Abstract
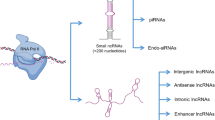
Emerging evidence from these studies suggested that the male germ cell transcriptome is more complex than previously envisioned. In addition to protein-coding genes, the transcriptome also encodes a significant number of nonprotein-coding transcripts. These noncoding (nc) RNAs appear to be involved in a variety of cellular activities, ranging from simple housekeeping to complex regulatory functions. A class of ncRNAs known as long ncRNAs (lncRNAs) were recently shown to be expressed in a developmentally regulated manner during brain and embryonic stem cell development. This protocol aims to predict and identify potential lncRNA candidates using Serial Analysis of Gene Expression (SAGE) data. We also illustrate how to validate the potential lncRNAs by expression analyses using real-time PCR and Northern Blot. Potential lncRNA candidates in male germ cells are identified using our previously established male germ cell SAGE database (GermSAGE).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Mattick JS (2001) Non-coding RNAs: the architects of eukaryotic complexity. EMBO Rep 2:986–991.
Mattick JS, Makunin IV (2005) Small regulatory RNAs in mammals. Hum Mol Genet 14 Spec No 1:R121–132.
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10:155–159.
Taft RJ, Pang KC, Mercer TR, Dinger M, Mattick JS (2010) Non-coding RNAs: regulators of disease. J Pathol 220:126–139.
Wu SM, Baxendale V, Chen Y, Pang AL, Stitely T, Munson PJ, Leung MY, Ravindranath N, Dym M, Rennert OM et al (2004) Analysis of mouse germ-cell transcriptome at different stages of spermatogenesis by SAGE: biological significance. Genomics 84:971–981.
Chan WY, Lee TL, Wu SM, Ruszczyk L, Alba D, Baxendale V, Rennert OM (2006) Transcriptome analyses of male germ cells with serial analysis of gene expression (SAGE). Mol Cell Endocrinol 250:8–19.
Chan WY, Wu SM, Ruszczyk L, Law E, Lee TL, Baxendale V, Lap-Yin Pang A, Rennert OM (2006) The complexity of antisense transcription revealed by the study of developing male germ cells. Genomics 87:681–692.
Lee TL, Li Y, Alba D, Vong QP, Wu SM, Baxendale V, Rennert OM, Lau YF, Chan WY (2009) Developmental staging of male murine embryonic gonad by SAGE analysis. J Genet Genomics 36:215–227.
Lee TL, Li Y, Cheung HH, Claus J, Singh S, Sastry C, Rennert OM, Lau YF, Chan WY (2010) GonadSAGE: a comprehensive SAGE database for transcript discovery on male embryonic gonad development. Bioinformatics 26:585–586.
Lee TL, Pang AL, Rennert OM, Chan WY (2009) Genomic landscape of developing male germ cells. Birth Defects Res C Embryo Today 87:43–63.
Lee TL, Cheung HH, Claus J, Sastry C, Singh S, Vu L, Rennert O, Chan WY (2009) GermSAGE: a comprehensive SAGE database for transcript discovery on male germ cell development. Nucleic Acids Res 37:D891–897.
Acknowledgments
This work was supported by the Intramural Research Program of the National Institutes of Health (NIH), Eunice Kennedy Shriver National Institute of Child Health and Human Development.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Lee, TL., Xiao, A., Rennert, O.M. (2012). Identification of Novel Long Noncoding RNA Transcripts in Male Germ Cells. In: Chan, WY., Blomberg, L. (eds) Germline Development. Methods in Molecular Biology, vol 825. Springer, New York, NY. https://doi.org/10.1007/978-1-61779-436-0_9
Download citation
DOI: https://doi.org/10.1007/978-1-61779-436-0_9
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-61779-435-3
Online ISBN: 978-1-61779-436-0
eBook Packages: Springer Protocols