Abstract
Pseudomonas aeruginosa is a Gram-negative bacterium which exacts a heavy burden on immunocompromised patients, but is non-pathogenic in a healthy host. Using small signaling molecules called acyl-homoserine lactones (AHLs), populations of P. aeruginosa can coordinate phenotypic changes, including biofilm formation and virulence factor secretion. This concentration-dependent process is called quorum sensing (QS). Interference with QS has been identified as a potential source of new treatments for P. aeruginosa infection. The human enzyme paraoxonase 1 (PON1) degrades AHL molecules, and is a promising candidate for QS interference therapy. Although paraoxonase orthologs exist in many species, genetic redundancy in humans and other mammals has made studying the specific effects of PON1 quite difficult. Arthropods, however, do not express any PON homologs. We generated a novel model to study the specific effects of PON1 by transgenically expressing human PON1 in Drosophila melanogaster. Using this model, we showed that P. aeruginosa infection lethality is QS-dependent, and that expression of PON1 has a protective effect. This work demonstrates the value of a D. melanogaster model for investigating the specific functions of members of the paraoxonase family in vivo, and suggests that PON1 plays a role in innateimmunity.
Time flies like an arrow; fruit flies like a banana
–Groucho Marx
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Paraoxonase
- Pseudomonas aeruginosa
- Drosophila melanogaster
- Quorum sensing
- 3OC12-HSL
- Biofilms
- Airway epithelium
- Host defense
- Evolution
1 Introduction
Pseudomonas aeruginosa is a Gram-negative bacterium which is ubiquitous in soil and water. Though generally non-pathogenic in a healthy host, P. aeruginosa represents a major cause of morbidity and mortality in immunocompromised patients. Burn victims, patients with diabetic foot ulcers, and those with chronic lung diseases such as cystic fibrosis (CF) are especially at risk of infection.
The ability of P. aeruginosa to form a biofilm contributes to antibiotic resistance and to poor outcomes in these populations. In planktonic cultures, P. aeruginosa is susceptible to a number of treatments. Within a biofilm, however, the bacteria are shielded from host immune responses and are much harder to treat; protected by a layer of secreted peptidogylcans, bacteria can more easily participate in horizontal gene transfer, and may evade drugs that target replication by entering a dormant phase (Fux et al., 2005). In P. aeruginosa, biofilm formation and virulence factor expression are regulated by quorum sensing.
Quorum sensing (QS) is a density-dependent process by which a population of bacteria can coordinate phenotypic changes by synthesizing, secreting, and responding to small signaling molecules known as autoinducers, or pheromones. QS was first identified in Vibrio fischeri, a marine bacterium which symbiotically colonizes the light organs of certain fish and squid (Waters and Bassler, 2005). Bacterial growth in this niche can become very dense, and when it reaches a threshold density, V. fischeri begins to transcribe luciferase and becomes biolumescent (Nealson, 1977). This behavior is accomplished via a positive feedback loop involving the production and detection of autoinducers, or quorum-sensing signals. Both V. fischeri and P. aeruginosa use acyl-homoserine lactones (AHLs) as their autoinducers.
Depending on the length of the carbon tail, AHLs can diffuse actively or passively across cell membranes (Pearson et al., 1999) in order to activate transcription factors in the LuxR family, which regulate certain target genes (Camilli and Bassler, 2006). AHL signaling is common in Gram-negative bacteria, but Gram-positive bacteria generally use short oligopeptide signals comprised of 5–17 amino acids. These signals do not diffuse freely through membranes, but may bind to cell-surface receptors. Finally, both Gram-negative and Gram-positive bacteria have been shown to produce and respond to a third category of quorum-sensing signal, known as autoinducer two (AI-2). As a result, AI-2 may facilitate “crosstalk” between different bacterial species (Lowery et al., 2008; Vlamakis and Kolter, 2005). Table 1 summarizes the three types of autoinducer molecules.
