Abstract
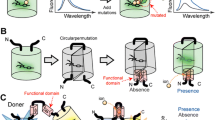
To track the activity of cellular signaling molecules within the endogenous cellular environment, researchers have developed a diverse set of genetically encodable fluorescent biosensors. These sensors, which can be targeted to specific subcellular regions to monitor specific pools of a given signaling molecule in real time, rely upon conformational changes in a sensor domain to alter the photophysical properties of green fluorescent protein (GFP) family members. In this introductory chapter, we first discuss the properties of GFP family members before turning our attention to the design and application of genetically encodable fluorescent biosensors to live cell imaging.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Tsien RY (1998) The green fluorescent protein. Annu Rev Biochem 67:509–544
Zimmer M (2002) Green fluorescent protein (GFP): applications, structure, and related photophysical behavior. Chem Rev 102:759–781
Remington SJ (2006) Fluorescent proteins: maturation, photochemistry and photophysics. Curr Opin Struct Biol 16:714–721
Heim R, Tsien RY (1996) Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr Biol 6:178–182
Miyawaki A, Griesbeck O, Heim R, Tsien RY (1999) Dynamic and quantitative Ca2+ measurements using improved cameleons. Proc Natl Acad Sci U S A 96:2135–2140
Davidson MW, Campbell RE (2009) Engineered fluorescent proteins: innovations and applications. Nat Methods 6:713–717
Shaner NC, Patterson GH, Davidson MW (2007) Advances in fluorescent protein technology. J Cell Sci 120:4247–4260
Newman RH, Fosbrink MD, Zhang J (2011) Genetically encodable fluorescent biosensors for tracking signaling dynamics in living cells. Chem Rev 111:3614–3666
Sample V, Newman RH, Zhang J (2009) The structure and function of fluorescent proteins. Chem Soc Rev 38:2852–2864
Day RN, Davidson MW (2009) The fluorescent protein palette: tools for cellular imaging. Chem Soc Rev 38:2887–2921
Pakhomov AA, Martynov VI (2008) GFP family: structural insights into spectral tuning. Chem Biol 15:755–764
Chattoraj M, King BA, Bublitz GU, Boxer SG (1996) Ultra-fast excited state dynamics in green fluorescent protein: multiple states and proton transfer. Proc Natl Acad Sci U S A 93:8362–8367
Brejc K, Sixma TK, Kitts PA, Kain SR et al (1997) Structural basis for dual excitation and photoisomerization of the Aequorea victoria green fluorescent protein. Proc Natl Acad Sci U S A 94:2306–2311
Ormo M, Cubitt AB, Kallio K, Gross LA et al (1996) Crystal structure of the Aequorea victoria green fluorescent protein. Science 273:1392–1395
Shaner NC, Steinbach PA, Tsien RY (2005) A guide to choosing fluorescent proteins. Nat Methods 2:905–909
Abad MF, Di Benedetto G, Magalhaes PJ, Filippin L et al (2004) Mitochondrial pH monitored by a new engineered green fluorescent protein mutant. J Biol Chem 279:11521–11529
Kneen M, Farinas J, Li Y, Verkman AS (1998) Green fluorescent protein as a noninvasive intracellular pH indicator. Biophys J 74:1591–1599
Llopis J, McCaffery JM, Miyawaki A, Farquhar MG et al (1998) Measurement of cytosolic, mitochondrial, and Golgi pH in single living cells with green fluorescent proteins. Proc Natl Acad Sci U S A 95:6803–6808
Griesbeck O, Baird GS, Campbell RE, Zacharias DA et al (2001) Reducing the environmental sensitivity of yellow fluorescent protein. Mechanism and applications. J Biol Chem 276:29188–29194
Nagai T, Ibata K, Park ES, Kubota M et al (2002) A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat Biotechnol 20:87–90
Heim R, Prasher DC, Tsien RY (1994) Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc Natl Acad Sci U S A 91:12501–12504
Rizzo MA, Springer GH, Granada B, Piston DW (2004) An improved cyan fluorescent protein variant useful for FRET. Nat Biotechnol 22:445–449
Goedhart J, van Weeren L, Hink MA, Vischer NO et al (2010) Bright cyan fluorescent protein variants identified by fluorescence lifetime screening. Nat Methods 7:137–139
Klarenbeek JB, Goedhart J, Hink MA, Gadella TW et al (2011) A mTurquoise-based cAMP sensor for both FLIM and ratiometric read-out has improved dynamic range. PLoS One 6:e19170
Verkhusha VV, Lukyanov KA (2004) The molecular properties and applications of Anthozoa fluorescent proteins and chromoproteins. Nat Biotechnol 22:289–296
Shu X, Shaner NC, Yarbrough CA, Tsien RY et al (2006) Novel chromophores and buried charges control color in mFruits. Biochemistry 45:9639–9647
Chudakov DM, Lukyanov S, Lukyanov KA (2005) Fluorescent proteins as a toolkit for in vivo imaging. Trends Biotechnol 23:605–613
Wachter RM, Watkins JL, Kim H (2010) Mechanistic diversity of red fluorescence acquisition by GFP-like proteins. Biochemistry 49:7417–7427
Merzlyak EM, Goedhart J, Shcherbo D, Bulina ME et al (2007) Bright monomeric red fluorescent protein with an extended fluorescence lifetime. Nat Methods 4:555–557
