Reconstruction methods depending on conserved gene adjacencies tend to break down in plants, largely because the history of whole genome doubling and tripling events (WGD and WGT, respectively) in the lineages of plants. All known flowering plant genomes (except Amborella trichopoda [1]) have at least one, and often several, WGDs or WGTs in their lineages since the ancestral angiosperm, followed by extensive loss of redundant genes, largely randomly distributed along one or other of the duplicated chromosomes. These processes effectively scramble gene order and disrupt most adjacencies. Subsequently, most of the sets of duplicate or triplicate genes created by WGD/WGT events are reduced sooner or later to a single gene, by the redundance-eliminating process known as gene fractionation. Because of this fractionation, duplication of a genome fragment containing genes in the order 1-2-3-4-5-6, for example, may result in two surviving orders 1-3-5 and 2-4-6, with none of the five fragment-internal adjacencies conserved, and only one adjacency at most conserved with the chromosomal regions surrounding each copy of the fragment. The situation is compounded if there are several WGD or WGT events in the history of some of the present-day genomes. All this is superimposed on a background of gene family expansion through tandem duplication or other mechanisms, and loss of genes from species for which they are no longer physiologically or ecologically essential, genome rearrangement and other processes, all of which disrupt adjacencies independently of the fractionation process.
For this paper, we developed a pipeline for ancestral plant genome inference, RACCROCHE, Reconstruction of AnCestral COntigs and CHromosomEs, including some intermediate ancestral genomes giving rise to major plant subgroupings. The new strategy implemented in our approach combines six fundamental components:
-
1.
The replacement of the traditional selection of 1-1 orthologs among input genomes, as a first step, by the identification of many-to-many correspondences among gene families of limited size within these genomes.
-
2.
The use of generalized adjacencies [17, 18], namely any pair of genes close to each other on a chromosome, instead of just immediately adjacent genes.
These first two components avoid premature decisions on which orthologies and which adjacencies should be incorporated in the final reconstruction, in contrast to approaches which insist on making these decisions early in the reconstruction process, e.g., [11].
-
3.
The compilation of oriented candidate adjacencies at each of the ancestral nodes of a given binary branching tree phylogeny using a “safe” criterion - that such an adjacency must be evidenced in genomes in two or three of the subtrees connected by this node, not just one or none.
-
4.
The large set of these candidates is then resolved, at each node, by maximum weight matching (MWM) to give an optimally compatible subset, which ipso facto defines linearly (or circularly) compatible “contigs” of the ancestral genomes to be constructed, thus avoiding the branching segments that plague other methods [14].
-
5.
A local sequence matching, satisfying proximity and contiguity conditions, of each contig on all of the chromosomes of the input genomes. This step includes the construction of a total chromosomal co-occurrence matrix of contigs belonging to each ancestral node.
-
6.
A clustering applied to the co-occurrence matrix. This is then decomposed into chromosomal sets of contigs, with the aid of a heat map comparison of the contigs as organized by the clustering. Within each contig, the order of the genes is already predetermined by the MWM step. Ordering the contigs along the chromosomes is carried out by a linear ordering algorithm. The assignment and ordering of contigs to construct entire chromosomes, and not just a collection of small regions, is an advance over previous methods. Corresponding chromosomes in different ancestral genomes can be identified by the similar contigs they contain.
The results of this pipeline are mapped back to the input genomes, indicating how these extant genomes were derived through chromosomal rearrangements from their immediate ancestral genome.
We provide an evaluation of the reconstruction in terms of the sizes of the ancient chromosomal fragments found, the coherence (or continuity) between adjacent ancestral genomes, the coverage of the ancestors when mapped to extant genomes, and the “choppiness” of this mapping in terms of ancestor-descendant rearrangement.
