Abstract
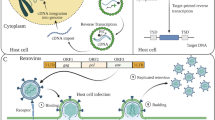
Retrotransposons are transposable elements that duplicate themselves by converting their transcribed RNA genome into cDNA, which is then integrated back into the genome. Retrotransposons can be divided into two major classes based on their mechanism of transposition and the presence or absence of long terminal repeats (LTRs). In contrast to mammalian genomes, in which non-LTR retrotransposons have proliferated, plant genomes show evolutionary evidence of an explosion in LTR retrotransposon copy number. These retrotransposons can comprise a large fraction of the genome (75 % in maize). Although often viewed as molecular parasites, retrotransposons have been shown to influence neighboring gene expression and play a structural and potential regulatory role in the centromere. To prevent retrotransposon activity, eukaryotic cells have evolved overlapping mechanisms to repress transposition. Plants are an excellent system for studying the mechanisms of LTR retrotransposon inhibition such as DNA methylation and small RNA-mediated degradation of retrotransposon transcripts. However, analysis of these multi-copy, mobile elements is considerably more difficult than analysis of single-copy genes located in stable regions of the genome. In this chapter we outline methods for analyzing the progress of LTR retrotransposons through their replication cycle in plants. We describe a mixture of traditional molecular biology experiments, such as Southern, Northern, and Western blotting, in addition to nontraditional techniques designed to take advantage of the specific mechanism of LTR retrotransposition.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Schnable PS et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Sharma A, Schneider KL, Presting GG (2008) Sustained retrotransposition is mediated by nucleotide deletions and interelement recombinations. Proc Natl Acad Sci U S A 105:15470–15474
Bennetzen JL (2002) Mechanisms and rates of genome expansion and contraction in flowering plants. Genetica 115:29–36
Slotkin RK, Martienssen R (2007) Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet 8:272–285
Lindroth AM, Cao X, Jackson JP, Zilberman D, McCallum CM, Henikoff S, Jacobsen SE (2001) Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science 292:2077–2080
Zilberman D, Cao X, Jacobsen SE (2003) ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299:716–719
Lippman Z, Martienssen R (2004) The role of RNA interference in heterochromatic silencing. Nature 431:364–370
Mirouze M, Reinders J, Bucher E, Nishimura T, Schneeberger K, Ossowski S, Cao J, Weigel D, Paszkowski J, Mathieu O (2009) Selective epigenetic control of retrotransposition in Arabidopsis. Nature 461:427–430
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE (2008) Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452:215–219
Lister R, O’Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH, Ecker JR (2008) Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133:523–536
Lisch D, Chomet P, Freeling M (1995) Genetic characterization of the Mutator system in maize: behavior and regulation of Mu transposons in a minimal line. Genetics 139:1777–1796
Kato M, Miura A, Bender J, Jacobsen SE, Kakutani T (2003) Role of CG and non-CG methylation in immobilization of transposons in Arabidopsis. Curr Biol 13:421–426
Cao X, Jacobsen SE (2002) Locus-specific control of asymmetric and CpNpG methylation by the DRM and CMT3 methyltransferase genes. Proc Natl Acad Sci U S A 99:16491–16498
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11:204–220
Huettel B, Kanno T, Daxinger L, Bucher E, van der Winden J, Matzke AJ, Matzke M (2007) RNA-directed DNA methylation mediated by DRD1 and Pol IVb: a versatile pathway for transcriptional gene silencing in plants. Biochim Biophys Acta 1769:358–374
Vaillant I, Schubert I, Tourmente S, Mathieu O (2006) MOM1 mediates DNA-methylation-independent silencing of repetitive sequences in Arabidopsis. EMBO Rep 7:1273–1278
Lippman Z, May B, Yordan C, Singer T, Martienssen R (2003) Distinct mechanisms determine transposon inheritance and methylation via small interfering RNA and histone modification. PLoS Biol 1:e67
Gruntman E, Qi Y, Slotkin RK, Roeder T, Martienssen RA, Sachidanandam R (2008) Kismeth: analyzer of plant methylation states through bisulfite sequencing. BMC Bioinforma 9:371
Slotkin RK, Vaughn M, Borges F, Tanurdzic M, Becker JD, Feijo JA, Martienssen RA (2009) Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136:461–472
