Abstract
Streptococcus mutans, a gram-positive facultative anaerobic bacterium, is one of the members of the mutans streptococci group, which is known to be a major pathogen of dental caries and a possible causative agent of infective endocarditis. Recent developments using molecular biological techniques enable to identify S. mutans in clinical specimens by PCR using species-specific sets of primers without direct isolation of the strains. S. mutans is classified into c/e/f/k serotypes based upon the chemical composition of the cell surface polysaccharide antigens. Serotypes in the clinical specimens can be easily determined by PCR using serotype-specific sets of primers. There are a large number of studies showing the association of the cell surface protein antigens and virulence for dental caries and infective endocarditis, whereas only a few reports document the molecular methods for identifying the strains with specific protein antigens related to the virulence of the specimens. The arbitrarily primed PCR (AP-PCR) method is one of the most common approaches for various types of studies with S. mutans, especially those investigating the strains transmitted from mothers to their children by comparing fingerprinting patterns. More recently, the multilocus sequence typing method, one of the molecular epidemiological approaches, was developed for S. mutans. Using this system, S. mutans could be classified into a large number of sequence types, which yields higher discriminatory power. Taken together, the molecular typing methods for S. mutans which have now been developed and widely used could lead to initiation of clinical applications as well as tools for basic research.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Properties of Streptococcus mutans
Streptococcus mutans, a Gram-positive facultative anaerobic bacterium, is generally known to be a pathogen of dental caries and also considered to be one of the oral streptococcal species that can cause infective endocarditis since it was reported to be recovered from 8 to 10% of patients with endocardial disease [1] (Fig. 9.1). S. mutans is one of the members of the “mutans streptococci” group, which also consists of Streptococcus sobrinus, Streptococcus cricetus, Streptococcus rattus, Streptococcus ferus, Streptococcus macacae, and Streptococcus downei. Mutans streptococci were previously classified into eight serotypes based on the chemical composition of their serotype-specific polysaccharides, among which five serotypes (a through e) were designated in 1970, followed by three additional serotypes ( f, g, h) determined during the next decade. S. mutans (c/e/f ) and S. sobrinus (d/g) are detected in humans, while S. cricetus (a) and S. rattus (b) strains are mainly identified in hamsters and rats, respectively. In addition, S. ferus (c) was reported to be isolated from rats, and S. macacae (c) and S. downei (h) were isolated from monkeys. Among the mutans streptococci, S. mutans is the most frequently identified in the human oral cavity, followed by S. sobrinus. On the other hand, the presence of the mutans streptococci in the oral cavities of animals is considered to result from the ingestion of sucrose contained in feed.
The serotype-specific polysaccharides of S. mutans are known to be composed of rhamnose-glucose polymers, with a backbone of rhamnose and side chains of glucose polymers [2]. The chemical linkage of each rhamnose unit is common in each c/e/f serotype (α-1,2 and α-1,3 repeatedly), while that for glucose side chains is different; α-1,2 for serotype c, β-1,2 for serotype e, and α-1,3 for serotype f. The distribution frequency of serotype c strains in the oral cavity is the highest with a rate of 70–80%, followed by serotype e (approximately 20%) and f (less than 5%). Strains, which could not be classified into any of c/e/f serotypes, have been described in the literature from subjects in several countries. However, details of the chemical composition of their serotype-specific polysaccharides have not been investigated. In 2004, the 9th serotype “k” was designated for the non-c/e/f serotype S. mutans strains which were isolated from the blood of the patients with bacteremia after tooth extraction and infective endocarditis complicated with subarachnoid hemorrhage [3].
Analysis of the distribution of serotype k strains revealed that the detection frequency in the oral cavity of Japanese children was 2–5%, which was shown to be consistent with that of Thai subjects [4]. In addition, several S. mutans strains of non-c/e/f serotypes isolated in Finland and UK were classified into serotype k [5, 6]. There have been several reports demonstrating the presence of non-c/e/f strains but an estimation of the presence of serotype k strains is not possible due to the lack of adequate description of the chemical composition of the serotype-specific rhamnose glucose polymers. On the other hand, a recent study in Chile considered the possibility of the presence of serotype k for the strains of non-c/e/f S. mutans serotypes isolated [7]. Taken together, these studies suggest that the serotype k strains are prevalent worldwide.
In addition to polysaccharide antigens, cell surface protein antigens are important for the virulence of S. mutans in dental caries. Among the various cell surface antigens, an approximately 190-kDa protein antigen (PA), glucosyltransferases (GTFs), and glucan-binding proteins (Gbps) are known to be major virulence factors for S. mutans [8–10]. PA, also referred to by other names (PAc, SpaP, antigen I/II, antigen B, SR, IF, P1, and MSL-1) is correlated with the sucrose-independent initial adhesion to tooth surfaces by the bacterium. In addition, GTFs are composed of three types (GTFB/GTFC/GTFD), and are known to be associated with sucrose-dependent adhesion. GTFB and GTFC, located on the cell surface, mainly synthesize water-insoluble glucans, which contain a high degree of branching of α-1,3-glucosidic linkages, whereas GTFD, released into the culture supernatant, produces water-soluble glucans that are predominantly linear polymers linked by α-1,6-glucosidic bonds, similar to dextran.
The complete genome of S. mutans strain UA159 (serotype c) was sequenced in 2002 by a team at the University of Oklahoma Health Sciences Center, which revealed that it was composed of 2,030,936 bp and contained 1,963 ORFs [11]. Detailed information is available in the Oral Pathogen Sequence Database provided by the Database Team at the Bioscience Division of Los Alamos National Laboratory (http://www.oralgen.lanl.gov/). Recently, the complete genome of another S. mutans strain NN2025 (serotype c) isolated from a Japanese child with severe dental caries was sequenced, which showed that it was composed of 2,013,587 bp and contained 1,869 ORFs [12]. When comparing the complete genomes of the two strains, core-genome was shown to be highly conserved, whereas a large genomic inversion between homologous ribosomal operons across the replication axis was identified. In addition, at least 25 different regions, which might be transferred following conjugation transfer or mediated by insertion elements, were identified in the two strains. At this moment, the complete genome of an additional S. mutans strain LJ23 (serotype k) is now being sequenced in order to identify the serotype-k specific genomic features by comparison with UA159 and NN2025.