There has been much recent work on the reconstruction of ancestral plant genomes [3, 4, 10, 12, 19]; on the computational side most of this has been based on common gene adjacencies in extant genomes, as summarized in such structures as sets of species trees and contiguous ancestral regions (CARS) [2]. The latter terminology, introduced successfully in the context of mammalian genomes [7], where there are no polyploidizations since the common ancestor, and then taken over to plant genomics [4, 5, 12], applies to a series of methods of which a recent improved exemplar is proCARs [11]. We will show that in the case of flowering plants, the avoidance of premature selection of gene adjacencies in RACCROCHE allows the recovery of more of the ancestral genome than proCARs.
The rest of the paper is organized as follows. Section 2 presents the features and procedure of the algorithm. (Most of the details appear in appendices.) An application of the RACCROCHE pipeline is shown in Sect. 3 with a focus on the reconstruction of the four monocot ancestors in the known phylogeny relating six extant monocot plant genomes. These include Acorus calamus (sweet flag) from the order Acorales, Spirodela polyrhiza (duckweed) from the order Alismatales, Dioscorea rotundata (yam) from the order Dioscorales, Asparagus officinalis (asparagus) from the order Aspargales, Elaeis guineensis (African oil palm) from the order Arecales and Ananas comosus (pineapple) from the order Poales. This includes an evaluation of the reconstruction in terms of the sizes of the ancient chromosomal fragments found, the coherence between adjacent ancestral genomes, the coverage of the ancestors when mapped to extant genomes, and the “choppiness” of this mapping in terms of ancestry descendant rearrangement. Section 4 concludes the paper and outlines some future directions.
2 Methods
2.1 Input
The input to RACCROCHE consists of N annotated extant genomes related by a given unrooted binary branching phylogeny, and a number of parameters, including
-
W: window size to include generalized as well as immediate adjacencies,
-
NF: largest total gene family size allowed in ortholog grouping in all extant genomes,
-
NG: largest gene family size allowed in any one genome,
-
NC: the number of longest contigs in ancestral genomes to be matched to extant genomes,
-
K: the desired number of chromosomes for each ancestor,
-
DIS: the maximum distance between two adjacent genes in an extant genome to be matched with adjacent genes in an ancestral contig.
Figure 1 depicts the overall flow of the RACCROCHE pipeline.
2.2 The Pipeline
Step 1: Pre-process gene families. Pre-processing for the RACCROCHE procedure starts with syntenically validated orthogroups, or gene families, constructed from \(\frac{1}{2}(N^2+N)\) between-genome and self-comparison sets of pairwise SynMap synteny blocks by accumulating all genes that are syntenically orthologous to at least one other gene in the family. It retains only those families with at most a preset number NF of members and at most NG members in any particular genome. Without loss of generality, \(NF\le N \times NG\).
The use of syntenically validated adjacencies only, restricted to genes appearing in synteny blocks identified by the comparison of some pair of the descendant genomes, avoids generating huge gene families and astronomical numbers of adjacencies not reflective of the ancestor.
An optional second “redistribution” step for genes in large families is described in Appendix A.
Step 2: List generalized adjacencies. For each of the N extant genomes, RACCROCHE compiles all generalized adjacencies, i.e., representatives of two gene families, occurring within a window of a preset size, W, in the order of genes on a chromosome. The adjacencies are oriented by the DNA strand or strands containing the two genes, so that we can distinguish the two ends of each gene and identify which ends are involved in the adjacency.
Step 3: List candidate adjacencies. For each ancestral tree node, allow only adjacencies in occurring in two or three of the three subtrees connected by a branch incident to that node as candidates to be adjacencies in the corresponding ancestral genome. Occurrence in a subtree means occurrence in at least one of the extant genomes in that subtree.
Step 4: Construct contigs. With candidate adjacencies weighted 2 or 3 according to whether they occur in 2 or 3 subtrees, use maximum weight matching to extract the highest weight set of compatible adjacencies, i.e., each gene end is matched to at most one other gene end, which automatically defines a set of disjoint linear contigs for the ancestral genome.
A method for improving the coherence of successive ancestors is discussed in Appendix B. This comes at the cost of other qualities of the contigs, and will not be discussed further here.
