Abstract
The mouse ooplasm is the ideal platform to study and compare induced and natural pluripotency because it can support both, after somatic cell nuclear transfer (cloning) and after fertilization, respectively. The amount of pluripotency induced after cloning is variable but always limited compared to fertilization. It can be visualized conveniently if the nucleus donor cells carry a green fluorescent protein (GFP) reporter under control of the pluripotency-associated gene Oct4 promoter. Thus we produced cloned and fertilized mouse embryos transgenic for Oct4-GFP (GOF18-∆PE-EGFP). We also developed and validated a live cell imaging method, whereby we resolve and selectively pick cloned embryos that hold distinct amounts of induced pluripotency as predicted by GFP intensity and measured by embryonic stem cell derivation. Currently we are developing a microinjection method to change the level of Oct4 without modifying the genome of the embryo. Here we discuss our findings in relation to the epigenetic reprogramming of the nucleus transplant and to cell fate decisions in the cloned or fertilized mouse embryo.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Natural Pluripotency
In mammalian development, pluripotency is the ability of a cell to give rise to a host of tissues belonging to the three primary germ layers: ectoderm, mesoderm, and endoderm, and the germ cells. Totipotency is the ability to give rise to all tissues including extraembryonic ones, e.g., the trophectoderm. It is believed that contribution of germ cells is sufficient to claim totipotency, since the gametes that form the totipotent zygote arise from germ cells. In mouse development, the zygote and the 2-cell stage blastomeres are totipotent. Isolated 4- and 8-cell stage blastomeres cannot form embryos that are viable in vivo, although they can contribute to all tissues of chimeric mouse embryos (1), form the whole mouse when supported with tetraploid blastomeres (2), and give rise to pluripotent cell lines (3). This indicates that cell number is important to implement pluripotency (4). At the blastocyst stage, the first cell lineage decision becomes apparent. The blastocyst is composed of an inner cell mass (ICM) and a trophectoderm (TE). The ICM but not the TE is pluripotent and can be derived in cell lines known as embryonic stem (ES) cells in mouse and human (5). Mouse ES cells cannot give rise to TE except after genetic manipulation (6) but can form germ cells both in vivo and in vitro (7), hence their potency is arguably almost full. Under current protocols, human ES cells can form TE in vitro (8) and probably give rise to germ cells, although the latter awaits formal proof.
In our current molecular understanding of pluripotency, which is mainly based on studies of mouse and human ES cells, the transcription factors Oct4, Sox2, and Nanog are crucial to keep cells self-renewing and preserve their pluripotency (9). These functions are implemented via a network of downstream transcriptional activators and repressors (5, 10–12). Studying how the network is regulated is complicated since responses are combinatorial (13) and depend on the protein level of more than a few factors. The case of Oct4 may be regarded paradigmatic for a dose-dependent effect on pluripotency, but it is not exclusive for Oct4. Mouse ES cells heterozygous for a mutation of Nanog are characterized by unstable pluripotency, as seen by in vitro cell differentiation in the absence of feeder cells (14). When Nanog is overexpressed, human ES cells acquire the ability to grow feeder-free, while features of the primitive ectoderm are induced (15), whereas hematopoietic stem cells suffer a disorder upon forced expression (16).
In this chapter we discuss the induction of pluripotency in somatic nuclei by means of the nuclear transfer technology that uses the ooplasm as a kind of bioreactor. Establishment of full pluripotency is the most defining aspect of the reprogramming process of somatic nuclei. Although complete reprogramming requires also the silencing of tissue-specific genes, we interchangeably use the terms “reprogramming” and “pluripotency induction.” Here we focus on Oct4 as the best (albeit not the only) known transcription factor associated with pluripotency in a dose-dependent manner. Because there can be no educated manipulation nor application without prior description of any biologic phenomena, first we summarize our observations from development of cloned and fertilized mouse embryos that hold different amounts of Oct4. In general, cloned embryos have lower levels of Oct4 than fertilized counterparts. Therefore, we hypothesize that the abnormal development of cloned mouse embryos is contributed, at least in part, by an insufficient level of Oct4. Based on these observations, it becomes a sensible deed to manipulate the level of Oct4 in cloned embryos without this necessarily making sense for fertilized embryos. In fact, the establishment of natural and induced pluripotency might follow different pathways, similar to the case of ES cells derived from blastocyst as opposed to induced pluripotent stem (iPS) cells formed directly from fibroblasts without an intermediate embryo (17). Finally, we anticipate an experimental approach to support nuclear reprogramming and pluripotency in mouse clones via manipulation of the cellular level of Oct4.