2 Detection of S. mutans
In the early 1990s, DNA probe methods targeting gtfs and other genes were constructed to detect S. mutans. However, these are hampered by complex procedures and low sensitivity and one of the studies demonstrated that more than 300 pg of DNA and as many as 2 × 105 cells would be required for detection [13]. In the middle of the 1990s, the PCR-based approach for S. mutans detection was introduced, which was regarded as rapid, sensitive and relatively simple method. The spaP gene encoding the 190 kDa-protein antigen (PA) [14] and the dexA gene encoding extracellular dextranase were targeted for construction of species-specific sets of primers [15] (Table 9.1). The sensitivity was drastically increased with a lower limit of 1 pg of chromosomal DNA or 12 colony-forming units of S. mutans cells. Subsequently, several molecular methods for detecting S. mutans DNA in specimens from dental plaque and saliva have been reported. In addition, cardiovascular specimens, such as those from heart valves and atheromatous plaques, have been examined [16, 17]. The lower limit of detection for S. mutans DNA was reported to be detected in heart valve specimens extirpated from infective endocarditis patients, in which multiple species were identified in each specimen [18]. Thus, it is possible to speculate that S. mutans is one of the possible etiological agents and/or it is incidentally detected during transient bacteremia. As for atheromatous plaque, the high detection rate of S. mutans DNA does not necessarily mean a direct association of S. mutans with atheromatous plaque formation [19]. It is advantageous that PCR methods are very sensitive, however, careful interpretation of the results is required since bacterial DNA from nonviable and/or incidentally disseminated strains can be identified as positive reactions.
Table 9.2 summarizes the commonly used primer sets for S. mutans detection, among which those designed based on the gtf genes are widely used. Many oral streptococcal species reside in dental plaque, and the glucan synthesizing by the glucosyltransferases encoded by the gtf genes is one of the major factors in dental plaque formation [9]. The species-specific sets of primers have been constructed based on the differences in the nucleotide alignments of the gtf genes among several oral streptococci. The primer sets for S. mutans designed based on the gtfB or the gtfD genes are widely used [20, 21]. As for the methods using the gtfD sequence, the lower limit of detection for S. mutans DNA was reported to be 1.5 pg, indicating that this method is very sensitive. In addition, the methods for quantifying the numbers of S. mutans cells were developed using real-time PCR with the primer set SmF5 (5′-AGC CAT GCG CAA TCA ACA GGT T-3′) and SmR4 (5′-CGC AAC GCG AAC ATC TTG ATC AG-3′) targeting the gtfB gene [22]. It was reported that high levels of S. mutans in the parents is one of the important factors for vertical transmission into children [23]. Thus, this method could be one of the possible tools for identifying subjects with high risk for transmission. In addition, it could also be used to determine the number of the bacterial cells in cardiovascular specimens, which might lead to the identification of specific pathogenic bacterial species when multiple species are identified in each specimen by PCR.
The other molecular approach for detecting S. mutans is the restriction fragment length polymorphism (RFLP) of amplified 16S rRNA fragments, in which approximately 1,500 bp fragments are amplified with the universal primers 8UA and 1492R followed by digestion with HpaII [24]. In addition, another primer set based on 16S rRNA alignments [25] has been modified to amplify approximately 1,500 bp lengths of 16S rRNA followed by nested PCR amplification of an internal 282 bp region [24]. However, it was reported that false positive could result since the 16S rRNA sequence of the mutans streptococci and neighboring group are quite similar [26]. On the other hand, the determination of the entire 16S rRNA sequence amplified by the primer sets 8UA (5′-AGA GTT TGA TCC TGG CTC AG-3′) and 1540R (5′-AAG GAG GTG ATC CAG CC-3′) was used to confirm that the tested strain was truly a S. mutans [27]. This obviates misleading results although it is more time consuming and expensive than PCR methods with species-specific primer sets.
The groESL genes encoding 10-kDa (GroES) and 60-kDa (GroEL) heat shock proteins are reported to be ubiquitous and evolutionary highly conserved genes [28]. However, the groESL sequence is known to be less conserved as compared to the 16S rRNA sequence, indicating the possibility for its application in the differentiation of species with high degrees of similarity in their 16S rRNA sequences. The primer set for identification of mutans streptococci using the groESL genes was designed to amplify the fragment containing a region of partial groES, partial groEL, and the groES-EL spacer [29]. Since the nucleotide length of the groES-groEL spacer is varied among each species (111 bp for S. mutans, 218 bp for S. sobrinus, 200 bp for S. cricetus, 125 bp for S. rattus, and 310 bp for S. downei), the molecular sizes of the positive bands are different depending on the species. The other species-specific sets of primers were also constructed based on the nucleotide alignment of the ddlA gene encoding d-alanine:d-alanine ligase, which is known to be essential for bacterial cell wall synthesis [30]. In addition, a method for sequencing the internal fragment representing 85% of the sodA gene encoding manganese-dependent superoxide dismutase, was also developed, which discriminates between a large numbers of the various streptococcal strains [31].
The 16S–23S ribosomal RNA intergenic spacer (ITS) region, known to contain low levels of intraspecies variation and high levels of interspecies divergence, can also be used for speciation of S. mutans. PCR using universal primers 13BF (5′-GTG AAT ACG TTC CCG GGC CT-3′) and 6R (5′-GGG TTY CRT TCR GAA AT-3′) was designed based on the sequence of the 3′-region of the 16S rRNA gene and the 5′-portion of the 23S rRNA gene [32]. This amplifies fragments of variable sizes depending on the species. When the specimens contain S. mutans DNA, the amplified fragments include the 387 bp or 388 bp regions of ITS. Determination of the nucleotide alignment is initially performed, after which the identification of S. mutans is made by comparing the sequence of species-specific ITS and that of the specimens.