Step 5: Match synteny blocks between ancestral genome and extant genomes. For each of the NC longest contigs of an ancestral genome, search for locally matched regions - synteny blocks - in all N extant genomes. This process is formally described in Appendix C.
Step 6: Cluster ancestral contigs into ancestral chromosomes. Clustering of ancestral chromosomes is based on co-occurrence of ancestral contigs of sufficient size on the same chromosomes of extant genomes. First, a co-occurrence matrix is constructed on the set of contigs counting the cumulative number of times two different contigs are matched on the same chromosome in one or more extant genomes. Next, a complete-link clustering of the contigs is performed in each ancestral genome, based on the co-occurrence matrix. The hierarchical cluster thus produced is decomposed either automatically (e.g., with a cut-off level or with a cluster size criterion) or with some biologically-motivated manual intervention into a preset number K of chromosomes. See Sect. 3.2 below for an example.
Contigs are ordered by applying the algorithm of linear ordering problem [13] based on the count of relative ordering, the number of times each contig appears upstream/downstream of the other contig for every pair of contigs within a cluster.
The clustering and ordering are detailed in Appendix D. These procedures have been validated through simulation studies [16].
2.3 Visualizing and Evaluating the Reconstruction
Step 7: Painting the extant genomes according to the ancestral chromosomes. Each of the K chromosomes of an ancestor genome is assigned a different colour. Each extant genome can then be painted by the colours of an ancestor based on the coordinates of synteny blocks calculated in Step 5. Unpainted regions less than 1Mb long between two blocks of the same colour are also painted with that colour. Although we can establish a general correspondence between the chromosomes of the successive ancestor genomes, the synteny blocks and the painting of the extant genomes will nevertheless depend on which ancestor is used. Generally the immediate ancestor of a genome gives the most meaningful painting.
Step 8: Adapting MCScanX to match ancestral genomes with extant genomes. We use MCScanX [15] to connect matching parts of each descendant and its immediate ancestor, as well as to calculate the optimal order of chromosomes.
MCScanX requires both gene location and gene sequence to search pairwise synteny. The “genes” in the constructed ancestors, however, are really gene familes, each represented by an integer label. For the purposes of MCScanX, we simply choose a member of the gene family, either randomly, or from a descendant of that ancestor.
For viewing purposes, the number of “crossing” lines in the trace diagram should be minimized. MCScanX searches for the ordering of the chromosomes that minimizes this, using a genetic algorithm.
Step 9: Measures of Quality. In the construction of the contigs, we count how many gene families and how many candidate adjacencies are incorporated in total by the MWM and in the longest NC chromosomes. We also document details of the contig length distribution, e.g., the longest contig and N50.
The coherence between all pairs of contig sets, each set associated with one ancestor is a way of more global way of assessing the reconstruction. To be credible, the contigs at one ancestral node should resemble to some extent the contigs at a neighbouring ancestor.
A measure of commonality between two contigs i and j from two ancestors I and J respectively, is given by
where \(x_{i.}, x_{.j}\) and \(x_{ij}\) are the numbers of gene families in contig i, in contig j and in both contigs, respectively.
Then, calculating the coherence between two tree nodes for the NC longest contigs.
Percent coverage is defined as the percentage that genome G is covered by the synteny block set of ancestor A. It also reflects how closely ancestor A is related to G.
Choppiness of painting in G is quantitatively measured by the number of different colours, T, the number of single-colour regions, R, and the number of small stripes, X, on each extant chromosome [9]. T is defined as the sum number of different colours on each chromosome of G minus 1, reflecting how much inter-chromosomal exchange, such as translocation, there has been; R is defined as the sum number of single-colour regions on each chromosome of G and is a measure of how much intra-chromosomal movement (e.g., reversals or transpositions) there has been; X is defined as the number of stripes less than a certain threshold size (i.e. 300 Kbp), which we deduct to avoid inflating R. The choppiness measure of painting in G is written as \(R-X\).