Oct4 as the Prototypic Determinant of Pluripotency
Oct4 is historically the pluripotency-associated factor (18–20). Somatic stem cells do not depend on Oct4 for self-renewal (20), and since they are also not pluripotent, this confirms the role of Oct4 in pluripotency rather than multi- or oligopotency. Yet we should resist the temptation to adopt an Oct4-centered view when dealing with pluripotency. Forced expression of Sox2 in the absence of Oct4 also keeps mouse ES cells pluripotent (21). In this chapter, we choose to confine our study to Oct4 in mouse development, being aware that there is a lot more to pluripotency than just Oct4.
Homozygous mouse embryos lacking the Oct4 locus form morphologically normal blastocysts but fail to form a pluripotent ICM (22). During embryogenesis, Oct4 knock-down in the ICM by RNA interference impairs cardiogenesis (23), whereas ubiquitous Oct4 expression driven by the chicken β-actin promoter and the cytomegalovirus enhancer affects midhindbrain patterning (24). In the adult mouse, epithelial cells respond to a bulk increase in Oct4 level and become neoplastic (25). In mouse ES cells, experimental manipulation of Oct4 protein levels below 50% or above 150% of the physiologic amount leads to differentiation along different lineages in vitro (6). Similarly, a bias in differentiation in vitro is observed in human ES cells upon manipulation of Oct4 level (26). In vivo, overproduction of Oct4 increases the malignant potential of mouse ES cell-derived tumors, while reduction of Oct4 induces tumor regression (27). In embryoid bodies, Oct4 overproduction inhibits hematopoietic differentiation in a dose-dependent manner (28). Maintenance of a certain protein level may be under post-translational control (29).
Induced Pluripotency
Pluripotency can be induced in somatic cells by nuclear transfer in the ooplasm, by cell–cell fusion where the partner cell is already pluripotent, by exposure to pluripotent cell extracts or specific molecules, and by retroviral delivery of genes encoding pluripotency factors. Induction of pluripotency becomes apparent from several landmarks, which include but are not limited to:
-
1.
Reactivation of somatic cell-encoded Oct4 and Nanog (14, 30) with demethylation of their relevant control genomic sequences
-
2.
Silencing of tissue-specific genes (31)
-
3.
Silencing of Xist and reactivation of the inactivated X-chromosome in female somatic cell (32, 33)
-
4.
Teratoma formation by injection of pluripotent cells into severe combined immunodeficiency (SCID) mice
-
5.
Formation of endodermal, mesodermal, and ectodermal tissues in embryoid bodies or contribution of such tissues in chimeric embryos
Historically, the chief method for pluripotency induction, known since 1928 from Hans Spemann and practiced since the 1950s by Briggs, King, and Gurdon, is to transplant the nucleus of a differentiated cell in an ooplasm. More than half a century later, transferring metaphase or interphase somatic genomes into metaphase zygotes or unfertilized oocytes, respectively, is still used to induce pluripotency (34). However, in humans this method is widely regarded as ethically objectionable. Cloned embryos are potentially implantable and may develop to term when transferred to a foster mother. In humans, in humans this potential of cloned embryos raises ethical concerns regarding the technique. Therefore, human reproductive cloning has not been permitted in any country to date. Another complication in oocyte-mediated nuclear reprogramming is the fact that special micromanipulation skills are needed, and freshly prepared unfertilized oocytes are required for making cloned blastocysts efficiently. In one day, only 100–200 nuclei may be successfully transplanted. The molecular and functional aspects of ooplasm-mediated nuclear reprogramming are dealt in the section “Ooplasm-Mediated Induction of Pluripotency.”