It is possible that the conventional methods for identification of bacterial species fail to identify phenotypically aberrant strains. On the other hand, the broad-range PCR and sequencing method, in which full and partial 16S rRNA nucleotide alignments are determined, is a reliable tool. As compared to PCR with species-specific sets of primers, the broad-range PCR and sequencing method enables the identification of multiple species in the specimens. Several primers for broad-range PCR methods have been developed and the amplified fragments are then sequenced, for comparison with those in the GenBank, EMBL, and DDBJ databases using the gapped BLASTN 2.0.5 program obtained from the National Center for Biotechnology Information server (http://www.ncbi.nlm.nih.gov/BLAST/) [33, 34] (Table 9.3). Identification to the species level is generally defined as a 16S rRNA sequence similarity of more than 99% with that of the prototype strain sequence in the databases. This approach is widely used for the investigation of bacterial profiles in the clinical specimens such as saliva and dental plaque as well as with cardiovascular specimens.
S. mutans strains are easily isolated from oral specimens, such as saliva and dental plaque, using selective medium and Mitis-salivarius agar plates containing bacitracin (MSB plates) [35]. On the other hand, S. mutans is occasionally isolated from the blood of the patients with bacteremia and infective endocarditis. However, it is difficult to isolate the strains from blood specimens once antibiotic treatments are initiated. The molecular methods enable detection of S. mutans using bacterial DNA extracted from even nonviable cells, which should be regarded as the most advantageous aspects in the use of molecular approaches.
3 Differentiation of S. mutans and S. sobrinus
S. mutans is highly prevalent in the oral cavity of humans, with a detection rate ranging from 74 to 94%, while S. sobrinus is known to be less prevalent [36]. In general, S. mutans is reported to be associated with coronal caries, whereas S. sobrinus is considered to be correlated to the lesions found on the smooth surfaces [36, 37]. It is generally known that subjects harboring both S. mutans and S. sobrinus have a significantly higher caries experience than those with only S. mutans [38, 39].
S. mutans and S. sobrinus are easily discriminated based on the rough and smooth colonies, respectively, on Mitis-salivarius agar plates. However, the GTF-defective strains of S. mutans are known to show smooth colony morphology on the agar although the distribution frequency is extremely low [40, 41]. Thus, it is possible that phenotypic variation prevents the appropriate discriminations of S. mutans and S. sobrinus. However, PCR-based approaches using species-specific primers for each species are not influenced by the phenotypic variations of S. mutans. The primer sets for S. mutans were designed based on the gtfB or gtfD genes of S. mutans, whereas gtfI was used for construction of the primer sets for S. sobrinus [20, 21].
One of the other molecular approaches is the PCR-RFLP method. In this method, the 4-kb region of the ribosomal RNA (rrn) operon, which most bacteria possess several copies for, with a high degree of homology, is amplified by PCR, followed by the comparison of the digestion patterns of the amplified fragments following HinfI, MboI, or TaqI digestion [42]. Another approach is the chromosomal DNA fingerprint (CDF) and arbitrarily primed (AP)-PCR methods [43]. The CDF method can discriminate between two major patterns after HaeIII digestion of the chromosomal DNA of each strain. One is designated as the CDF-1 group with restriction fragments equal to or greater than 6.6 kb in size and the other the CDF-2 group with fragments less than 6.6 kb. All of the tested S. mutans strains are classified as CDF-1, whereas most of the S. sobrinus strains were classified as CDF-2.
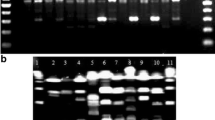
As for AP-PCR approach characterized by the short length of primers and low annealing temperatures, the primer OPA-02 (5′-TGC CGA GCT G-3′) is the most commonly used for the analysis of S. mutans, followed by OPA-13 (5′-CAG CAC CCA C-3′). This choice is primarily due to the appearance of readily identifiable electrophoretic products with the former compared to the other 40 sets of the primers [44] (Fig. 9.2). S. mutans and S. sobrinus strains showed similar patterns in each group, which are reported to consist of the major common amplified fragments of 782 bp and 1,070 bp, respectively. In addition, OPA-03 (5′-AGT CAG CCA C-3′), OPA-05 (5′-AGG GGT CTT G-3′) and OPA-18 (5′-AGG TGA CCG T-3′) were also used in several studies [44].
4 Classification of Serotypes
The genes involved in the biosynthesis of serotype-specific polysaccharides are estimated to be located in four different regions and those in strain UA159 (serotype c) are illustrated in Fig. 9.3. There are multiple enzymes required for the biosynthesis of the polysaccharides and the biochemical steps and their relevant genes have been identified [1]. The enzyme RgpG encoded by rgpG is proposed to be involved in the first step in the formation of the polysaccharides catalyzing the transfer of N-acetylglucosamine-1-phosphate to a lipid carrier [45]. On the other hand, the basic units of the rhamnose polymers and glucose side chains are considered to be dTDP-l-rhamnose and UDP-d-glucose, respectively. These units are synthesized from the UDP-d-glucose-1-phosphate by the actions of multiple enzymes encoded by their respective genes [46, 47]. The rmlA, rmlB, rmlC, and rmlD genes are known to encode the enzymes that catalyze the pathway from UDP-d-glucose-1-phosphate to the rhamnose units [46, 48], and the gluA gene encoding GluA is known to be involved in the biosynthesis of the units of the glucose side chain [49].