Quorum sensing signals/autoinducers
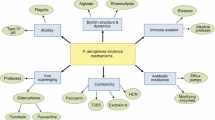
Pseudomonas aeruginosa has been shown to respond to both AHL and AI-2 signaling, but cannot synthesize AI-2 (Duan et al., 2003); we will therefore focus on the AHL pathway. AHL signaling in P. aeruginosa is controlled by a two-component regulatory system, which is depicted in Fig. 1. The AHL autoinducer N-3-oxodocecanoyl homoserine lactone (3OC12-HSL) diffuses across the cell membranes and activates the transcription factor LasR, which triggers a positive feedback loop by initiating the transcription of LasI, an enzyme which produces 3OC12-HSL. LasR also turns on the RhlR/RhlI regulatory circuit. The RhlI gene encodes an enzyme which produces another AHL, N-butanoyl homoserine lactone (C4-HSL); RhlR is a transcription factor which, when activated by C4-HSL, turns on transcription of RhlI. Numerous other genes, perhaps as much as 6% of the approximately 6,000 encoded by P. aeruginosa (Schuster et al., 2003; Wagner et al., 2003; Whiteley et al., 1999), are targeted by LasR and RhlR, either independently or together. Among these genes are those related to biofilm formation, and those that encode numerous virulence factors, including exotoxin A, alkaline protease, superoxide dismutase, elastase, and pyocyanin (Smith and Iglewski, 2003). In vivo experiments in animal models of wound and pulmonary infection have demonstrated attenuated virulence in QS-deficient strains of P. aeruginosa (Imamura et al., 2005; Pearson et al., 2000; Rumbaugh et al., 1999; Tang et al., 1996; Wu et al., 2001).
The las/rhl quorum-sensing system. P. aeruginosa uses a two-component QS system to regulate virulence factor expression and biofilm formation. The 3OC12-HSL autoinducer diffuses across cell membranes and activates the transcription factor LasR, which initiates the transcription of LasI, an enzyme which produces 3OC12-HSL. LasR also activates the rhlR and rhlI genes. RhlI is an enzyme that produces C4-HSL, the autoinducer associated with the rhlI/rhlR circuit. C4-HSL activates RhlR, a transcription factor encoded by the rhlR gene. Upon activation, RhlR initiates transcription of rhlI. Both LasR and RhlR also activate genes associated with virulence factors and biofilm formation
Interference with QS signaling, or “quorum quenching,” has been identified in nature and provides a promising avenue for new therapies against a range of bacterial pathogens. For example, soft-rot infection by the AHL quorum-sensing bacterium Erwinia carotovora was inhibited when AiiA, a lactonase naturally produced by Bacillus sp. bacteria, was transgenically expressed in tobacco and cabbage plants (Dong et al., 2001; Dong et al., 2000). Similarly, treatment with a synthetic analog of the furanones produced by the algae Delisea pulchra was shown to improve survival and clearance of pulmonary P. aeruginosa infection in a murine model (Hentzer et al., 2003; Wu et al., 2004).
The paraoxonase family of enzymes, including PON1, PON2, and PON3, has been associated with multiple enzymatic activities. In humans and other mammals, PON1 is secreted into the blood. It is capable of degrading paraoxon and other organophosphates, is active as a lactonase, and has been shown to protect against lipid oxidation (Draganov et al., 2005). Although the native substrate(s) of the PON family remain unknown, all three hydrolyze lactones. Recent work suggests that this may be PON’s endogenous enzymatic role (Khersonsky and Tawfik, 2005).
PON1 has been proposed as a candidate for P. aeruginosa QS signaling interference. Our laboratory has previously shown that human and murine airway epithelial cells, which endogenously express all three PONs, degrade 3OC12-HSL (Chun et al., 2004). In addition, mouse serum-PON1 degrades 3OC12-HSL in vitro (Ozer et al., 2005). Although P. aeruginosa QS signaling was not enhanced in mice with a PON1 deficiency, mRNA expression levels suggest that this may be due to compensation by other members of the PON family (Ozer et al., 2005). The redundancy of PON in humans and other mammals has made it difficult to tease apart the specific activities of individual members of the family in existing models. This chapter will describe a novel D. melanogaster model for PON1, and will summarize our recent data.
2 Paraoxonases are Widely Conserved, but Have Not Been Found in Arthropods
Previous work has shown that the paraoxonases are highly conserved among mammals, and indeed across many species (Draganov and La Du, 2004; Yang et al., 2005). These studies further suggest that redundancy in the paraoxonase family is the result of gene duplication, and that PON2 is the oldest member of the family while PON1 is the youngest. Interestingly, PON2 is primarily active as a lactonase, supporting the notion that lactonase activity is among the oldest endogenous functions of PON species (Draganov and La Du, 2004).
