Abstract
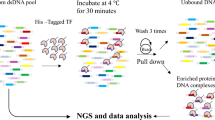
DNA-binding proteins, including transcription factors, play essential roles in many biological processes. The identification of the DNA sequences to which these proteins bind is a first, yet still challenging, step for determining their functions. SELEX provides an excellent tool for deciphering protein DNA-binding sequence specificity, and it has been widely adopted for addressing fundamental biological questions (1, 2). SELEX is an experimental procedure that involves the progressive selection, from a large combinatorial double-stranded oligonucleotide library, of DNA ligands with variable DNA-binding affinities and specificities by repeated rounds of partition and amplification. In this chapter, we describe a SELEX protocol that we have successfully applied to both plant and animal MYB transcription factors.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ng, E. W., Shima, D. T., Calias, P., Cunningham, E. T., Jr., Guyer, D. R., and Adamis, A. P. (2006) Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nat. Rev. Drug Discov. 5, 123–132.
Tuerk, C., and Gold, L. (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510.
Essers, J., Vermeulen, W., and Houtsmuller, A. B. (2006) DNA damage repair: anytime, anywhere? Curr. Opin. Cell Biol. 18, 240–246.
Sarai, A., and Kono, H. (2005) Protein-DNA recognition patterns and predictions. Annu. Rev. Biophys. Biomol. Struct. 34, 379–398.
Dervan, P. B. (1986) Design of sequence-specific DNA-binding molecules. Science 232, 464–471.
Matthews, B. W. (1988) Protein-DNA interaction. No code for recognition. Nature 335, 294–295.
Persikov, A. V., Osada, R., and Singh, M. (2009) Predicting DNA recognition by Cys2His2 zinc finger proteins. Bioinformatics 25, 22–29.
Collas, P., and Dahl, J. A. (2008) Chop it, ChIP it, check it: the current status of chromatin immunoprecipitation. Front. Biosci. 13, 929–943.
Grotewold, E., and Springer, N. (2009) Decoding the transcriptional hardwiring of the plant genome. In Annual Plant Reviews: Systems Biology. Coruzzi, G., and Guttierrez, R. (eds), Blackwell Publishing, Oxford, UK, vol. 35, pp. 196–227.
Haring, M., Offermann, S., Danker, T., Horst, I., Peterhansel, C., and Stam, M. (2007) Chromatin immunoprecipitation: optimization, quantitative analysis and data normalization. Plant Methods 3, 11.
Yang, Y., Yang, D., Schluesener, H. J., and Zhang, Z. (2007) Advances in SELEX and application of aptamers in the central nervous system. Biomol. Eng. 24, 583–592.
Ellington, A. D., and Szostak, J. W. (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822.
Stoltenburg, R., Reinemann, C., and Strehlitz, B. (2007) SELEX–a (r)evolut-ionary method to generate high-affinity nucleic acid ligands. Biomol. Eng. 24, 381–403.
Jayasena, S. D. (1999) Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 45, 1628–1650.
Djordjevic, M. (2007) SELEX experiments: new prospects, applications and data analysis in inferring regulatory pathways. Biomol. Eng. 24, 179–189.
Gopinath, S. C. (2007) Methods developed for SELEX. Anal. Bioanal. Chem. 387, 171–182.
Grotewold, E., Drummond, B. J., Bowen, B., and Peterson, T. (1994) The myb-homologous P gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset. Cell 76, 543–553.
Huang, H., Mizukami, Y., Hu, Y., and Ma, H. (1993) Isolation and characterization of the binding sequences for the product of the Arabidopsis floral homeotic gene AGAMOUS. Nucleic Acids Res. 21, 4769–4776.
Nole-Wilson, S., and Krizek, B. A. (2000) DNA binding properties of the Arabidopsis floral development protein AINTEGUMENTA. Nucleic Acids Res. 28, 4076–4082.
Acknowledgments
Support in the Grotewold lab for projects involving regulation of gene expression is provided by NRI Grant 2007-35318-17805 from the USDA CSREES, DOE Grant DE-FG02-07ER15881, and NSF grant DBI-0701405. ZX was supported by a 1-year predoctoral Excellence in Plant Molecular Biology & Biotechnology fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Chai, C., Xie, Z., Grotewold, E. (2011). SELEX (Systematic Evolution of Ligands by EXponential Enrichment), as a Powerful Tool for Deciphering the Protein–DNA Interaction Space. In: Yuan, L., Perry, S. (eds) Plant Transcription Factors. Methods in Molecular Biology, vol 754. Humana Press. https://doi.org/10.1007/978-1-61779-154-3_14
Download citation
DOI: https://doi.org/10.1007/978-1-61779-154-3_14
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-61779-153-6
Online ISBN: 978-1-61779-154-3
eBook Packages: Springer Protocols