2.4 Ancestral Gene Function
To aid in future studies of the genomic organization of gene function, a GO-term enrichment analysis of the members of each gene family is implemented to produce a functional annotation for the inferred ancestral genes. The details are reported in Appendix E, but are not applied in this paper.
3 Reconstruction of Monocot Ancestors
We applied our method to the reconstruction of four monocot ancestors, given six extant monocot plant genomes from Acorus calamus (sweet flag), Spirodela polyrhiza (duckweed), Dioscorea rotundata (yam), Asparagus officinalis (asparagus), Elaeis guineensis (African oil palm) and Ananas comosus (pineapple). The phylogenetic tree is shown in Fig. 2. The divergence time from Ancestor 1 to any of the extant genomes is about 130 Mya [6]. The reconstruction problem is difficult due not only to this lengthy elapsed time, since the early Cretaceous, comparable to that of the early divergence of placental mammals, but also to the occurrence of at least one WGD in every order, and generally two or more.
One question we aimed to answer was whether both ancient WGD detected in the extant Dioscorea genome occurred after its branching off the stem lineage to Asparagales, Arecales and Poales, or whether one of these WGD occurred earlier, between Ancestors 1 and 2, and is identical to the “tau” event known to affect all these later branching orders.
3.1 Properties of the Contig Reconstruction
After numerous trials, input parameters that seemed (somewhat subjectively) to balance contig length properties, coherence and coverage were chosen to be window size \(W=7\), maximum total family size \(NF=50\) and within-genome maximum family size \(NG=10\). Table 1 summarizes the gene content of each of the input genomes, first, syntenically validated genes (i.e., in synteny blocks); second, after removing very large gene families; third, after filtering for within-genome family size; fourth, genes present in a candidate adjacency; fifth, genes incorporated in the 250 longest contigs for any ancestor.
Recall that to be a candidate, an adjacency must appear at least once in at least two different genomes, thus satisfying the safety criterion for at least one ancestor. Applying the MWM algorithm to the set of candidates greatly reduces the number in selecting the best linearized subset, as documented in Table 2.
The contigs that are formed by the MWM matches are of moderate length, as suggested by Table 3. The longest one contains 84–89 genes and the last one retained (\(NC=250\)) contains around 10 genes. We then locate all the matches of these contigs on the chromosomes of the extant genomes.
A good proportion of the MWM adjacencies will be shared by successive (or all) ancestors, and many contigs will be similar from ancestor to ancestor. Table 4 displays the coherence among the contig sets for the four ancestor genomes.
3.2 Clustering
The choice of complete link method of hierarchical clustering is appropriate in the context of searching for balanced clusters at all levels, and avoiding an asymmetric “chaining” effect. Chromosomes in a genome tend to be roughly the same order of magnitude, which therefore suggests complete link.
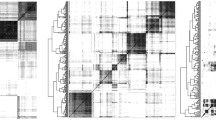
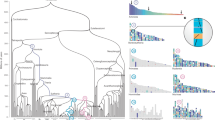
The hierarchical cluster of the 250 longest contigs according to their chromosomal co-occurrence (Sect. 2.2) is seen beside each panel in Fig. 3. The intensity of the shading of each cell in the heat map reflects how frequently the corresponding contigs co-occur in the extant genomes. In each case seven large, darkly shaded, blocks emerge neatly from the map, thus constituting the chromosomes of the ancestral genome. Table 5 contains statistics on the chromosomes and contigs.
3.3 Painting the Chromosomes of the Present-Day Genomes
Each chromosome in an ancestor genome is assigned a colour. Despite the genome rearrangements intervening between an earlier ancestor and a later one, corresponding chromosomes in different ancestral genomes can be identified by similarity in the gene content of their constituent contigs. This correspondence, though it disrupted in many places by interchromosomal exchanges, is reflected in the chromosomal colour assignment in the four ancestors. The colours are then projected onto the chromosomes of the extant genomes that served as inputs to the pipeline, based on the contig matches detected in Sect. 3.1. Painting is carried out as described in Sect. 2.3 and is depicted in Fig. 4.