Shaner NC, Lin MZ, McKeown MR, Steinbach PA et al (2008) Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat Methods 5:545–551
Shcherbo D, Merzlyak EM, Chepurnykh TV, Fradkov AF et al (2007) Bright far-red fluorescent protein for whole-body imaging. Nat Methods 4:741–746
Shcherbo D, Murphy CS, Ermakova GV, Solovieva EA et al (2009) Far-red fluorescent tags for protein imaging in living tissues. Biochem J 418:567–574
Lin MZ, McKeown MR, Ng HL, Aguilera TA et al (2009) Autofluorescent proteins with excitation in the optical window for intravital imaging in mammals. Chem Biol 16:1169–1179
Lam AJ, St Pierre F, Gong Y, Marshall JD et al (2012) Improving FRET dynamic range with bright green and red fluorescent proteins. Nat Methods 9:1005–1012
Meyer AJ, Dick TP (2010) Fluorescent protein-based redox probes. Antioxid Redox Signal 13:621–650
Hanson GT, Aggeler R, Oglesbee D, Cannon M et al (2004) Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J Biol Chem 279:13044–13053
Hung YP, Albeck JG, Tantama M, Yellen G (2011) Imaging cytosolic NADH-NAD(+) redox state with a genetically encoded fluorescent biosensor. Cell Metab 14:545–554
Dittmer PJ, Miranda JG, Gorski JA, Palmer AE (2009) Genetically encoded sensors to elucidate spatial distribution of cellular zinc. J Biol Chem 284:16289–16297
Park JG, Qin Y, Galati DF, Palmer AE (2012) New sensors for quantitative measurement of mitochondrial Zn(2+). ACS Chem Biol 7:1636–1640
Qin Y, Dittmer PJ, Park JG, Jansen KB et al (2011) Measuring steady-state and dynamic endoplasmic reticulum and Golgi Zn2+ with genetically encoded sensors. Proc Natl Acad Sci U S A 108:7351–7356
Miyawaki A, Llopis J, Heim R, McCaffery JM et al (1997) Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388:882–887
Nagai T, Sawano A, Park ES, Miyawaki A (2001) Circularly permuted green fluorescent proteins engineered to sense Ca2+. Proc Natl Acad Sci U S A 98:3197–3202
Nagai T, Yamada S, Tominaga T, Ichikawa M et al (2004) Expanded dynamic range of fluorescent indicators for Ca(2+) by circularly permuted yellow fluorescent proteins. Proc Natl Acad Sci U S A 101:10554–10559
Ohkura M, Matsuzaki M, Kasai H, Imoto K et al (2005) Genetically encoded bright Ca2+ probe applicable for dynamic Ca2+ imaging of dendritic spines. Anal Chem 77:5861–5869
Palmer AE, Giacomello M, Kortemme T, Hires SA et al (2006) Ca2+ indicators based on computationally redesigned calmodulin-peptide pairs. Chem Biol 13:521–530
Tallini YN, Ohkura M, Choi BR, Ji G et al (2006) Imaging cellular signals in the heart in vivo: cardiac expression of the high-signal Ca2+ indicator GCaMP2. Proc Natl Acad Sci U S A 103:4753–4758
Zhang J, Allen MD (2007) FRET-based biosensors for protein kinases: illuminating the kinome. Mol Biosyst 3:759–765
Zhang J, Ma Y, Taylor SS, Tsien RY (2001) Genetically encoded reporters of protein kinase A activity reveal impact of substrate tethering. Proc Natl Acad Sci U S A 98:14997–15002
Allen MD, Zhang J (2006) Subcellular dynamics of protein kinase A activity visualized by FRET-based reporters. Biochem Biophys Res Commun 348:716–721
Depry C, Allen MD, Zhang J (2011) Visualization of PKA activity in plasma membrane microdomains. Mol Biosyst 7:52–58
Zhang J, Hupfeld CJ, Taylor SS, Olefsky JM et al (2005) Insulin disrupts beta-adrenergic signalling to protein kinase A in adipocytes. Nature 437:569–573
Zhou X, Herbst-Robinson KJ, Zhang J (2012) Visualizing dynamic activities of signaling enzymes using genetically encodable FRET-based biosensors from designs to applications. Methods Enzymol 504:317–340
Knopfel T, Tomita K, Shimazaki R, Sakai R (2003) Optical recordings of membrane potential using genetically targeted voltage-sensitive fluorescent proteins. Methods 30:42–48
Lundby A, Mutoh H, Dimitrov D, Akemann W et al (2008) Engineering of a genetically encodable fluorescent voltage sensor exploiting fast Ci-VSP voltage-sensing movements. PLoS One 3:e2514
Zhang J, Campbell RE, Ting AY, Tsien RY (2002) Creating new fluorescent probes for cell biology. Nat Rev Mol Cell Biol 3:906–918
Komatsu N, Aoki K, Yamada M, Yukinaga H et al (2011) Development of an optimized backbone of FRET biosensors for kinases and GTPases. Mol Biol Cell 22:4647–4656
Yang F, Moss LG, Phillips GN Jr (1996) The molecular structure of green fluorescent protein. Nat Biotechnol 14:1246–1251
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Newman, R.H., Zhang, J. (2014). The Design and Application of Genetically Encodable Biosensors Based on Fluorescent Proteins. In: Zhang, J., Ni, Q., Newman, R. (eds) Fluorescent Protein-Based Biosensors. Methods in Molecular Biology, vol 1071. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-622-1_1
Download citation
DOI: https://doi.org/10.1007/978-1-62703-622-1_1
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-621-4
Online ISBN: 978-1-62703-622-1
eBook Packages: Springer Protocols