Teixeira FK, Heredia F, Sarazin A, Roudier F, Boccara M, Ciaudo C, Cruaud C, Poulain J, Berdasco M, Fraga MF, Voinnet O, Wincker P, Esteller M, Colot V (2009) A role for RNAi in the selective correction of DNA methylation defects. Science 323:1600–1604
Johnson L, Cao X, Jacobsen S (2002) Interplay between two epigenetic marks. DNA methylation and histone H3 lysine 9 methylation. Curr Biol 12:1360–1367
Jackson JP, Lindroth AM, Cao X, Jacobsen SE (2002) Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416:556–560
Jacob Y, Stroud H, Leblanc C, Feng S, Zhuo L, Caro E, Hassel C, Gutierrez C, Michaels SD, Jacobsen SE (2010) Regulation of heterochromatic DNA replication by histone H3 lysine 27 methyltransferases. Nature 466:987–991
Gendrel AV, Lippman Z, Martienssen R, Colot V (2005) Profiling histone modification patterns in plants using genomic tiling microarrays. Nat Methods 2:213–218
Sundaresan V, Springer P, Volpe T, Haward S, Jones JD, Dean C, Ma H, Martienssen R (1995) Patterns of gene action in plant development revealed by enhancer trap and gene trap transposable elements. Genes Dev 9:1797–1810
Henderson IR, Zhang X, Lu C, Johnson L, Meyers BC, Green PJ, Jacobsen SE (2006) Dissecting Arabidopsis thaliana DICER function in small RNA processing, gene silencing and DNA methylation patterning. Nat Genet 38:721–725
Pall GS, Codony-Servat C, Byrne J, Ritchie L, Hamilton A (2007) Carbodiimide-mediated cross-linking of RNA to nylon membranes improves the detection of siRNA, miRNA and piRNA by northern blot. Nucleic Acids Res 35:e60
Kasschau KD, Fahlgren N, Chapman EJ, Sullivan CM, Cumbie JS, Givan SA, Carrington JC (2007) Genome-wide profiling and analysis of Arabidopsis siRNAs. PLoS Biol 5:e57
Takeda S, Sugimoto K, Kakutani T, Hirochika H (2001) Linear DNA intermediates of the Tto1 retrotransposon in Gag particles accumulated in stressed tobacco and Arabidopsis thaliana. Plant J 28:307–317
Bachmair A, Garber K, Takeda S, Sugimoto K, Kakutani T, Hirochika H (2004) Biochemical Analysis of Long Terminal Repeat Retrotransposons. In: Mobile Genetic Elements, Methods in Molecular Biology, eds. Miller WJ and Capy P. Humana Press 260:73–82
Maudru T, Peden K (1997) Elimination of background signals in a modified polymerase chain reaction-based reverse transcriptase assay. J Virol Methods 66:247–261
Lovatt A, Black J, Galbraith D, Doherty I, Moran MW, Shepherd AJ, Griffen A, Bailey A, Wilson N, Smith KT (1999) High throughput detection of retrovirus-associated reverse transcriptase using an improved fluorescent product enhanced reverse transcriptase assay and its comparison to conventional detection methods. J Virol Methods 82:185–200
Lucas H, Feuerbach F, Kunert K, Grandbastien MA, Caboche M (1995) RNA-mediated transposition of the tobacco retrotransposon Tnt1 in Arabidopsis thaliana. EMBO J 14:2364–2373
Kikuchi K, Terauchi K, Wada M, Hirano HY (2003) The plant MITE mPing is mobilized in anther culture. Nature 421:167–170
Fukai E, Umehara Y, Sato S, Endo M, Kouchi H, Hayashi M, Stougaard J, Hirochika H (2010) Derepression of the plant Chromovirus LORE1 induces germline transposition in regenerated plants. PLoS Genet 6:e1000868
Lund J, Tedesco P, Duke K, Wang J, Kim SK, Johnson TE (2002) Transcriptional profile of aging in C. elegans. Curr Biol 12:1566–1573
Casa AM, Nagel A, Wessler SR (2004) MITE display. Methods Mol Biol 260:175–188
Jiang N, Bao Z, Zhang X, Hirochika H, Eddy SR, McCouch SR, Wessler SR (2003) An active DNA transposon family in rice. Nature 421:163–167
Rangwala SH, Kazazian HH Jr (2009) The L1 retrotransposition assay: a retrospective and toolkit. Methods 49:219–226
Xie Y, Rosser JM, Thompson TL, Boeke JD, An W (2011) Characterization of L1 retrotransposition with high-throughput dual-luciferase assays. Nucleic Acids Res 39:e16
Sabot F, Schulman AH (2006) Parasitism and the retrotransposon life cycle in plants: a hitchhiker’s guide to the genome. Heredity 97:381–388
Acknowledgements
The authors gratefully acknowledge Andrea McCue and Savageethi Nuthikattu for helpful discussions and critical comments. This work was supported by the National Science Foundation grant MCB-1020499.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media, New York
About this protocol
Cite this protocol
DeFraia, C., Slotkin, R.K. (2014). Analysis of Retrotransposon Activity in Plants. In: Spillane, C., McKeown, P. (eds) Plant Epigenetics and Epigenomics. Methods in Molecular Biology, vol 1112. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-773-0_13
Download citation
DOI: https://doi.org/10.1007/978-1-62703-773-0_13
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-772-3
Online ISBN: 978-1-62703-773-0
eBook Packages: Springer Protocols