Genes involved in the biosynthesis of the serotype-specific polysaccharide of S. mutans. The genes were located four different regions (a–d). rgpG (a) and gluA (b) genes are completely conserved between two sequenced serotype c strains (UA159 and NN2025). However, genes located downstream of rgpF are quite diverse (c). rmlA-C genes are also conserved between the two serotype c strains, but several differences are found in the intergenetic regions (d)
The rgpA, rgpB, and rgpF genes encoding RgpA, RgpB, and RgpF, respectively, are reported to function in the polymerization of the rhamnose units [47]. Specifically, RgpA is proposed to be associated with the first rhamnose unit, whereas RgpB and RgpF are presumed to be correlated with the polymerization of the even and odd numbers of rhamnose units from the second unit, respectively [50]. In addition, the rgpE gene encoding RgpE is considered to be involved in the side-chain formation by glucose units [47]. Furthermore, RgpH was shown to encode a glucosyltransferase, while RgpI is thought to control the frequency of branching [51]. As for polysaccharide export, the rgpC and rgpD genes encoding RgpC and RgpD were demonstrated to regulate this function [47].
As compared to serotype c/e/f strains, the region from downstream of rgpF to the upstream of ORF12 was demonstrated to be highly variable among each serotype [38]. Using the differences in the nucleotide alignments in these regions, primer sets for identification of each c/e/f serotype were constructed. On the other hand, no drastic differences in the nucleotide alignments between serotype c and k strains were identified in that region [52]. However, a serotype k-specific alignment is present in the 5′ one-third end of the rgpF gene, upon which the serotype-k specific set of primers was constructed. It should be noted that these alterations in the nucleotide alignments in this region for the serotype k strains are considered to be inconsequential for the observed variety of glucose side chains in the serotype-specific polysaccharides [53]. Table 9.4 lists the primers for determination of each serotype c/e/f/k strain of S. mutans [38, 52]. PCR detection system using these primer sets were demonstrated to be very sensitive, with the minimum number of cells detected being 5–50 per reaction.
5 Identification of Virulent Strains
Considering the prevention of dental caries, the approaches used should be ideally based on the common risk factors for dental caries [54]. Thus, the identification of subjects with highly virulent strains could be beneficial for the prevention of dental caries. This should be true also when considering the pathogenesis of S. mutans in blood. It is generally accepted that considerable phenotypic variations exist within S. mutans species, which is derived from a consequence of a variety of genetic events, such as point mutations, translocations and inversions [35]. Therefore, some of the strains show strong virulence and others are regarded as weak virulent strains. In order to develop molecular methods to identify subjects who harbor the highly virulent S. mutans strains, several PCR approaches using the extracted bacterial DNA from the specimens have been evaluated.
One of the methods for possible clinical use in estimating the risk of dental caries in subjects is the identification of the multiple serotypes of S. mutans in specimens from the oral cavity by PCR with serotype-specific sets of primers [38]. The use of such methods is supported by the evidence that dental caries scores for preschool children with multiple serotypes of S. mutans were shown to be significantly higher than those with a single serotype or with no detectable S. mutans. It should be noted that the risk for subjects should be estimated based upon their clinical conditions, such as the number of lesions or fillings, in addition to the results of the molecular analyses. The other study showed the unique distribution of the serotypes in the oral and cardiovascular specimens in subjects with cardiovascular diseases (Fig. 9.4) [55]. The serotype distribution patterns in the subjects with cardiovascular diseases were demonstrated to be totally different compared with healthy subjects. Thus, it might be possible to also use serotype determinations as a means of identifying subjects at risk for developing cardiovascular diseases although additional confirmation of such a relationship is still required.
Although there are a large number of studies attempting to identify the association of the cell surface protein antigens and the pathogenesis of dental caries, the development of molecular methods to analyze for virulence genes is relatively uncommon. In this regard, one of the approaches is RFLP analysis of the gtf genes, which is based upon the high diversity of the gtf genes [56]. In that method, the 5.2-kb gtfB and 4.3-kb gtfC genes amplified by PCR are digested with BsrI and SspI, respectively. Ten and five genotypes were designated based on the digestion patterns for gtfB and gtfC, respectively. However, there were no correlations found between specific genotypes and the GTF enzymatic activities.
Recently, the cnm gene, encoding a 120-kDa cell-surface collagen-binding adhesin of S. mutans, was cloned and sequenced [57], which has received attention due to the possible association of dental caries with infective endocarditis [58]. S. mutans strains with the cnm gene are estimated to be present in approximately 10–20% of individuals. The cnm-positive strains possess significantly higher activities for binding type I collagen than the cnm-negative strains (Fig. 9.5a). Thus, S. mutans strains with cnm were predicted to show high virulence for dental caries since type I collagen is also a major organic component of dentin. It was also proposed that the cnm-positive strains could bind with higher affinity than the cnm-negative strains once the dentin is exposed as caries progresses. In fact, clinical parameters indicate that dental caries in children with cnm-positive S. mutans in saliva was significantly higher than those with cnm-negative S. mutans strains as well as S. mutans-negative children (Fig. 9.5b).
The cnm gene consists of the conserved collagen-binding domain in the 5′-region, followed by a region containing multiple B-repeats, whose length varied among the different strains. The primer set specific for the S. mutans cnm gene was constructed based on the nucleotide alignment of the cnm gene (cnm-1F 5′-GAC AAA GAA ATG AAA GAT GT-3′ and cnm-1R 5′-GCA AAG ACT CTT GTC CCT GC-3′). The size of the amplified fragments varied from approximately 1,650–1,750 bp due to the number of repeats within the amplicon. The distribution frequency of the cnm gene in S. mutans strains in the oral cavity is estimated to be approximately 10–20%, with the cnm-positive strains showing a predominant distribution among strains with the minor serotypes f and k.
