Abstract
In the past several years, a host of new technologies have made it possible to visualize single molecules within cells and organisms (Raj et al., Nat Methods 5:877–879, 2008; Paré et al., Curr Biol 19:2037–2042, 2009; Lu and Tsourkas, Nucleic Acids Res 37:e100, 2009; Femino et al., Science 280:585–590, 1998; Rodriguez et al., Semin Cell Dev Biol 18:202–208, 2007; Betzig et al., Science 313:1642–1645, 2006; Rust et al., Nat Methods 3:793–796, 2006; Fusco et al., Curr Biol 13:161–167, 2003). Many of these are based on fluorescence, either fluorescent proteins or fluorescent dyes coupled to a molecule of interest. In many applications, the fluorescent signal is limited to a few pixels, which poses a classic signal processing problem: how can actual signal be distinguished from background noise?
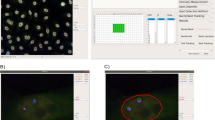
In this chapter, I present a MATLAB (MathWorks (2010) MATLAB. Retrieved from http://www.mathworks.com) software suite designed to work with these single-molecule visualization technologies (Rifkin (2010) spotFinding Suite. http://www.biology.ucsd.edu/labs/rifkin/software.html). It takes images or image stacks from a fluorescence microscope as input and outputs locations of the molecules. Although the software was developed for the specific application of identifying single mRNA transcripts in fixed specimens, it is more general than this and can be used and/or customized for other applications that produce localized signals embedded in a potentially noisy background. The analysis pipeline consists of the following steps: (a) create a gold-standard dataset, (b) train a machine-learning algorithm to classify image features as signal or noise depending upon user defined statistics, (c) run the machine-learning algorithm on a new dataset to identify mRNA locations, and (d) visually inspect and correct the results.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Raj A, van den Bogaard P, Rifkin SA et al (2008) Imaging individual mRNA molecules using multiple singly labeled probes. Nature Methods 5:877–879
Paré A, Lemons D, Kosman D et al (2009) Visualization of individual Scr mRNAs during drosophila embryogenesis yields evidence for transcriptional bursting. Curr Biol 19:2037–2042
Lu J, Tsourkas A (2009) Imaging individual microRNAs in single mammalian cells in situ. Nucleic Acids Res 37:e100
Femino AM, Fay FS, Fogarty K et al (1998) Visualization of single RNA transcripts in situ. Science 280:585–590
Rodriguez AJ, Condeelis J, Singer RH et al (2007) Imaging mRNA movement from transcription sites to translation sites. Seminars Cell Dev Biol 18:202–208
Betzig E, Patterson GH, Sougrat R et al (2006) Imaging intracellular fluorescent proteins at nanometer resolution. Science 313:1642–1645
Rust MJ, Bates M, Zhuang X (2006) Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nature Methods 3:793–796
Fusco D, Accornero N, Lavoie B et al (2003) Single mRNA molecules demonstrate probabilistic movement in living mammalian cells. Curr Biol 13:161–167
MathWorks (2010) MATLAB. Retrieved from http://www.mathworks.com
Rifkin SA (2010). spotFinding Suite. http://www.biology.ucsd.edu/labs/rifkin/software.html
Raj A, Tyagi S (2010) Detection of individual endogenous RNA transcripts in situ using multiple singly labeled probes. In: Walter NG (ed), Single Molecule Tools: Fluorescence Based Approaches, Part A. Academic Press
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2:18–22
Breiman L (2001) Random forests. Machine Learning 45:5–32
Breiman L, Cutler A (2001) Random Forests. http://www.stat.berkeley.edu/∼breiman/RandomForests/cc_home.htm
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Rifkin, S.A. (2012). Identifying Fluorescently Labeled Single Molecules in Image Stacks Using Machine Learning. In: Orgogozo, V., Rockman, M. (eds) Molecular Methods for Evolutionary Genetics. Methods in Molecular Biology, vol 772. Humana Press. https://doi.org/10.1007/978-1-61779-228-1_20
Download citation
DOI: https://doi.org/10.1007/978-1-61779-228-1_20
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-61779-227-4
Online ISBN: 978-1-61779-228-1
eBook Packages: Springer Protocols