We generated a systematic phylogeny of PON orthologs based on identified and predicted amino acid reference sequences in the NCBI/BLASTp (Altschul et al., 1997) and Superfamily 1.73 (Gough et al., 2001) databases. Figure 2 shows the number of species per Domain, Phylum, Class, or Order which generated a match in these databases. PON-like domains are conserved across all three major Domains: Archaea, Bacteria, and Protozoa. We next generated an amino acid alignment using the T-Coffee program, and examined several important functional sites within the PON1 sequence (Labarga et al., 2007). We found that these sites are generally conserved as well. Figure 3 identifies key glycosylation sites and active site residues thought to be important for lactonase activity. The PON family is catalytically promiscuous, and has evolved to develop divergent enzymatic functions in many species (Khersonsky and Tawfik, 2006). These data suggest that PON possesses a long evolutionary history. Finally, although PON orthologs are present in most animals, it is not able that PON orthologs are absent from all arthropod sequences in both databases, including that of Drosophila melanogaster.
Conservation of PON1 functional sites in eukaryotes. Portion of PON1 amino acid sequence. Predicted functional sites are conserved across many species. Highlighted sites are conserved, according to the ClustalX algorithm. [*] indicates glycosylation site, and [#] indicates key active site residue; numbers are given according to the human PON1 amino acid sequence
3 Generating a PON1 Transgenic Drosophila melanogaster Infection Model
The finding that insects express no PON orthologs suggested that D. melanogaster might provide an ideal model system to study the distinct functions of individual members of the PON family. The D. melanogaster model has other experimental advantages as well, including a short generation time and well-characterized genetics. The D. melanogaster genome contains orthologs for as much as 80% of human disease genes (Gilbert, 2008), and it has been an important model for the discovery of key components of human innate immunity, including the Toll, Imd, and JAK/STAT pathways (Lemaitre and Hoffmann, 2007).
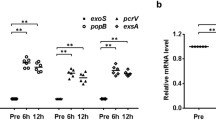
We assessed D. melanogaster as a model for P. aeruginosa infection. Using a previously-described abdomen wound infection model (Avet-Rochex et al., 2005; D‘Argenio et al., 2001; Lau et al., 2003), we first showed that P. aeruginosa infection lethality in wild-type D. melanogaster is QS-dependent. Briefly, we dipped a needle in a suspension of P. aeruginosa expressing green fluorescent protein (GFP) under the control of a QS-dependent promoter, and pricked the fly’s abdomen. Intense GFP expression 18 h post-inoculation indicated that P. aeruginosa was indeed expressing QS-dependent genes.
Next, we showed that QS increases P. aeruginosa lethality by infecting with mutant P. aeruginosa strains deficient in either AHL production (ΔlasI/rhlI) or detection (ΔlasR/rhlR). Compared to controls inoculated with wild-type P. aeruginosa, flies inoculated with either mutant strain had greater rates of survival. Interestingly, feeding 3OC12-HSL and C4-HSL to flies restored P. aeruginosa virulence in those infected with the AHL production mutant (ΔlasI/rhlI) but not the detection mutant (ΔlasR/rhlR).
Finally, we generated a novel PON whole-organism model in order to test whether the lactonase activity of PON1 would confer a protective effect against P. aeruginosa infection in vivo. We generated transgenic D. melanogaster ubiquitously expressing human PON1 (hPON1) using the GAL4-UAS system with a da promoter (Brand and Perrimon, 1993).
4 PON1 Transgenic Flies are Protected from Organophosphate Poisoning
After confirming PON1 expression and protein production by qPCR and Western blot, we set out to validate the in vivo function of this protein. The ability of PON1 to degrade organophosphates was among the first enzymatic activities identified for this family. We were therefore able to measure PON1 activity by exposing both transgenic and wild-type flies to the organophosphate chlorpyrifos, and comparing rates of survival between the two groups. Chlorpyrifos lethality was dose-dependent (Fig. 4a), and survival rates were significantly higher among PON1 transgenic (UAS-PON1/da-GAL4) flies than among (da-GAL4 +/+) controls (Fig. 4b). We also conducted control experiments with an organophosphate not degraded by PON1, demeton-S-methyl (DSM). As shown in Fig. 4c, DSM survival was similar in both groups.
Organophosphate lethality. Transgenic expression of human PON1 protects D. melanogaster from organophosphate toxicity. (a) Chlorpyrifos toxicity is dose-dependent in wild-type flies. (b) PON1 flies are protected from chlorpyrifos toxicity. (c) Similar DSM lethality is observed in PON1 and wild-type flies
5 PON1 Expression Protects Against Pseudomonas aeruginosa Lethality
Using the abdominal wound infection model we previously validated, we demonstrated that PON1 improved survival after infection with the PA01 strain of P. aeruginosa. We showed similar results with two clinical isolates of P. aeruginosa. Survival rates were similar to those of control flies infected with the AHL production-mutant ΔlasI/rhlI. Moreover, the PON1 transgenic flies were also protected against infection with Serratia marcescens, another AHL-sensing bacterium which is lethal to flies. However, when flies were infected with Staphylococcus aureus, which does not use AHL-mediated QS, the PON1 transgenic and control populations showed identical survival rates. Figure 5 summarizes this data (Harel et al., 2004; Tavori et al., 2008).