3.4 Evaluation
Tables 6 and 7 provide quality assessments of the reconstruction as manifest in the painted extant genomes. In Table 6 we see a high degree of coverage of the extant genomes, while Table 7 shows a degree of choppiness that is moderate, given the time scale involved. Ancestors 1 and 2 achieve better coverage of all the extant genomes, even though most of the genomes were more directly involved in the reconstruction of Ancestors 3 and 4. This may be an artifact of the sparsity of matches from Ancestors 1 and 2, so that the inter-block colouring discussed in Sect. 2.3 can cover longer, uninterrupted, regions of the chromosomes. A similar sparsity explanation can also be entertained for the low degree of choppiness of the paintings on the Spirodela genome, despite its higher degree of polyploidy than Acorus.
3.5 MCScanX Visualization
A different view of the evolution of the monocot genomes via ancestral intermediates is obtained through connecting homologous synteny blocks in a MCScanX visualization, as laid out in Fig. 5. Consistent with the history of extensive rearrangement evident in Fig. 4 and Table 7, the patterns of MCScanX connections is rather complex. Nevertheless, we can find important relationships using the “highlight” feature of the software.
Thus, the comparison between Ancestor 1 and Acorus shows several chromosomal regions in the ancestor each linked to two regions in the extant genome, whereas the opposite pattern is non-existent. Similarly the comparison between Ancestor 1 and Spirodela also shows instances of a 1:4 pattern, consistent with the two WGDs inherited by this species.
The most interesting pattern, however, is that between Ancestors 1 and 2, which strongly suggests a duplication event occurring before the branching of the Dioscorales from the main monocot stem lineage. In contrast the Ancestor 2-Ancestor 3 and Ancestor 3-Ancestor 4 comparisons both show 1-1 patterns. Moreover, though dot-plot examination of Dioscorea evidences four subgenomes, thus two WGD in its history, the MCScanX diagram of Ancestor 2-Dioscorea only shows evidence of one event, confirming that one event must have predated Ancestor 2. This latter event is the one shared by all the more recently branching orders, known as “tau”.
4 Discussions and Conclusions
This work explored an alternative approach to genome reconstruction by stepwise piecing together of small units. Instead, we compile a large number of potential components and use a combinatorial optimization approach to combining them, an approach explicitly disavowed by, e.g., [11]. We were motivated by the special case of plant comparative genomics, which has to deal with the aftermath or recurrent polyploidization and fractionation. Compared to approaches like proCARs [11] which is very successful in reconstructing ancestral animal genomes, RACCROCHE may work better with plant genomes, since it is designed to be robust against the gene order scrambling effect of fractionation.
Since the entities reconstructed by proCARs are not meant to be individual ancestral genes, but blocks of syntenically related genes identified at the level of extant genomes, it is hard to compare our inferred ancestral genomes, composed of hypothetical genes with identifiable functions, with the output of proCARs. In our hands proCARs identified 214 synteny blocks in our data, organized into “CARs” (contiguous ancestral regions) making up the ancestral genomes. These contained a total of 3,248 “universal seeds”, which may be comparable to our ancestral genes, although our ancestors contained about twice as many. Insofar as these comparisons are valid, they confirm a role for RACCROCHE in plant comparative genomics.
One particular feature that stands out in this work, is the innovative clustering of counts of contig co-occurrences on extant chromosomes, followed by heatmap construction to identify ancestral chromosomes. Another is the use of MCScanX to locate a WGD on an internal branch of a phylogeny.
Availability
The annotated genomic data is accessible on the CoGe platform https://genomevolution.org/coge/ and Phytozome. The pipeline is available at https://github.com/jin-repo/RACCROCHE.