Alternatively, pluripotency can be induced in somatic cells by fusion with ES cells (30, 35) or by delivering via retroviruses specific genes encoding pluripotency-associated factors. Cell fusion and direct gene transfer give rise to iPS cells without generating an intermediate embryo. However, these methods entail modification of the donor genome, i.e., tetraploidy after fusion or insertional mutagenesis after retroviral infection. Strategies are being developed to eliminate the ES cells’ chromosomes after cell fusion (36) and to deliver the factors without insertional mutagenesis. So far, four factors have proven sufficient to induce pluripotency in mouse and human somatic cells after retrovirus-mediated delivery (OCT4, SOX2, C-MYC, KLF4 (17, 37); OCT4, SOX2, NANOG, LIN28 (38)). Most recently the number of factors has been reduced to three (OCT4, SOX2, KLF4 (39)).
Multi- and pluripotency can also be induced by exposing somatic cells to pluripotent cell extracts (40) or synthetic molecules (41) and by transduction of defined factors as proteins (42). When 293T cells are permeabilized and exposed to extracts of NCCIT human carcinoma cells, they form colonies that can be maintained for at least 23 passages (40). In these colonies, markers of differentiation such as Lamin A are down-regulated, pluripotency associated genes Oct4 and Sox2 are up-regulated, the Oct4 protein is detected in the cell nucleus, and the Oct4 promoter is demethylated (positions −1534 to −1773). Despite these remarkable observations, the nature of reprogramming induced via cell extracts is not yet firmly established. In particular, functional proof is lacking that cells reprogrammed by extracts can form unrelated tissues when placed in a proper proper developmental system, as is known for ES cells when injected into blastocysts. It is also unclear how the reprogramming factors reach their targets in the nucleus. This is an issue in protein transduction since a substantial proportion of the factor remains trapped inside cytoplasmic vesicles and thus is not (promptly) available (43).
Ooplasm-Mediated Induction of Pluripotency
Although nuclear transfer in an ooplasm is ethically objectionable, it remains the most efficient method to induce pluripotency, as it allows up to 60% blastocyst formation and from these about 20% ES cell line derivation (44). Full development of cloned embryos in the uterus provides the ultimate proof of effective reprogramming, as evidenced by the thousands of cloned animals produced worldwide (45).
A central question in ooplasm-mediated induction of pluripotency is why gene expression of resultant cloned embryos does not resemble that of fertilized embryos. The obvious answer is that a somatic genome does not have the epigenetic makeup of a gametic genome. The ooplasm machinery modifies the somatic chromatin in a way the chromatin was not prepared for (46); hence, gene expression tends to be abnormal. Another possibility is that nuclear reprogramming is random or stochastic and thereby generates gene expression patterns that are mostly abnormal, no matter which type of input. Bortvin et al. (47) analyzed the expression of Oct4 and 10 Oct4-related genes in individual cloned mouse blastocysts derived from cumulus cells. They found that 62% of these correctly expressed all tested genes. In contrast, ES cell-cloned embryos expressed these genes quite normally, although later studies exposed that levels of Oct4 mRNA in ES cell clones are actually lower than in somatic cell clones (48, 49). Additionally, ES cells derived from cumulus cell-cloned mouse embryos accumulate chromosomal aneuploidies (50). Although these views imply that nuclear reprogramming is bound to go wrong, they do not mean that it cannot be normalized, rescued, or prevented from getting worse. Recent observations suggest that cell fate in mouse embryos may be manipulated by changing the level of defined factors.
In a recent paper, Torres-Padilla et al. (51) showed that microinjection of mRNA encoding CARM1 (coactivator arginine methyltransferase 1) in one mouse blastomere at the 2-cell stage causes a developmental bias. At the blastocyst stage, the cell progeny of the injected blastomere is found enriched, albeit not exclusively localized, in the ICM. Notably, deletion of the CARM1 gene allows mouse development to E18.5 (52). Because depletion of CARM1 does not impair development to near term, increase, rather than reduction, of this factor appears to be more consequential for pluripotency. As CARM1 is normally present in the embryo, it would be even more interesting to introduce factors that are lacking in cloned embryos, such as Oct4.
In our so far unpublished studies, we have found that the level of Oct4 protein is 40% lower in cloned than in fertilized mouse embryos produced by intracytoplasmic sperm injection (ICSI) (Fig. 1). This is in line with the report that cloned mouse embryos exhibit correct timing but the wrong level of gene expression (53). The immunoblotting method used to measure the amount of Oct4 in embryos is obviously destructive, thereby preventing test hypotheses on subsequent development. Therefore, we sought to measure embryonic pluripotency without consuming the embryos for the assay. This requires a suitable reporter.