It was reported that a S. mutans strain with defects in the expression of all three GTFs has been isolated [58]. This defect of caused a drastic reduction in its virulence potential for inducing dental caries [40], however, the concomitant decrease in GTF antigenicity was speculated to result in lower susceptibility to phagocytosis by polymorphonuclear leukocytes. This could result in the enhanced survival of such strains in blood compared to GTF-expressing strains. Using the specific nucleotide alignment in gtf regions, primer sets specific for detection of similar non-GTF expressing strains were constructed [59]. The detection rate for such strains was shown to be quite low. It has been hypothesized that alterations of cell surface structures of S. mutans are considered to be related to the survival in blood as well as the pathogenicity for infective endocarditis [60]. Therefore, further studies focused on the relationship between the cell surface antigens and pathogenesis as well as the development of PCR methods to identify subjects with these highly virulent strains should be considered.
6 Transmission Studies
Acquisition of S. mutans is considered to be initiated after the first primary tooth erupts, which provides a location for the bacterium to be colonized [61]. The first tooth which erupts is the generally mandibular primary central incisor, which emerges into the oral cavity at an age between 6 and 12 months. The number of erupted teeth increases as children grow, and they face a critical period for colonization at the age between 19 and 31 months called the “window of infectivity” [62], although the speed of colonization could depend on the caries activity of the bacterial population. More recent studies indicate that S. mutans can colonize the mouths of predentate infants [63], which indicates the possibility that the predentate children receive S. mutans frequently in their mouth and some are transient and some can colonize on the tongue. They are then able to attach to the surface of the first erupting tooth when circumstances are favorable such as with sugar ingestion. The original sources of S. mutans have been demonstrated to be mainly their mothers from a large number of studies conducted worldwide. Longitudinal studies regarding the genotypes of S. mutans in children demonstrate that most of the initially acquired genotypes generally transmitted from the mothers persist and some are lost and new strains are also acquired. In addition, the sharing of S. mutans genotypes between siblings has also been reported, which suggests the possibility of horizontal transfer of strains acquired from mothers between siblings [64]. Furthermore, the transmission of S. mutans strains from other family members or other care givers has also been considered.
Saliva is considered to be the major vehicle for oral bacterial transmission, and a high level of salivary S. mutans in mothers results in the earlier colonization of the bacterium in their children [65]. In addition, saliva specimens are thought to reflect the composition of the whole oral cavity, whereas dental plaque specimens primarily indicate localized colonization [66]. Studies regarding the transmission of S. mutans have been performed by comparison of the isolated strains using various subtyping strategies, such as serotyping, bacteriocin activity profiles, and molecular typings [4]. As for molecular biological methods, CDF techniques commonly employed with HaeIII digestion, ribotyping, AP-PCR assays, or random-amplified polymorphic DNA (RAPD) analyses are generally used [43, 67]. In addition, the diversity of S. mutans strains from children and their mothers was investigated by RFLP of the gtfB gene digested by HaeIII [68].
It should be advantageous to analyze as many strains as possible in a single subject when performing transmission studies. Since there exist time and financial limitations, the number of the estimated genotypes in the populations should be carefully considered when constructing the study design. As for Japanese, approximately 90% of the subjects are estimated to harbor one or two genotypes (average; 1.9 genotypes) [69]. Another study conducted in China demonstrated that 95% of the subjects aged 9–14 years possess one or two genotypes (average; 1.5 genotypes) [70]. Thus, 3–5 randomly selected representative strains should be sufficient for S. mutans to be analysis when performing transmission studies.
On the other hand, a study carried out in Sweden showed that only 60% of the subjects between the ages of 20–40 possessed fewer than two genotypes (average; 2.6 genotypes), and the maximum number of the genotypes was shown to be seven [67]. In addition, analysis of the subjects aged 18–29 years held in Brazil showed that a caries-free group possessed one to four genotypes (average; 3.0 genotypes) and that the caries-active group possessed two to eight genotypes (average; 5.5 genotypes). When analyzing these populations, the numbers of the strains are recommended for the study should be as large as possible [71]. In addition, it should be noted that the number of genotypes of S. mutans could be influenced by the dental caries status of each individual. AP-PCR analyses revealed that the children with severe dental caries based on the inappropriate usage of nursing bottles showed higher numbers of genotypes as compared with that of caries-free children [72]. This finding suggests that during favorable circumstances, such as sugar exposure, it is easier for new genotypes to be colonized.
The intra-familial transmission rates have been reported by a large number of groups worldwide and could be influenced by many factors such as cultural background, even within similar populations in the same country. Although most of the studies focus on the mother–child transmission of S. mutans, there is one study considering father–child transmission [69]. In that study, analysis of 1908 isolates from 76 subjects with 20 Japanese families including children below the age of 12 demonstrated that the transmission ratio from mothers and fathers were demonstrated to be 51.4% and 31.4%, respectively.
7 Multilocus Sequencing Typing Approach
Multilocus sequencing typing (MLST) is a generic typing method, employed to date principally, but not solely with bacterial pathogens, which aims to be a robust and portable method for the characterization of bacterial isolates at the molecular level. This method differs from many other approaches for characterization in that it is based explicitly on population genetic concepts [73]. MLST usually employs allele fragments of housekeeping genes approximately 400–600 bp in length and 6–10 loci were selected for sequencing because MLST provides sufficient discrimination for bacterial typing without being subject to diversifying selections which could obscure relationships among isolates. This method was first developed in 1998 in a study of Neisseria meningitides [74] and it has been applied for examination of approximately 40 species of microorganisms.
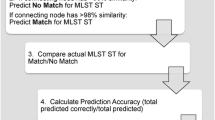
As mentioned above, several genotypic typing methodologies have been used to subtype S. mutans including multilocus enzyme electrophoresis (MLEE), ribotyping, and RAPD [64, 67, 71]. More discriminating methods for the subtyping of S. mutans include pulsed-field gel electrophoresis (PFGE) [75]. However, these methods differ in their discriminatory abilities power and reproducibility. Therefore, we have developed the MLST method for S. mutans typing (Fig. 9.6). Table 9.5 lists the eight housekeeping gene loci applied for a MLST scheme for S. mutans [6].