References
Amborella Genome Project: The Amborella genome and the evolution of flowering plants. Science 342(6165), 1241089 (2013)
Anselmetti, Y., Luhmann, N., Bérard, S., Tannier, E., Chauve, C.: Comparative methods for reconstructing ancient genome organization. In: Setubal, J.C., Stoye, J., Stadler, P.F. (eds.) Comparative Genomics. MMB, vol. 1704, pp. 343–362. Springer, New York (2018). https://doi.org/10.1007/978-1-4939-7463-4_13
Avdeyev, P., Alexeev, N., Rong, Y., Alekseyev, M.A.: A unified ILP framework for core ancestral genome reconstruction problems. Bioinformatics 36(10), 2993–3003 (2020)
Badouin, H., et al.: The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 546(7656), 148–152 (2017)
Chauve, C., Tannier, E.: A methodological framework for the reconstruction of contiguous regions of ancestral genomes and its application to mammalian genomes. PLoS Comput. Biol. 4(11), e1000234 (2008)
Givnish, T.J., et al.: Monocot plastid phylogenomics, timeline, net rates of species diversification, the power of multi-gene analyses, and a functional model for the origin of monocots. Am. J. Bot. 105(11), 1888–1910 (2018)
Ma, J., et al.: Reconstructing contiguous regions of an ancestral genome. Genome Res. 16(12), 1557–1565 (2006)
Martí, R., Reinelt, G., Duarte, A.: A benchmark library and a comparison of heuristic methods for the linear ordering problem. Comput. Optim. Appl. 51(3), 1297–1317 (2012). https://doi.org/10.1007/s10589-010-9384-9
Mazowita, M., Haque, L., Sankoff, D.: Stability of rearrangement measures in the comparison of genome sequences. J. Comput. Biol. 13(2), 554–566 (2006)
Murat, F., Armero, A., Pont, C., Klopp, C., Salse, J.: Reconstructing the genome of the most recent common ancestor of flowering plants. Nat. Genet. 49, 490–496 (2017)
Perrin, A., Varré, J.S., Blanquart, S., Ouangraoua, A.: ProCARs: progressive reconstruction of ancestral gene orders. BMC Genomics 16(S5) (2015). Article number: S6. https://doi.org/10.1186/1471-2164-16-S5-S6
Rubert, D.P., Martinez, F.V., Stoye, J., Doerr, D.: Analysis of local genome rearrangement improves resolution of ancestral genomic maps in plants. BMC Genomics 21, 1–11 (2020). https://doi.org/10.1186/s12864-020-6609-x
Schiavinotto, T., Stützle, T.: The linear ordering problem: instances, search space analysis and algorithms. J. Math. Model. Algorithms 3(4), 367–402 (2004). https://doi.org/10.1007/s10852-005-2583-1
Tannier, E., Bazin, A., Davín, A., Guéguen, L., Bérard, S., Chauve, C.: Ancestral genome organization as a diagnosis tool for phylogenomics (2020)
Wang, Y., et al.: MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40(7), e49 (2012)
Xu, Q., Jin, L., Zheng, C., Leebens-Mack, J.H., Sankoff, D.: Validation of automated chromosome recovery in the reconstruction of ancestral gene order. Algorithms 14, 160 (2021)
Xu, X., Sankoff, D.: Tests for gene clusters satisfying the generalized adjacency criterion. In: Bazzan, A.L.C., Craven, M., Martins, N.F. (eds.) BSB 2008. LNCS, vol. 5167, pp. 152–160. Springer, Heidelberg (2008). https://doi.org/10.1007/978-3-540-85557-6_14
Yang, Z., Sankoff, D.: Natural parameter values for generalized gene adjacency. J. Comput. Biol. 17(9), 1113–1128 (2010)
Zheng, C., Chen, E., Albert, V.A., Lyons, E., Sankoff, D.: Ancient eudicot hexaploidy meets ancestral eurosid gene order. BMC Genomics 14(S7), S3 (2013)
Acknowledgements
We thank the Department of Energy Joint Genome Institute staff and collaborators including David Kudrna, Jerry Jenkins, Jane Grimwood, Shengqiang Shu, and Jeremy Schmutz for pre-publication access to the Acorus genome sequence and annotation. Thanks to Aîda Ouangraoua for much help in implementing ProCARs [11] and Haibao Tang for prompt replies to queries about MCScanX [15].