Quantitation of Oct4 in nuclear transfer (NT) and intracytoplasmic sperm injection (ICSI) mouse embryos by immunoblotting. Calibrator: mouse embryonic stem (ES) cells. Antibody anti-Oct4 from Santa Cruz (sc-5279). (a) Western blotting after SDS-PAGE (b) histogram of the western blot in (a) after densitometry, presenting the amount of Oct4 per developmental unit (one embryonic stem cell [ESC] or one oocyte or one embryo). Lanes of the gel: 1: 5,000 ESC; 2: 10,000 ESC; 3: 25,000 ESC; 4: 50,000 ESC; 5: 120 NT morulae; 6: 120 ICSI morulae; 7: 120 NT blastocysts; 8: 120 ICSI blastocysts; 9: 50,000 ESC; 10: 25,000 ESC; 11: 10,000 ESC; 12: 5,000 ESC; 13: 140 oocytes; 14: 140 8-cell NT embryos; 15: 140 8-cell ICSI embryos; 16: 50,000 ESC; 17: 25,000 ESC; 18: 10,000 ESC; 19: 5,000 ESC
Visualization of Mouse Embryo Pluripotency Using the Oct4-GFP Transgene
Transgenic strains of mice with the green fluorescent protein (GFP) as a reporter allow tracking of embryonic blastomeres and discrimination between the G1 and G2 phase of their cell cycle (H2B-GFP (54)), monitoring of X-chromosome activity (X-linked GFP (55)), or monitoring of organogenesis (Hox-GFP (56)). As of May 2007, as many as 53 GFP mouse strains were available from the Jackson Laboratory (Bar Harbor, Maine, USA). Among them is one strain that carries GFP under control of the Oct4 promoter.
An 8.5 kb DNA region upstream the start codon of mouse Oct4 gene was found to be capable to drive expression of β-galactosidase (LacZ) indistinguishable from endogenous Oct4 (57). This region contains regulatory sequences, chiefly the proximal enhancer (PE) which is relevant to expression in the epiblast, and the distal enhancer (DE), which is relevant to expression in preimplantation stages and in germ cells. To allow for live cell imaging, the LacZ-encoding sequence was replaced with enhanced GFP (EGFP). The construct was used to generate a transgenic mouse strain known as GOF18 (58). In a subsequent variant, the 8.5 kb DNA control region was shortened by excision of the PE (59). Linked to EGFP, the shortened region was used to generate a transgenic mouse strain known as OG2 (GOF18-∆PE-EGFP; deposited at the Jackson Laboratory as B6;CBA-Tg(Pou5f1-EGFP)2Mnn/J (60)) (Fig. 2). By further reducing the size of the Oct4 control region in the transgene, Hübner et al. (7) obtained a marker solely for germ cells, termed gc-Oct4, which provides another tool to track primordial germ cells (61, 62).
It should be emphasized that both the GOF18 and OG2 mouse strains are transgenic models that have both advantages and disadvantages. Although knock-in reporters obtained by homologous recombination allow faithful regulation of the reporter by the full set of genomic control sequences, this usually impairs one of the two alleles and thus introduces possible dosage effects. Some genes are indeed haplo-insufficient (e.g., Berf-1 (63)). Transgenes instead may not always be faithful mirrors of the endogenous gene expression, but they also do not directly interfere with its function. The Oct4 promoter is silent in sperm and somatic cells, and its activation provides a direct measure of the reprogramming process. Work from our laboratory until recently (64) has shown that the quantitative expression of GOF18-∆PE-EGFP transgene is proportional to that of the endogenous Oct4. This, together with the fact that OG2 probably has only two insertions in the genome (as detected by Southern blotting of EcoR1 digest with GFP probe; Konstantinos Anastassiadis, personal communication), portrays GOF18-∆PE-EGFP as a valid tool to monitor nuclear reprogramming.
Live Cell Confocal Imaging of Cloned Mouse Embryos Expressing Oct4-GFP (GOF18-∆PE-EGFP)
To study embryonic potential in relation to Oct4 level, we produced mouse embryos that carry the GOF18-∆PE-EGFP transgene from either sperm of homozygous (t/t) donors and wild-type oocytes (ICSI) or cumulus cell of hemizygous (t/+) donors (nuclear transfer; NT) so as to have the same number of copies of GOF18-∆PE-EGFP in both cloned and fertilized embryos.
