Abstract
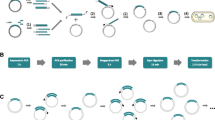
In-Fusion™ cloning is a flexible DNA ligase-independent cloning technology that has wide-ranging uses in molecular biology. In this chapter we describe the protocols used in the OPPF-UK to design and construct expression vectors using In-Fusion™. Our method for small scale expression screening in Escherichia coli of constructs generated by In-Fusion™ is also outlined.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Berrow NS, Bussow K, Coutard B et al (2006) Recombinant protein expression and solubility screening in Escherichia coli: a comparative study. Acta Crystallogr D Biol Crystallogr 62: 1218–1226
Savitsky P, Bray J, Cooper CD et al (2010) High-throughput production of human proteins for crystallization: the SGC experience. J Struct Biol 172:3–13
Graslund S, Nordlund P, Weigelt J et al (2008) Protein production and purification. Nat Methods 5:135–146
Irwin CR, Farmer A, Willer DO et al (2012) In-fusion(R) cloning with vaccinia virus DNA polymerase. Methods Mol Biol 890:23–35
Hamilton MD, Nuara AA, Gammon DB et al (2007) Duplex strand joining reactions catalyzed by vaccinia virus DNA polymerase. Nucleic Acids Res 35:143–151
Aslandis C, de Jong PJ (1990) Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res 18:6069–6074
Haun RS, Serventi IM, Moss J (1992) Rapid, reliable ligation-independent cloning of PCR products using modified plasmid vectors. Biotechniques 13:515–518
Li MZ, Elledge SJ (2012) SLIC: a method for sequence- and ligation-independent cloning. Methods Mol Biol 852:51–59
Li MZ, Elledge SJ (2007) Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat Methods 4: 251–256
Jeong JY, Yim HS, Ryu JY et al (2012) One-step sequence- and ligation-independent cloning as a rapid and versatile cloning method for functional genomics studies. Appl Environ Microbiol 78:5440–5443
Gibson DG, Young L, Chuang RY et al (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6: 343–345
Gibson DG, Smith HO, Hutchison CA III et al (2010) Chemical synthesis of the mouse mitochondrial genome. Nat Methods 7:901–903
Berrow NS, Alderton D, Sainsbury S et al (2007) A versatile ligation-independent cloning method suitable for high-throughput expression screening applications. Nucleic Acids Res 35:e45
Berrow NS, Alderton D, Owens RJ (2009) The precise engineering of expression vectors using high-throughput In-Fusion PCR cloning. Methods Mol Biol 498:75–90
Bird LE (2011) High throughput construction and small scale expression screening of multi-tag vectors in Escherichia coli. Methods 55:29–37
Chen JH, Jung JW, Wang Y et al (2010) Immunoproteomics profiling of blood stage Plasmodium vivax infection by high-throughput screening assays. J Proteome Res 9:6479–6489
Howland SW, Poh CM, Renia L (2011) Directional, seamless, and restriction enzyme-free construction of random-primed complementary DNA libraries using phosphorothioate-modified primers. Anal Biochem 416:141–143
Nettleship JE, Ren J, Rahman N et al (2008) A pipeline for the production of antibody fragments for structural studies using transient expression in HEK 293T cells. Protein Expr Purif 62:83–89
Au K, Berrow NS, Blagova E et al (2006) Application of high-throughput technologies to a structural proteomics-type analysis of Bacillus anthracis. Acta Crystallogr D Biol Crystallogr 62:1267–1275
Marsischky G, LaBaer J (2004) Many paths to many clones: a comparative look at high-throughput cloning methods. Genome Res 14:2020–2028
Zhu B, Cai G, Hall EO et al (2007) In-fusion assembly: seamless engineering of multidomain fusion proteins, modular vectors, and mutations. Biotechniques 43:354–359
Zhou B, Donnelly ME, Scholes DT et al (2009) Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza a viruses. J Virol 83:10309–10313
Sleight SC, Bartley BA, Lieviant JA et al (2010) Designing and engineering evolutionary robust genetic circuits. J Biol Eng 4:12
Sleight SC, Bartley BA, Lieviant JA et al (2010) In-Fusion BioBrick assembly and re-engineering. Nucleic Acids Res 38:2624–2636
