Abstract
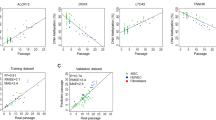
Somatic cells change continuously during culture expansion—long-term culture evokes increasing cell size, declining differentiation potential, and ultimate cell cycle arrest upon senescence. These changes are of particular relevance for cellular therapy which necessitates standardized products and reliable quality control. Recently, replicative senescence has been shown to be associated with highly reproducible epigenetic modifications. Here, we describe a simple method to track the state of senescence in mesenchymal stromal cells (MSCs) or fibroblasts by monitoring continuous DNA methylation (DNAm) changes at specific sites in the genome. Six CpG sites have been identified which reveal either linear hypermethylation or hypomethylation with respect to the number of cumulative population doublings (cPDs). Conversely, the DNAm level at these CpG sites can be analyzed—for example, by pyrosequencing of bisulfite-converted DNA—and then used for linear regression models to predict cPDs. Our method provides an epigenetic biomarker to determine the state of senescence in cell preparations.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wagner W, Horn P, Castoldi M et al (2008) Replicative senescence of mesenchymal stem cells—a continuous and organized process. PLoS One 5:e2213
Hayflick L (1965) The limited in vitro lifetime of human diploid cell strains. Exp Cell Res 37:614–636
Kang TW, Yevsa T, Woller N et al (2011) Senescence surveillance of pre-malignant hepatocytes limits liver cancer development. Nature 479:547–551
Campisi J, d’Adda di FF (2007) Cellular senescence: when bad things happen to good cells. Nat Rev Mol Cell Biol 8:729–740
Kiyono T, Foster SA, Koop JI, McDougall JK, Galloway DA, Klingelhutz AJ (1998) Both Rb/p16INK4a inactivation and telomerase activity are required to immortalize human epithelial cells. Nature 396:84–88
Bork S, Pfister S, Witt H et al (2010) DNA methylation pattern changes upon long-term culture and aging of human mesenchymal stromal cells. Aging Cell 9:54–63
Schellenberg A, Lin Q, Schueler H et al (2011) Replicative senescence of mesenchymal stem cells causes DNA-methylation changes which correlate with repressive histone marks. Aging (Albany NY) 3:873–888
Koch CM, Joussen S, Schellenberg A, Lin Q, Zenke M, Wagner W (2012) Monitoring of cellular senescence by DNA-methylation at specific CpG sites. Aging Cell 11:366–369
Koch C, Suschek CV, Lin Q et al (2011) Specific age-associated DNA methylation changes in human dermal fibroblasts. PLoS One 6:e16679
Koch C, Reck K, Shao K et al (2012) Pluripotent stem cells escape from senescence-associated DNA methylation changes. Genome Res 23(2):248–259
Roobrouck VD, Ulloa-Montoya F, Verfaillie CM (2008) Self-renewal and differentiation capacity of young and aged stem cells. Exp Cell Res 314:1937–1944
Wagner W, Bork S, Lepperdinger G et al (2010) How to track cellular aging of mesenchymal stromal cells. Aging (Albany NY) 2:224–230
Wagner W, Ho AD, Zenke M (2010) Different facets of aging in human mesenchymal stem cells. Tissue Eng Part B Rev 16:445–453
Schellenberg A, Stiehl T, Horn P et al (2012) Population dynamics of mesenchymal stromal cells during culture expansion. Cytotherapy 14:401–411
Dimri GP, Lee X, Basile G et al (1995) A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proc Natl Acad Sci USA 92:9363–9367
Debacq-Chainiaux F, Erusalimsky JD, Campisi J, Toussaint O (2009) Protocols to detect senescence-associated beta-galactosidase (SA-betagal) activity, a biomarker of senescent cells in culture and in vivo. Nat Protoc 4:1798–1806
Schallmoser K, Bartmann C, Rohde E et al (2010) Replicative senescence-associated gene expression changes in mesenchymal stromal cells are similar under different culture conditions. Haematologica 95:867–874
Pavlidis P, Noble WS (2001) Analysis of strain and regional variation in gene expression in mouse brain. Genome Biol 2:42
Bocklandt S, Lin W, Sehl ME et al (2011) Epigenetic predictor of age. PLoS One 6:e14821
Koch CM, Wagner W (2011) Epigenetic-aging-signature to determine age in different tissues. Aging (Albany NY) 3:1018–1027
Shao K, Koch CM, Gupta MK et al (2013) Induced pluripotent mesenchymal stromal cell clones retain donor-derived differences in DNA methylation profiles. Mol Ther 21(1):240–250
Schallmoser K, Bartmann C, Rohde E et al (2007) Human platelet lysate can replace fetal bovine serum for clinical-scale expansion of functional mesenchymal stromal cells. Transfusion 47:1436–1446
Horn P, Bokermann G, Cholewa D et al (2010) Comparison of individual platelet lysates for isolation of human mesenchymal stromal cells. Cytotherapy 12:888–898
Bibikova M, Le J, Barnes R et al (2009) Genome-wide DNA methylation profiling using Infinium assay. Epigenomics 1:177–200
Ehrich M, Nelson MR, Stanssens P et al (2005) Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc Natl Acad Sci USA 102:15785–15790
Cholewa D, Stiehl T, Schellenberg A et al (2011) Expansion of adipose mesenchymal stromal cells is affected by human platelet lysate and plating density. Cell Transplant 20:1409–1922
Lohmann M, Walenda G, Hemeda H et al (2012) Donor Age of human platelet lysate affects proliferation and differentiation of mesenchymal stem cells. PLoS One 7:e37839
Bibikova M, Barnes B, Tsan C et al (2011) High density DNA methylation array with single CpG site resolution. Genomics 98:288–295
Acknowledgments
We thank Anne Schellenberg for critical reading of the manuscript and Qiong Lin for contribution of the multivariate model. RWTH Aachen Medical School has applied for a patent application for the above mentioned method. This work was supported by the excellence initiative of the German federal and state governments within the START Program of the Faculty of Medicine, RWTH Aachen (W.W.), by the Stem Cell Network North Rhine Westphalia (W.W.) and by the Else-Kröner Fresenius Stiftung (W.W. and C.K.).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, New York
About this protocol
Cite this protocol
Koch, C.M., Wagner, W. (2013). Epigenetic Biomarker to Determine Replicative Senescence of Cultured Cells. In: Tollefsbol, T. (eds) Biological Aging. Methods in Molecular Biology, vol 1048. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-556-9_20
Download citation
DOI: https://doi.org/10.1007/978-1-62703-556-9_20
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-555-2
Online ISBN: 978-1-62703-556-9
eBook Packages: Springer Protocols