Abstract
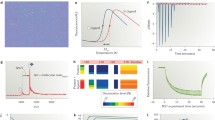
Discovering small-molecule chemical probes of protein function has great potential to elucidate biological pathways and to provide early-stage proof-of-concept for target validation. Discovery of such probes therefore underpins many of the chemical biology and drug discovery efforts in both academia and the pharmaceutical industry. The process generally begins with screening small molecules to identify bona fide “hits” that bind non-covalently to a target protein. This chapter is concerned with the application of biophysical and structural techniques to small-molecule ligand screening, and with the validation of hits from both structural (binding mode) and energetic (binding affinity) stand-points. The methods discussed include differential scanning fluorimetry (thermal shift), fluorescence polarization (FP), surface plasmon resonance, ligand-observed NMR spectroscopy, isothermal titration calorimetry, and protein X-ray crystallography. The principles of these techniques and the fundamental nature of the observables used to detect macromolecule-ligand binding are briefly outlined. The practicalities, advantages, and disadvantages of each technique are described, particularly in the context of detecting weak affinities, as relevant to fragment screening. Fluorescence-based methods, which offer an attractive combination of high throughput and low cost are discussed in detail. It is argued that applying a combination of different methods provides the most robust and effective way to identify high-quality starting points for follow-up medicinal chemistry and to build structure–activity relationships that better inform effective development of high-quality, cell-active chemical probes by structure-based drug design.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Crews CM (2010) Targeting the undruggable proteome: the small molecules of my dreams. Chem Biol 17:551–555
Schreiber SL (2005) Small molecules: the missing link in the central dogma. Nat Chem Biol 1:64–66
Frye SV (2010) The art of the chemical probe. Nat Chem Biol 6:159–161
Edwards AM, Bountra C, Kerr DJ, Willson TM (2009) Open access chemical and clinical probes to support drug discovery. Nat Chem Biol 5:436–440
Cole PA (2008) Chemical probes for histone-modifying enzymes. Nat Chem Biol 4:590–597
Hopkins AL, Groom CR (2002) The druggable genome. Nat Rev Drug Discov 1:727–730
Russ AP, Lampel S (2005) The druggable genome: an update. Drug Discov Today 10:1607–1610
Broach JR, Thorner J (1996) High-throughput screening for drug discovery. Nature 384:14–16
Spencer RW (1999) High-throughput screening of historic collections: observations on file size, biological targets, and file diversity. Biotechnol Bioeng 61:61–67
Macarron R, Banks MN, Bojanic D et al (2011) Impact of high-throughput screening in biomedical research. Nat Rev Drug Discov 10:188–195
Bleicher KH, Böhm H-J, Müller K, Alanine AI (2003) Hit and lead generation: beyond high-throughput screening. Nat Rev Drug Discov 2:369–378
Shoichet BK (2006) Interpreting steep dose–response curves in early inhibitor discovery. J Med Chem 49:7274–7277
Babaoglu K, Simeonov A, Irwin JJ et al (2008) Comprehensive mechanistic analysis of hits from high-throughput and docking screens against beta-lactamase. J Med Chem 51:2502–2511
McGovern SL, Caselli E, Grigorieff N et al (2002) A common mechanism underlying promiscuous inhibitors from virtual and high-throughput screening. J Med Chem 45:1712–1722
Jhoti H, Cleasby A, Verdonk M, Williams G (2007) Fragment-based screening using X-ray crystallography and NMR spectroscopy. Curr Opin Chem Biol 11:485–493
Ciulli A, Blundell TL, Abell C (2008) Discovery and extrapolation of fragment structures towards drug design. In: Stroud RM, Finer-Moore J (eds) Computational and structural approaches to drug discovery: ligand–protein interactions. The Royal Society of Chemistry, Cambridge
Shuker SB, Hajduk PJ, Meadows RP, Fesik SW (1996) Discovering high-affinity ligands for proteins: SAR by NMR. Science 274:1531–1534
Hann MM, Leach AR, Harper G (2001) Molecular complexity and its impact on the probability of finding leads for drug discovery. J Chem Inf Comput Sci 41:856–864
Blundell TL, Jhoti H, Abell C (2002) High-throughput crystallography for lead discovery in drug design. Nat Rev Drug Discov 1:45–54
Rees DC, Congreve M, Murray CW, Carr R (2004) Fragment-based lead discovery. Nat Rev Drug Discov 3:660–672
Hajduk PJ, Greer J (2007) A decade of fragment-based drug design: strategic advances and lessons learned. Nat Rev Drug Discov 6:211–219
Congreve M, Chessari G, Tisi D, Woodhead AJ (2008) Recent developments in fragment-based drug discovery. J Med Chem 51:3661–3680
Ciulli A, Abell C (2007) Fragment-based approaches to enzyme inhibition. Curr Opin Biotechnol 18:489–496
Murray CW, Rees DC (2009) The rise of fragment-based drug discovery. Nat Chem 1:187–192
Erlanson DA (2012) Introduction to fragment-based drug discovery. Top Curr Chem 317:1–32
Lundqvist T (2005) The devil is still in the details—driving early drug discovery forward with biophysical experimental methods. Curr Opin Drug Discov Devel 8:513–519
Ciulli A, Williams G, Smith AG, Blundell TL, Abell C (2006) Probing hot spots at protein–ligand binding sites: a fragment-based approach using biophysical methods. J Med Chem 49:4992–5000