In previous work, we used wide-field fluorescence microscopy and qualitative criteria to score the pattern of GOF18-∆PE-EGFP in live mouse blastocysts (96 hpa) cloned from OG2 cumulus cells (65). Because our checkpoint was at least 24 h past the onset of Oct4 expression at the morula stage as we scored the blastocysts, we might have missed information about Oct4 induction. One problem with early assessment is that early stages of development are more sensitive to manipulation including imaging and photo damage. Confocal microscopy might serve our purpose as it has been shown to preserve the full developmental potential of preimplantation embryos (66).
Using the Perkin-Elmer UltraView RS3 system, we first detected GOF18-∆PE-EGFP fluorescence at the 4-cell stage and were able to score the pattern of GOF18-∆PE-EGFP in live cloned morulae (78 hpa) in a quantitative manner (Fig. 3a–c). Thanks to a spinning (>1,800 rpm) disk with 20,000 pinholes that let only 1–4% of the incident light pass through, the energy delivered to the embryo is very low (below the saturation threshold of the fluorophore), yet an image is produced due to the high quantum efficiency (up to 75%) of a charge-coupled device (CCD) light detector. A Nikon CFI Plan Apochromat VC 60× WI objective lens (N.A. 1.20) was used to convey 488-nm laser excitation to the GFP-expressing morula. The source was a three-line (488 nm, 568 nm, 647 nm) Argon/Krypton laser (Melles Griot). No auxiliary lens (e.g., optovar) or optical device (e.g., filter) was present in the optical path except for the UltraView’s own, and the laser was not attenuated (100% AOTF). Optical sections were captured using a 1.3 megapixel Hamamatsu ORCA ER digital camera with standardized settings (999 ms camera exposure, 2 × 2 binning hence 672 × 512 image pixels, no electronic gain).
A total of 794 morulae cloned from OG2 t/+ cumulus cells and 546 morulae obtained from fertilization with sperm of OG2 t/t males were imaged. Morulae were placed individually in 1.0 μL drops of α-MEM arrayed 5 × 5 on a 50-mm thin-bottom plastic dish (Greiner Bio-One, Lumox dish, catalog 96077303) overlaid with mineral oil (Sigma catalog M5310). A Tokai-Hit environmental minichamber maintained the dish in a gas phase of 5% CO2 at the temperature of 37°C during the time of imaging (about 20 min. per dish).
Photodamage to the embryo would be readily apparent from the inability of the morula to form a blastocyst (cavitation) within 10 h (Fig. 3a–c). To test this, we compared imaged to non-imaged morulae. Not only the rates of morula-to-blastocyst transition were very similar in imaged versus nonimaged morulae, but cloned morulae exposed to GFP excitation formed fetuses at a similar rate to unexposed controls (64).
GFP Image Analysis
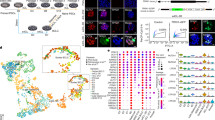
For each morula, five fluorescence confocal sections were captured 5 μm apart from each other in the equatorial region. In order to match the pattern of GFP in morulae with the ability to form blastocyst, combined (maximum projection) images were analyzed with the software ImageJ (67). It was obvious that images had complex patterns (Fig. 3). Our group has shown that “patches” of higher or lower GFP intensity in the morula are not contributed by mosaic aneuploidy (50), thereby warranting further analysis with ImageJ. A region of interest (ROI) was drawn by hand using the polygon selection tool of ImageJ including only pixels belonging to the embryo. For each maximum projection, we measured 15 image parameters extracted by ImageJ and we performed a principal component analysis (PCA) to check whether the images of embryos contain information about the probability of embryo survival. The proportion of variability explained by the first three principal components was 84.6% for the ICSI morulae and 80.6% for the NT morulae. We plotted the components in three-dimensional projections (Fig. 4a, a′) and observed that the nonsurvival cases (in black) tend to have lower values of their first principal component than the survival cases (in white). To better expose the degree to which the first three principal components of the survival population overlap with the components of the nonsurvival population, we performed a Delaunay triangulation. Although the tetrahedrons of the non-survival (in black) and survival cases (in white) overlap in a central region, they cover distinguishable nonsurvival and survival regions (Fig. 4b, b′).