Funding
Research supported by Discovery grants to LJ and DS from the Natural Sciences and Engineering Research Council of Canada. DS holds the Canada Research Chair in Mathematical Genomics.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendices
Appendices
A Redistributing Genes from Families Exceeding Upper Size Limits
As an optional second “redistribution” step, all families with more than NF members or more than NG members in any particular genome, are flagged. Then the construction of the families is repeated, with the restriction that no gene can be recruited to a family by virtue only of a similarity of less than some threshold homology level \(\theta \) to a gene already in the family. The intent is to break up large families held together by a few weak links, and thus to retrieve some better supported smaller families.
B Modes of Contig Construction
RACCROCHE executes for a single set of W, NF, NG parameters, or for a range of values of W and NG. In the latter case, there is an option, designed to increase coherence among sets of contigs for successive ancestors, that the MWM for any combination of W and NG must be restricted to include all adjacencies already recovered for lesser values of W or NG, insofar as possible. Thus, starting with some small W and NG, we can construct MWM solutions for larger window size and/or larger gene family size, and hence sets of contigs, by incrementing one or the other of the parameters.
It is possible, however, to have conflicts between \(W,NG-1\), and \(W-1,NG\) analyses. For example if adjacencies (a, b) and (b, c) are in the MWM for \((W, NG-1)\) and (a, b) and (b, d) are in the MWM for \((W-1, NG)\), then a matching for W, G cannot be forced to include all matchings from the two previous MWM. To accommodate this possibility, when we restrict the MWM for (W, NG) to include all adjacencies from \((W,NG-1)\) and \((W-1,NG)\), we make an exception for any adjacencies from either that are in potential conflict with adjacencies from the other. Thus (a, b) in the example above might be obligatorily included, but (b, c) and (b, d) would not. Thus the MWM for (W, NG) might include (b, c) or (b, d), but not both.
C Matching Contigs to Chromosomes of Extant Genomes
For the ancestor genome, A, computed from a set of extant genomes neighbouring A, \(G_{1\cdots n}\), perform the following steps.
-
1.
Extract gene features of ancestor A in descendant genomes.
For every gene, g, in ancestor A computed from Step 2, retrieve six features of this gene in every extant genome \(G_{1\cdots n}\) involved in constructing ancestor A. The features of a gene include chromosome ID, start and end chromosomal positions, distance between g to its next adjacent gene in \(G_i\), gene family ID labelled in Step 1, and contig ID in A, denoted as \(g^{A\rightarrow G_i } (chr,start,end,distance,gf,ctg)\).
-
2.
Map ancestor A to each of the descendant genomes.
The ancestor will be mapped as ancestral syntenic blocks on the descendant genome in two steps. The first step initializes a syntenic block by merging two adjacent genes given a distance threshold DIS: merge two genes, \(g_1\) and \(g_2\), forming one ancestral syntenic block on \(G_i\) if \(g_1\) and \(g_2\) satisfy the following conditions:
-
(a)
\(g_1\) and \(g_2\) locate the same chromosome of \(G_i\);
-
(b)
\(g_1\) and \(g_2\) are adjacent to each other; in other words, there could be a non-coding region but no other gene(s) between \(g_1\) and \(g_2\);
-
(c)
The distance between the two adjacent genes must be less than or equal to the distance threshold DIS (i.e. \(DIS=1\) Mbp).
The second step extends the above identified ancestral syntenic block by merging flanking gene(s) into the block if the gene(s) satisfies the above three conditions. It stops extending the block if no flanking gene could be merged into the block. After the two steps, an ancestral synteny block mapping A to \(G_i\) is denoted as syntenyBlk(chr, start, end, ctg, len). The set of synteny blocks between A and \(G_i\) is
\(syntenyBlkSet^{A\rightarrow G_i}=\{syntenyBlk_k(chr,start,end,ctg,len) | 1\le k\le m,\) where m is the total number of synteny blocks mapping from A to \(G_i\)}
-
(a)
D Construction of Ancestral Chromosomes
-
1.