Ericsson UB, Hallberg BM, Detitta GT, Dekker N, Nordlund P (2006) Thermofluor-based high-throughput stability optimization of proteins for structural studies. Anal Biochem 357:289–298
Cummings M, Farnum M, Nelen M (2006) Universal screening methods and applications of ThermoFluor®. J Biomol Screen 11:854–863
Lo M-C, Aulabaugh A, Jin G et al (2004) Evaluation of fluorescence-based thermal shift assays for hit identification in drug discovery. Anal Biochem 332:153–159
Kranz JK, Schalk-Hihi C (2011) Protein thermal shifts to identify low molecular weight fragments. Methods Enzymol 493:277–298
Reindl W, Strebhardt K, Berg T (2008) A high-throughput assay based on fluorescence polarization for inhibitors of the polo-box domain of polo-like kinase 1. Anal Biochem 383:205–209
Huang X (2003) Fluorescence polarization competition assay: the range of resolvable inhibitor potency is limited by the affinity of the fluorescent ligand. J Biomol Screen 8:34–38
Wiseman T, Williston S, Brandts JF, Lin LN (1989) Rapid measurement of binding constants and heats of binding using a new titration calorimeter. Anal Biochem 179:131–137
Turnbull WB, Daranas AH (2003) On the value of c: can low affinity systems be studied by isothermal titration calorimetry? J Am Chem Soc 125:14859–14866
Van der Merwe PA (2001) Surface plasmon resonance. In: Chowdhry B, Harding S (eds) Protein–ligand interactions: hydrodynamics and calorimetry. Oxford University Press, Oxford
Pellecchia M, Bertini I, Cowburn D et al (2008) Perspectives on NMR in drug discovery: a technique comes of age. Nat Rev Drug Discov 7(9):738–745
Lepre CA, Moore JM, Peng JW (2004) Theory and applications of NMR-based screening in pharmaceutical research. Chem Rev 104:3641–3676
Hajduk PJ, Sheppard G, Nettesheim D et al (1997) Discovery of potent nonpeptide inhibitors of stromelysin using SAR by NMR. J Am Chem Soc 119:5818–5827
Śledź P, Abell C, Ciulli A (2012) Ligand-observed NMR in fragment-based approaches. In: Bertini I, McGreevy K, Parigi G (eds) NMR of biomolecules: towards mechanistic systems biology. Wiley-VCH, Weinheim
Hajduk PJ, Olejniczak E, Fesik S (1997) One-dimensional relaxation- and diffusion-edited NMR methods for screening compounds that bind to macromolecules. J Am Chem Soc 119:12257–12261
Mayer M, Meyer B (1999) Characterization of ligand binding by saturation transfer difference NMR spectroscopy. Angew Chem Int Ed Engl 38:1784–1788
Dalvit C, Fogliatto G, Stewart A, Veronesi M, Stockman B (2001) WaterLOGSY as a method for primary NMR screening: practical aspects and range of applicability. J Biomol NMR 21:349–359
Nienaber VL, Richardson PL, Klighofer V et al (2000) Discovering novel ligands for macromolecules using X-ray crystallographic screening. Nat Biotechnol 18:1105–1108
Blundell TL, Abell C, Cleasby A et al (2002) High throughput X-ray crystallography for drug discovery. In: Flower DR (ed) Drug design: cutting edge approaches. The Royal Society of Chemistry, Cambridge
Niesen FH, Berglund H, Vedadi M (2007) The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat Protocols 2:2212–2221
Zhang J, Chung T, Oldenburg K (1999) A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J Biomol Screen 4:67–73
Berman HM, Westbrook J, Feng Z et al (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Philpott M, Yang J, Tumber T et al (2011) Bromodomain-peptide displacement assays for interactome mapping and inhibitor discovery. Mol BioSyst 7:2899–2908
Ciulli A, Scott DE, Ando M et al (2008) Inhibition of Mycobacterium tuberculosis pantothenate synthetase by analogues of the reaction intermediate. ChemBioChem 9:2606–2611
Hung AW, Silvestre HL, Wen S et al (2009) Application of fragment growing and fragment linking to the discovery of inhibitors of Mycobacterium tuberculosis pantothenate synthetase. Angew Chem Int Ed Engl 48:8452–8456
Buckley DL, Van Molle I, Gareiss PC et al (2012) Targeting the von Hippel-Lindau E3 ubiquitin ligase using small molecules to disrupt the VHL/HIF-1α interaction. J Am Chem Soc 134:4465–4468
Acknowledgments
The author wishes to thank several members of the Ciulli and Abell research groups (Department of Chemistry) and the Blundell research group (Department of Biochemistry) for productive collaboration and invaluable discussions over the years. The Ciulli laboratory is funded primarily but not exclusively by the UK Biotechnology and Biological Sciences Research Council (BBSRC). The author thanks the BBSRC for the award of a David Phillips Fellowship (BB/G023123/1). The author did not receive any financial contribution or writing assistance for the production of this chapter and declares no competing financial interests.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this protocol
Cite this protocol
Ciulli, A. (2013). Biophysical Screening for the Discovery of Small-Molecule Ligands. In: Williams, M., Daviter, T. (eds) Protein-Ligand Interactions. Methods in Molecular Biology, vol 1008. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-398-5_13
Download citation
DOI: https://doi.org/10.1007/978-1-62703-398-5_13
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-397-8
Online ISBN: 978-1-62703-398-5
eBook Packages: Springer Protocols