Principal component analysis of mouse morulae expressing GOF18-∆PE-EGFP. (a, a′) nuclear transfer (NT) morulae. (b, b′) intracytoplasmic sperm injection (ICSI) morulae. The spheres in (a, b) correspond to the three-dimensional projection of the principal components, and the tetrahedrons in (a′, b′) represent their connection using the Delaunay triangulation. White colors represent survival cases; black colors, nonsurvival cases
Given the results of the PCA, we focused on searching the more informative image parameter and found that this was GFP intensity. Distributions of GFP intensity were noticeably different in clones and ICSI morulae, and the average GFP intensity was lower in clones than ICSI (Fig. 5). This was confirmed by immunoblotting of the endogenous Oct4 protein. A total of 794 cloned and 546 ICSI morulae were allowed to develop to blastocyst, while the corresponding images were processed with ImageJ.
Intervals of GOF18-ΔPE-EGFP Intensity Define Morulae that Have Distinct Blastocyst Potentials (Subsets)
In the following experiments, we used only GFP intensity as the image component that gives the most information about embryonic potential. Absolute intensity values of GFP depend on biological as well as non-biological factors, such as type and age of the light source, objective lens, and camera, which are difficult to reproduce exactly in different laboratories. For better reproducibility, we ranked the morulae by GFP intensity (Fig. 5) and allocated them into quartile intervals:
-
1.
0–25th percentile, low GFP, subset 1
-
2.
26th–50th percentile, low-medium GFP, subset 2
-
3.
51th–75th percentile, medium-high GFP, subset 3
-
4.
76th–100th percentile, high GFP, subset 4
On the day following imaging, the records of image analysis and blastocyst formation were matched. Table 1 shows the data from 176 ICSI and 316 NT morulae that were analyzed by quartile allocation, until practical reasons emerged and called for the introduction of tertiles (see the section “Success in Deriving ES Cell Lines Increases with Increased GOF18-∆PE-EGFP Intensity of Morulae”). Blastocyst formation was associated with GFP intensity in NT morulae (r 2 = 0.42) more than in ICSI morulae (r 2 = 0.26) (Table 1).
Subsets of Morulae Defined by GOF18-∆PE-EGFP Contain Specific Levels of Oct4, Nanog, Sox2, and Cdx2 Transcripts
An important question in oocyte-mediated nuclear reprogramming is whether genes are reprogrammed independently of one another. Thus we analyzed the GFP subsets for expression of the endogenous genes Oct4, Nanog, and Sox2 (pluripotency- and ICM-associated), and Cdx2 (TE-associated) (64). Levels of mRNA (cDNA) were measured using TaqMan real-time PCR and normalized to the endogenous control Hprt (Table 2).
In general, correlation with GFP intensity was substantial for all genes analyzed. Except for Cdx2, cloned morulae had lower mRNA values than those of ICSI morulae, with Sox2 mRNA at bare levels (p < 0.01). Differences of mRNA levels between cloned and ICSI blastocysts were less prominent; only for Sox2 was the difference significant. This led us to speculate (64) that embryonic potential might be limited by the factor present in the least amount, reminiscent of the Liebig’s Law of the Minimum, a principle developed in agricultural science whereby plant growth is limited by the scarcest resource available. It would therefore be interesting to increase the level of Sox2 in the embryo.
Success in Deriving ES Cell Lines Increases with Increased GOF18-∆PE-EGFP Intensity of Morulae
Since the blastocyst rate increases with higher intensity of GOF18-∆PE-EGFP, we hypothesized that the level of Oct4 in the embryo could have an effect on the pluripotent founder cells in the ICM and ES cell derivation. Indeed, a statistically not significant increase of the total cell number was observed in cloned blastocysts that had Oct4 riboprobe signal localized more prominently to the putative ICM. We set out to follow this up by deriving ES cells.
ES cell derivation was conducted according to a standard protocol (44, 64). Since derivation is labor-intensive and a substantial number of lines is required for each subset, we reduced the number of subsets from four to three. Practically, the morulae were scored for GFP as described, and allocated to tertile instead of quartile intervals of GFP intensity (0–33%, subset 1; 34–67%, subset 2; 68–100%, subset 3).