6 Concluding Remarks
The paraoxonases are a versatile family of enzymes, which have developed divergent catalytic specificities in many species over a long evolutionary history. Recent evidence suggests that the endogenous role of paraoxonase may be as a lactonase, and that this is the primary activity of PON2, the oldest member of the PON family. PON1 is the youngest member of the family. It is present in many tissues, and is secreted into the blood in humans and other mammals (Marsillach et al., 2008). PON1 is also active as a lactonase, but also degrades paraoxons. In vitro evidence has suggested that the former function may allow PON1 to play a role in host defense against infection by AHL-sensing bacteria. However, confirmation of these PON1-specific observations in vivo has been difficult due to genetic redundancy in humans and other mammals.
Taking advantage of the unique absence of paraoxonase orthologs in D. melanogaster and other arthropods, we developed a novel whole-organism model in which to investigate the specific effects of human PON1 on P. aeruginosa infection and QS. We expressed human PON1 in D. melanogaster and confirmed PON1 mRNA and protein expression in our transgenic model with biochemical assays, and demonstrated in vivo activity using chlorpyrifos susceptibility as a functional end point.
Using an abdominal wound infection model, we showed that P. aeruginosa lethality is dependent on QS, and that transgenic expression of hPON1 significantly increases survival after inoculation. PON1 was similarly protective after infection with another AHL-sensing bacterium, S. marcescens, but did not improve the survival rates of flies inoculated with S. aureus, a Gram-positive bacterium which does not use AHLs for QS. Taken together, our data suggest that PON1 may play an important role in innate immunity, particularly in preventing Gram-negative bacterial infection and biofilm formation. Moreover, we showed that D. melanogaster is a valuable model for investigating the specific functions of members of the paraoxonase family in vivo (see Fig. 6).
References
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., Miller, W., Lipman, D. J. (1997) Nucleic Acids Res 25, 3389–3402.
Avet-Rochex, A., Bergeret, E., Attree, I., Meister, M., Fauvarque, M. O. (2005) Cell Microbiol 7, 799–810.
Brand, A. H., Perrimon, N. (1993) Development 118, 401–415.
Camilli, A., Bassler, B. L. (2006) Science 311, 1113–1116.
Chun, C. K., Ozer, E. A., Welsh, M. J., Zabner, J., Greenberg, E. P. (2004) Proc Natl Acad Sci USA 101, 3587–3590.
Dong, Y. H., Wang, L. H., Xu, J. L., Zhang, H. B., Zhang, X. F., Zhang, L. H. (2001) Nature 411, 813–817.
Dong, Y. H., Xu, J. L., Li, X. Z., Zhang, L. H. (2000) Proc Natl Acad Sci USA 97, 3526–3531.
Draganov, D. I., La Du, B. N. (2004) Naunyn Schmiedebergs Arch Pharmacol 369, 78–88.
Draganov, D. I., Teiber, J. F., Speelman, A., Osawa, Y., Sunahara, R., La Du, B. N. (2005) J Lipid Res 46, 1239–1247.
Duan, K., Dammel, C., Stein, J., Rabin, H., Surette, M. G. (2003) Mol Microbiol 50, 1477–1491.
D‘Argenio, D. A., Gallagher, L. A. , Berg, C. A., Manoil, C. (2001) J Bacteriol 183, 1466–1471.
Fux, C. A., Costerton, J. W., Stewart, P. S., Stoodley, P. (2005) Trends Microbiol 13, 34–40.
Gilbert, L. I. (2008) Mol Cell Endocrinol 293, 25–31.
Gough, J., Karplus, K., Hughey, R., Chothia, C. (2001) J Mol Biol 313, 903–919.
Harel, M., Aharoni, A., Gaidukov, L., Brumshtein, B., Khersonsky, O., Meged, R., Dvir, H., Ravelli, R. B., McCarthy, A., Toker, L., et al. (2004) Nat Struct Mol Biol 11, 412–419.