Initially, the internal fragments of the housekeeping genes were amplified and their nucleotide sequences determined (GenBank accession numbers AB281702-AB282509 and AB427220-AB428307). The sequences for each allele are compared with those in nonredundant databases (http://pubmlst.org/oralstrep/) and allele numbers were assigned for each strain, which defines the allelic profile. Finally, the sequence types (STs) for each strain are assigned. Numbers of the alleles for eight kinds of housekeeping genes are between 14 and 26 and STs 1–108 are assigned for 238 strains from 142 subjects at present [4, 6].
Figure 9.7 shows a phylogenetic tree based on hierarchical clustering of MLST results with 108 STs from 238 strains isolated in Japan and Finland. This result indicated that S. mutans contains a diverse population. This method was proven to theoretically distinguish more than 1.2 × 1010 sequence types. The serotype c strains are shown to be widely distributed in the tree, whereas the serotype e, f, and k strains were differentiated into clonal complexes, suggesting that the original ancestral strain of S. mutans was serotype c. Although no geographic specificity was identified, the distribution of the cnm gene was demonstrated to be clearly evident.
Cluster analysis of ST profiles and the relationship among serotypes, year of isolation, and areas of isolation. Phylogenetic tree constructed based on the 164 strains isolated from Japan and Finland using CLUSTER3 software (http://bonsai.ims.u-tokyo.ac.jp/%7Emdehoon/software/cluster/software.htm) and Java TreeView (http://jtreeview.sourceforge.net/). Area—U, USA; J, Japan; F, Finland
The superior discriminatory capacity of this MLST method for S. mutans may have important practical implications. Although various kinds of subtyping methods have been applied for transmission studies of S. mutans, the high discriminatory power gained by the MLST method is considered to result in greater sensitivity. Using MLST, 20 Japanese mother–child pairs whose children were between 2 and 10 years of ages showed that transmission could be observed with 70% of the pairs [4]. The MLST method could be applied for various epidemiological studies, which possibly could lead to grouping of the virulent strains of S. mutans into specific clusters to aid clinical assessment in the near future.
8 Summary
In summary, recent development of molecular biological techniques enables the detection of target bacterial species and their virulence genes without direct isolation of the strains. As for S. mutans, there have been a large number of such approaches developed. Identification of S. mutans followed by speciation of the highly virulent strains for dental caries as well as other diseases could be one of the powerful tools for clinical interventions in the future. Accumulation of data from clinical studies using molecular biological techniques might lead to the development of novel relevant systems for clinical use.
References
Nakano K, Ooshima T (2009) Serotype classification of Streptococcus mutans and its detection outside the oral cavity. Future Microbiol 4:891–902
Hamada S, Slade HD (1980) Biology, immunology, and cariogenicity of Streptococcus mutans. Microbiol Rev 44:331–384
Nakano K, Nomura R, Nakagawa I, Hamada S, Ooshima T (2004) Demonstration of Streptococcus mutans with a cell wall polysaccharide specific to a new serotype, k, in the human oral cavity. J Clin Microbiol 42:198–202
Lapirattanakul J, Nakano K, Nomura R, Hamada S, Nakagawa I, Ooshima T (2008) Demonstration of mother-to-child transmission of Streptococcus mutans using multilocus sequence typing. Caries Res 42:466–474
Waterhouse JC, Russell RR (2006) Dispensable genes and foreign DNA in Streptococcus mutans. Microbiology 152:1777–1788
Nakano K, Lapirattanakul J, Nomura R et al (2007) Streptococcus mutans clonal variation revealed by multilocus sequence typing. J Clin Microbiol 45:2616–2625
Linossier AG, Valenzuela CY, Toledo H (2008) Differences of the oral colonization by Streptococcus of the mutans group in children and adolescents with Down syndrome, mental retardation and normal controls. Med Oral Patol Oral Cir Bucal 13:E536–E539
Koga T, Okahashi N, Takahashi I, Kanamoto T, Asakawa H, Iwaki M (1990) Surface hydrophobicity, adherence, and aggregation of cell surface protein antigen mutants of Streptococcus mutans serotype c. Infect Immun 58:289–296
Kuramitsu HK (1993) Virulence factors of mutans streptococci: role of molecular genetics. Crit Rev Oral Biol Med 4:159–176
Banas JA, Vickerman MM (2003) Glucan-binding proteins of the oral streptococci. Crit Rev Oral Biol Med 14:89–99
Ajdić D, McShan WM, McLaughlin RE et al (2002) Genome sequence of Streptococcus mutans UA159, a cariogenic dental pathogen. Proc Natl Acad Sci USA 99:14434–14439
Maruyama F, Kobata M, Kurokawa K et al (2009) Comparative genomic analyses of Streptococcus mutans provide insights into chromosomal shuffling and species-specific content. BMC Genomics 10:358
Smorawinska M, Kuramitsu HK (1992) DNA probes for detection of cariogenic Streptococcus mutans. Oral Microbiol Immunol 7:177–181
Ono T, Hirota K, Nemoto K, Fernandez EJ, Ota F, Fukui K (1994) Detection of Streptococcus mutans by PCR amplification of spaP gene. J Med Microbiol 41:231–235
Igarashi T, Yamamoto A, Goto N (1996) Direct detection of Streptococcus mutans in human dental plaque by polymerase chain reaction. Oral Microbiol Immunol 11:294–298