Filter the set of blocks longer than a block length threshold.
Given a block length threshold, blockLEN, \(\overline{syntenyBlkSet}^{A\rightarrow G_i}\) is a subset of \(syntenyBlkSet^{A\rightarrow G_i}\), where each block in the set is longer than blockLEN (i.e. \(blockLEN = 150\) Kbp).
-
2.
Count co-occurrence of ancestral contigs on same chromosomes.
Based on syntenyBlk.chr and syntenyBlk.ctg of each pair of synteny block in \(\overline{syntenyBlkSet}^{A\rightarrow G_i}\), gather the co-occurrence of ancestral contigs on the same extant chromosome. Write the co-occurrence result into the lower triangle of a \(NC\times NC\) matrix, m, where the rows and columns are contigs with ID from 0 to \((NC-1)\), \(m_{i,j}\) is the number of co-occurrence between contigs i and j, where \(0<j<i<NC-1\). The maximum co-occurrence frequency in m is denoted as \(\max _{freq}\).
-
3.
Cluster ancestral contigs into ancestral chromosomes according to pairwise distance matrix based on co-occurrence.
A NC by NC distance matrix, dmat, is calculated as
$$dmat_{i,j}= -\log (\frac{\max _{freq}-m_{i,j} }{\max _{freq}}).$$This distance matrix is fed into the complete-link clustering algorithm. This can then be composed into K clusters, according to users’ preferences. The resultant clusters of contigs correspond to ancestral chromosomes and their compositions.
Last, attach ancestral chromosome number as an attribute to each of the synteny block:
where \(ancestral\_chr\) corresponds to the cluster ID which blk.ctg belong to.
To order the contigs along each chromosome, we proceed as follows.
After the \(syntenyBlkSet^{A\rightarrow G_{1\cdots N}}\) is generated in Step 3, relative ordering between every pair of contigs is counted. The number of times each contig appears upstream/downstream of other contig is structured into an \(NC\times NC\) ordering matrix, C, where the rows and columns are contig IDs from 0 to \(NC-1\). \(c_{i,j}\) represents the number of times contig i occurred in upstream of contig j in the extant chromosomes.
Given the ordering matrix C, the linear ordering problem (LOP) is the problem of finding a permutation \(\pi \) of the column and row indices \(\{1, \cdots , NC\}\), such that the value
is maximized [13]. In other words, the goal is to find a permutation of the columns and rows of C such that the sum of the elements in the upper triangle is maximized.
By applying a meta-heuristic solver of LOP, Tabu Search [8], the solution order corresponds to the ordering/permutation of contigs sorted by their positions along ancestral chromosomes.
E Functional Annotation of Ancestral Genes
We create a set of all genes in all families represented by ancestral genes in the reconstructed ancestor. This is the background set. For each gene family, all the genes in the family constitute a query set for GO-term enrichment analysis against the background set. Significant terms that emerge constitute the functional annotation for the ancestral gene.
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Xu, Q., Jin, L., Zheng, C., Leebens Mack, J.H., Sankoff, D. (2021). RACCROCHE: Ancestral Flowering Plant Chromosomes and Gene Orders Based on Generalized Adjacencies and Chromosomal Gene Co-occurrences. In: Jha, S.K., Măndoiu, I., Rajasekaran, S., Skums, P., Zelikovsky, A. (eds) Computational Advances in Bio and Medical Sciences. ICCABS 2020. Lecture Notes in Computer Science(), vol 12686. Springer, Cham. https://doi.org/10.1007/978-3-030-79290-9_9
Download citation
DOI: https://doi.org/10.1007/978-3-030-79290-9_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-79289-3
Online ISBN: 978-3-030-79290-9
eBook Packages: Computer ScienceComputer Science (R0)