Hentzer, M., Wu, H., Andersen, J. B., Riedel, K., Rasmussen, T. B., Bagge, N., Kumar, N., Schembri, M. A., Song, Z., Kristoffersen, P., et al. (2003) Embo J 22, 3803–3815.
Imamura, Y., Yanagihara, K., Tomono, K., Ohno, H., Higashiyama, Y., Miyazaki, Y., Hirakata, Y., Mizuta, Y., Kadota, J., Tsukamoto, K., et al. (2005) J. Med. Microbiol 54, 515–518.
Khersonsky, O., Tawfik, D. S. (2005) Biochemistry 44, 6371–6382.
Khersonsky, O., Tawfik, D. S. (2006) Chembiochem 7, 49–53.
Labarga, A., Valentin, F., Anderson, M., Lopez, R. (2007) Nucleic Acids Res 35, W6–11.
Lau, G. W., Goumnerov, B. C., Walendziewicz, C. L., Hewitson, J., Xiao, W., Mahajan-Miklos, S., Tompkins, R. G., Perkins, L. A., Rahme, L. G. (2003) Infect Immun 71, 4059–4066.
Lemaitre, B., Hoffmann, J. (2007) Annu Rev Immunol 25, 697–743.
Lowery, C. A., Dickerson, T. J., Janda, K. D. (2008) Chem Soc Rev 37, 1337–1346.
Marsillach, J., Mackness, B., Mackness, M., Riu, F., Beltran, R., Joven, J., Camps, J. (2008) Free Radic Biol Med 45, 146–157.
Nealson, K. H. (1977) Arch Microbiol 112, 73–79.
Ozer, E. A., Pezzulo, A., Shih, D. M., Chun, C., Furlong, C., Lusis, A. J., Greenberg, E. P., Zabner, J. (2005) FEMS Microbiol Lett 253, 29–37.
Pearson, J. P., Feldman, M., Iglewski, B. H., Prince, A. (2000) Infect Immun 68, 4331–4334.
Pearson, J. P., Van Delden, C., Iglewski, B. H. (1999) J Bacteriol 181, 1203–1210.
Rumbaugh, K. P., Griswold, J. A., Iglewski, B. H., Hamood, A. N. (1999) Infect. Immun 67, 5854–5862.
Schuster, M., Lostroh, C. P., Ogi, T., Greenberg, E. P. (2003) J. Bacteriol 185, 2066–2079.
Smith, R. S., Iglewski, B. H. (2003) Curr Opin Microbiol 6, 56–60.
Tang, H. B., DiMango, E., Bryan, R., Gambello, M., Iglewski, B. H., Goldberg, J. B., Prince, A. (1996) Infect Immun 64, 37–43.
Tavori, H., Khatib, S., Aviram, M., Vaya, J. (2008) Bioorg Med Chem 16, 7504–7509.
Vlamakis, H. C., Kolter, R. (2005) Nat Cell Biol 7, 933–934.
Wagner, V. E., Bushnell, D., Passador, L., Brooks, A. I., Iglewski, B. H. (2003) J Bacteriol 185, 2080–2095.
Waters, C. M., Bassler, B. L. (2005) Annu Rev Cell Dev Biol 21, 319–346.
Whiteley, M., Lee, K. M., Greenberg, E. P. (1999) Proc Natl Acad Sci USA 96, 13904–13909.
Wu, H., Song, Z., Givskov, M., Doring, G., Worlitzsch, D., Mathee, K., Rygaard, J., Hoiby, N. (2001) Microbiology 147, 1105–1113.
Wu, H., Song, Z., Hentzer, M., Andersen, J. B., Molin, S., Givskov, M., Hoiby, N. (2004) J Antimicrob Chemother 53, 1054–1061.
Yang, F., Wang, L. H., Wang, J., Dong, Y. H., Hu, J. Y., Zhang, L. H. (2005) FEBS Lett 579, 3713–3717.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Humana Press, a part of Springer Science+Business Media, LLC
About this paper
Cite this paper
Estin, M.L., Stoltz, D.A., Zabner, J. (2010). Paraoxonase 1, Quorum Sensing, and P. aeruginosa Infection: A Novel Model. In: Reddy, S. (eds) Paraoxonases in Inflammation, Infection, and Toxicology. Advances in Experimental Medicine and Biology, vol 660. Humana Press. https://doi.org/10.1007/978-1-60761-350-3_17
Download citation
DOI: https://doi.org/10.1007/978-1-60761-350-3_17
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-60761-349-7
Online ISBN: 978-1-60761-350-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)