Alzari PM, Berglund H, Berrow NS et al (2006) Implementation of semi-automated cloning and prokaryotic expression screening: the impact of SPINE. Acta Crystallogr D Biol Crystallogr 62:1103–1113
Esposito D, Chatterjee DK (2006) Enhancement of soluble protein expression through the use of fusion tags. Curr Opin Biotechnol 17:353–358
Marblestone JG, Edavettal SC, Lim Y et al (2006) Comparison of SUMO fusion technology with traditional gene fusion systems: enhanced expression and solubility with SUMO. Protein Sci 15:182–189
Ohana RF, Encell LP, Zhao K et al (2009) HaloTag7: a genetically engineered tag that enhances bacterial expression of soluble proteins and improves protein purification. Protein Expr Purif 68:110–120
Chudakov DM, Matz MV, Lukyanov S et al (2010) Fluorescent proteins and their applications in imaging living cells and tissues. Physiol Rev 90:1103–1163
Li C, Schwabe JW, Banayo E et al (1997) Coexpression of nuclear receptor partners increases their solubility and biological activities. Proc Natl Acad Sci U S A 94:2278–2283
Alexandrov A, Vignali M, LaCount DJ et al (2004) A facile method for high-throughput co-expression of protein pairs. Mol Cell Proteomics 3:934–938
Scheich C, Kummel D, Soumailakakis D et al (2007) Vectors for co-expression of an unrestricted number of proteins. Nucleic Acids Res 35:e43
Kholod N, Mustelin T (2001) Novel vectors for co-expression of two proteins in E. coli. Biotechniques 31(322–323):326–328
Yang W, Zhang L, Lu Z et al (2001) A new method for protein coexpression in Escherichia coli using two incompatible plasmids. Protein Expr Purif 22:472–478
Hinnebusch AG (2006) eIF3: a versatile scaffold for translation initiation complexes. Trends Biochem Sci 31:553–562
Busso D, Peleg Y, Heidebrecht T et al (2011) Expression of protein complexes using multiple Escherichia coli protein co-expression systems: a benchmarking study. J Struct Biol 175:159–170
Economou A, Christie PJ, Fernandez RC et al (2006) Secretion by numbers: protein traffic in prokaryotes. Mol Microbiol 62:308–319
Desvaux M, Hebraud M, Talon R et al (2009) Secretion and subcellular localizations of bacterial proteins: a semantic awareness issue. Trends Microbiol 17:139–145
Choi JH, Lee SY (2004) Secretory and extracellular production of recombinant proteins using Escherichia coli. Appl Microbiol Biotechnol 64:625–635
Mergulhao FJ, Summers DK, Monteiro GA (2005) Recombinant protein secretion in Escherichia coli. Biotechnol Adv 23:177–202
Binet R, Letoffe S, Ghigo JM et al (1997) Protein secretion by Gram-negative bacterial ABC exporters—a review. Gene 192:7–11
Steiner D, Forrer P, Stumpp MT et al (2006) Signal sequences directing cotranslational translocation expand the range of proteins amenable to phage display. Nat Biotechnol 24:823–831
Benoit RM, Wilhelm RN, Scherer-Becker D et al (2006) An improved method for fast, robust, and seamless integration of DNA fragments into multiple plasmids. Protein Expr Purif 45:66–71
Ruther U (1980) Construction and properties of a new cloning vehicle, allowing direct screening for recombinant plasmids. Mol Gen Genet 178:475–477
Acknowledgements
The Oxford Protein Production Facility-UK is supported by the UK Medical Research Council and the Biotechnology and Biology Research Council (MRC Grant MR/K018779/1). We thank Jo Nettleship for helpful discussions on E. coli secretion vectors.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media, New York
About this protocol
Cite this protocol
Bird, L.E., Rada, H., Flanagan, J., Diprose, J.M., Gilbert, R.J.C., Owens, R.J. (2014). Application of In-Fusion™ Cloning for the Parallel Construction of E. coli Expression Vectors. In: Valla, S., Lale, R. (eds) DNA Cloning and Assembly Methods. Methods in Molecular Biology, vol 1116. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-764-8_15
Download citation
DOI: https://doi.org/10.1007/978-1-62703-764-8_15
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-763-1
Online ISBN: 978-1-62703-764-8
eBook Packages: Springer Protocols