Nakano K, Inaba H, Nomura R et al (2006) Detection of cariogenic Streptococcus mutans in extirpated heart valve and atheromatous plaque specimens. J Clin Microbiol 44:3313–3317
Nakano K, Nemoto H, Nomura R et al (2009) Detection of oral bacteria in cardiovascular specimens. Oral Microbiol Immunol 24:64–68
Nomura R, Nakano K, Nemoto H et al (2009) Molecular analyses of bacterial DNA in extirpated heart valves from patients with infective endocarditis. Oral Microbiol Immunol 24:43–49
Hokamura K, Inaba H, Nakano K et al (2010) Molecular analysis of aortic intimal hyperplasia caused by Porphyromonas gingivalis infection in mice with endothelial damage. J Periodontal Res 45:337–344
Oho T, Yamashita Y, Shimazaki Y, Kushiyama M, Koga T (2000) Simple and rapid detection of Streptococcus mutans and Streptococcus sobrinus in human saliva by polymerase chain reaction. Oral Microbiol Immunol 15:258–262
Hoshino T, Kawaguchi M, Shimizu N, Hoshino N, Ooshima T, Fujiwara T (2004) PCR detection and identification of oral streptococci in saliva samples using gtf genes. Diagn Microbiol Infect Dis 48:195–199
Yano A, Kaneko N, Ida H, Yamaguchi T, Hanada N (2002) Real-time PCR for quantification of Streptococcus mutans. FEMS Microbiol Lett 217:23–30
Hameş-Kocabaş EE, Uçar F, Kocataş Ersin N, Uzel A, Alpöz AR (2008) Colonization and vertical transmission of Streptococcus mutans in Turkish children. Microbiol Res 163:168–172
Sato T, Matsuyama J, Kumagai T et al (2003) Nested PCR for detection of mutans streptococci in dental plaque. Lett Appl Microbiol 37:66–69
Rupf S, Merte K, Eschrich K, Stösser L, Kneist S (2001) Peroxidase reaction as a parameter for discrimination of Streptococcus mutans and Streptococcus sobrinus. Caries Res 35:258–264
Al-Ahmad A, Auschill TM, Braun G, Hellwig E, Arweiler NB (2006) Overestimation of Streptococcus mutans prevalence by nested PCR detection of the 16S rRNA gene. J Med Microbiol 55:109–113
Fujiwara T, Nakano K, Kawaguchi M et al (2001) Biochemical and genetic characterization of serologically untypable Streptococcus mutans strains isolated from patients with bacteremia. Eur J Oral Sci 109:330–334
Teng LJ, Hsueh PR, Tsai JC et al (2002) groESL sequence determination, phylogenetic analysis, and species differentiation for viridans group streptococci. J Clin Microbiol 40:3172–3178
Hung WC, Tsai JC, Hsueh PR, Chia JS, Teng LJ (2005) Species identification of mutans streptococci by groESL gene sequence. J Med Microbiol 54:857–862
Garnier F, Gerbaud G, Courvalin P, Galimand M (1997) Identification of clinically relevant viridans group streptococci to the species level by PCR. J Clin Microbiol 35:2337–2341
Poyart C, Quesne G, Coulon S, Berche P, Trieu-Cuot P (1998) Identification of streptococci to species level by sequencing the gene encoding the manganese-dependent superoxide dismutase. J Clin Microbiol 36:41–47
Chen CC, Teng LJ, Chang TC (2004) Identification of clinically relevant viridans group streptococci by sequence analysis of the 16S-23S ribosomal DNA spacer region. J Clin Microbiol 42:2651–2657
Rovery C, Greub G, Lepidi H et al (2005) PCR detection of bacteria on cardiac valves of patients with treated bacterial endocarditis. J Clin Microbiol 43:163–167
Marques da Silva R, Caugant DA, Eribe ER et al (2006) Bacterial diversity in aortic aneurysms determined by 16S ribosomal RNA gene analysis. J Vasc Surg 44:1055–1060
Russell RR (2008) How has genomics altered our view of caries microbiology? Caries Res 42:319–327
Loesche WJ (1986) Role of Streptococcus mutans in human dental decay. Microbiol Rev 50:353–380
Ooshima T, Sobue S, Hamada S, Kotani S (1981) Susceptibility of rats, hamsters, and mice to carious infection by Streptococcus mutans serotype c and d organisms. J Dent Res 60:855–859
Shibata Y, Ozaki K, Seki M et al (2003) Analysis of loci required for determination of serotype antigenicity in Streptococcus mutans and its clinical utilization. J Clin Microbiol 41:4107–4112
Seki M, Yamashita Y, Shibata Y, Torigoe H, Tsuda H, Maeno M (2006) Effect of mixed mutans streptococci colonization on caries development. Oral Microbiol Immunol 21:47–52
Ooshima T, Matsumura M, Hoshino T, Kawabata S, Sobue S, Fujiwara T (2001) Contributions of three glucosyltransferases to sucrose-dependent adherence of Streptococcus mutans. J Dent Res 80:1672–1677
Nomura R, Nakano K, Taniguchi N et al (2009) Molecular and clinical analyses of the gene encoding the collagen-binding adhesin of Streptococcus mutans. J Med Microbiol 58:469–475
Shiroza T, Shinozaki N, Watanabe T, Ikemi T, Fukushima K, Abiko Y (1998) Rapid isolation of chromosomal DNA from oral streptococci and polymerase chain reaction-oriented restriction fragment-length polymorphism analysis for genetic heterogeneity. Oral Microbiol Immunol 13:11–16
Li Y, Caufield PW (1998) Arbitrarily primed polymerase chain reaction fingerprinting for the genotypic identification of mutans streptococci from humans. Oral Microbiol Immunol 13:17–22
Saarela M, Hannula J, Mättö J, Asikainen S, Alaluusua S (1996) Typing of mutans streptococci by arbitrarily primed polymerase chain reaction. Arch Oral Biol 41:821–826
Yamashita Y, Shibata Y, Nakano Y et al (1999) A novel gene required for rhamnose-glucose polysaccharide synthesis in Streptococcus mutans. J Bacteriol 181:6556–6559
Tsukioka Y, Yamashita Y, Nakano Y, Oho T, Koga T (1997) Identification of a fourth gene involved in dTDP-rhamnose synthesis in Streptococcus mutans. J Bacteriol 179:4411–4414
Yamashita Y, Tsukioka Y, Tomihisa K, Nakano Y, Koga T (1998) Genes involved in cell wall localization and side chain formation of rhamnose-glucose polysaccharide in Streptococcus mutans. J Bacteriol 180:5803–5807
Tsukioka Y, Yamashita Y, Oho T, Nakano Y, Koga T (1997) Biological function of the dTDP-rhamnose synthesis pathway in Streptococcus mutans. J Bacteriol 179:1126–1134
Yamashita Y, Tsukioka Y, Nakano Y, Tomihisa K, Oho T, Koga T (1998) Biological functions of UDP-glucose synthesis in Streptococcus mutans. Microbiology 144:1235–1245
Shibata Y, Yamashita Y, Ozaki K, Nakano Y, Koga T (2002) Expression and characterization of streptococcal rgp genes required for rhamnan synthesis in Escherichia coli. Infect Immun 70:2891–2898
Ozaki K, Shibata Y, Yamashita Y, Nakano Y, Tsuda H, Koga T (2002) A novel mechanism for glucose side-chain formation in rhamnose-glucose polysaccharide synthesis. FEBS Lett 532:159–163
Nakano K, Nomura R, Shimizu N, Nakagawa I, Hamada S, Ooshima T (2004) Development of a PCR method for rapid identification of new Streptococcus mutans serotype k strains. J Clin Microbiol 42:4925–4930
Nomura R, Nakano K, Ooshima T (2005) Molecular analysis of the genes involved in the biosynthesis of serotype specific polysaccharide in the novel serotype k strains of Streptococcus mutans. Oral Microbiol Immunol 20:303–309
Selwitz RH, Ismail AI, Pitts NB (2007) Dental caries. Lancet 369:51–59
Nakano K, Nemoto H, Nomura R et al (2007) Serotype distribution of Streptococcus mutans, a pathogen of dental caries in cardiovascular specimens from Japanese patients. J Med Microbiol 56:551–556
Mattos-Graner RO, Napimoga MH, Fukushima K, Duncan MJ, Smith DJ (2004) Comparative analysis of Gtf isozyme production and diversity in isolates of Streptococcus mutans with different biofilm growth phenotypes. J Clin Microbiol 42:4586–4592
Sato Y, Okamoto K, Kagami A, Yamamoto Y, Igarashi T, Kizaki H (2004) Streptococcus mutans strains harboring collagen-binding adhesin. J Dent Res 83:534–539
Nomura R, Nakano K, Nemoto H et al (2006) Isolation and characterization of Streptococcus mutans in heart valve and dental plaque specimens from a patient with infective endocarditis. J Med Microbiol 55:1135–1140
Nemoto H, Nakano K, Nomura R, Ooshima T (2008) Molecular characterization of Streptococcus mutans strains isolated from the heart valve of an infective endocarditis patient. J Med Microbiol 57:891–895
Lu J, Zhang W, Hao Y, Zhu Y (2009) Defect of cell wall construction may shield oral bacteria’s survival in bloodstream and cause infective endocarditis. Med Hypotheses 73:1055–1057
Berkowitz RJ (2003) Acquisition and transmission of mutans streptococci. J Calif Dent Assoc 31:135–138
Caufield PW, Cutter GR, Dasanayake AP (1993) Initial acquisition of mutans streptococci by infants: evidence for a discrete window of infectivity. J Dent Res 72:37–45
Berkowitz RJ (2006) Mutans streptococci: acquisition and transmission. Pediatr Dent 28:106–109
Köhler B, Lundberg AB, Birkhed D, Papapanou PN (2003) Longitudinal study of intrafamilial mutans streptococci ribotypes. Eur J Oral Sci 111:383–389
Thorild I, Lindau-Jonson B, Twetman S (2002) Prevalence of salivary Streptococcus mutans in mothers and in their preschool children. Int J Paediatr Dent 12:2–7
Miyamoto E, Nakano K, Fujita K et al (2009) Bacterial profiles of oral streptococcal and periodontal bacterial species in saliva specimens from Japanese subjects. Arch Oral Biol 54:374–379
Redmo Emanuelsson IM, Carlsson P, Hamberg K, Bratthall D (2003) Tracing genotypes of mutans streptococci on tooth sites by random amplified polymorphic DNA (RAPD) analysis. Oral Microbiol Immunol 18:24–29
Toi CS, Cleaton-Jones P, Fatti P (2005) Characterization of Streptococcus mutans diversity by determining restriction fragment-length polymorphisms of the gtfB gene of isolates from 5-year-old children and their mothers. Antonie Van Leeuwenhoek 88:75–85
Kozai K, Nakayama R, Tedjosasongko U et al (1999) Intrafamilial distribution of mutans streptococci in Japanese families and possibility of father-to-child transmission. Microbiol Immunol 43:99–106
Liu J, Bian Z, Fan M et al (2004) Typing of mutans streptococci by arbitrarily primed PCR in patients undergoing orthodontic treatment. Caries Res 38:523–529
Napimoga MH, Kamiya RU, Rosa RT et al (2004) Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol 53:697–703
Alaluusua S, Mättö J, Grönroos L et al (1996) Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol 41:167–173
Maiden MC (2006) Multilocus sequence typing of bacteria. Annu Rev Microbiol 60:561–588
Maiden MC, Bygraves JA, Feil E et al (1998) Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc Natl Acad Sci USA 95:3140–3145
Mineyama R, Yoshino S, Maeda N (2007) DNA fingerprinting of isolates of Streptococcus mutans by pulsed-field gel electrophoresis. Microbiol Res 162:244–249
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Nakano, K., Nakagawa, I., Alaluusua, S., Ooshima, T. (2013). Molecular Typing of Streptococcus mutans . In: de Filippis, I., McKee, M. (eds) Molecular Typing in Bacterial Infections. Infectious Disease. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-185-1_9
Download citation
DOI: https://doi.org/10.1007/978-1-62703-185-1_9
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-184-4
Online ISBN: 978-1-62703-185-1
eBook Packages: MedicineMedicine (R0)