Overall, 104 and 21 ES cell lines were derived from NT and ICSI embryos, respectively. NT-ES cell derivation was more efficient (χ 2 test, p = 0.000012) for clones of GFP subsets 3 and 2 (36%, 27%) than for subset 1 (9% of blastocysts form ES cells). ES cell derivability was almost equal for ICSI embryos across the three groups (χ 2 test, p = 0.49). All ES cell lines displayed characteristic morphology of undifferentiated ES cells. The pluripotent status was confirmed by immunohistochemical analysis of SSEA-1, alkaline phosphatase (AP), and Oct4 proteins.
ES cells derived from cloned or fertilized embryos are held to be transcriptionally and functionally indistinguishable (68, 69). However, Gidekel et al. (27) reported that ES cells transplanted in vivo behave differently depending on the residual amount of Oct4. To probe if the different rates of ES cell derivation from embryos that had different amounts of Oct4 corresponded to qualitative differences of the ES cells, we allowed them to differentiate in vivo by injecting them into recipient blastocysts followed by embryo transfer.
Six lines per subset were selected for prevalent normal karyotype (4 of 6 lines, 3 of 6 lines, and 4 of 6 lines of GFP intervals 1, 2, 3, respectively), each presenting greater than 50% euploid chromosome sets. Approximately 15 cells were injected into fertilization-derived blastocysts: 286, 347, and 334 blastocysts were injected with ES cells of GFP subset 1, 2, and 3, respectively. The injected blastocysts were transferred into uteri of pseudopregnant mice. On average, 17% of the injected blastocysts had formed proper fetuses at 14.5 dpc (166 of 967) and 28% fetuses had GFP-positive gonads (46 of 166). Distribution of the chimeric fetuses according to the subset of origin of the ES cells was unbiased (n = 54, 61, 51). Germline contribution was observed regardless of the GFP subset (subset 1, 28%; subset 2, 38%; subset 3, 16%; defined as the proportion of fetuses with GFP+ gonads).
Although our data corroborate the view that ES cells derived from cloned or fertilized embryos are indeed indistinguishable from each other, we maintain that further investigation is necessary. In two recent studies, NT-ES cells were found to have higher aneuploidy rates than ES cells from fertilized embryos (50), and the nuclear transfer procedure was found to leave a specific mark on NT-ES cells (70).
Manipulating Pluripotency in Cloned Mouse Embryos
Oct4 is deficient in cloned mouse embryos after ooplasm-mediated induction of pluripotency (broad literature, e.g., Sebastiano et al. (53)) and clones with lower or higher Oct4 level attain lower or higher developmental rates, respectively (64). Oct4 exhibits a dose–effect response in pluripotent ES cells (6), and it is among the three or four factors that converted mouse and human fibroblasts into ES-like cells after retrovirus-mediated delivery to the nucleus (17, 37–39). For these reasons, we became interested whether the level of Oct4 is a determinant of blastomere fate in normal and cloned mouse embryos.
It is possible that subsets of cloned mouse embryos with low, intermediate, and higher levels of GOF18-∆PE-EGFP represent discrete stages in the induction of pluripotency. Microinjection of the arginine methyltransferase CARM1 mRNA into blastomeres has recently been shown to be feasible and resulted in alteration of their fate (51). Therefore, using a similar approach we decided to examine the impact of changing the Oct4 level on ooplasm-induced pluripotency. To avoid any genetic manipulation, we did not use retrovirus-mediated delivery of Oct4 cDNA, which could result in insertional mutagenesis. Instead, we implemented a dual strategy based on microinjection of known amounts of Oct4 mRNA or recombinant protein into the ooplasm prior to somatic nuclear transfer or into a blastomere of a 2-cell stage embryo. This approach allows us to monitor the effects of changing Oct4 levels without modifying the genome of the embryo.
For recombinant Oct4 (rOct4) production, we expressed a GST-Oct4-His construct in the Escherichia coli BL21 strain and purify the recombinant protein in two-steps, using the Äkta Purifier chromatographic system (GE Healthcare). First, GST-Oct4-His was captured by Ni-affinity chromatography, eluted by high concentrations of imidazole and the GST tag cleaved by thrombin. In the second step, the resulting Oct4-His protein was purified from thrombin, imidazole, and other contaminants by size-exclusion chromatography. To verify the activity of the rOct4 obtained in this way, we made use of 2-cell stage embryos derived from the GOF18-(∆PE)-EGFP transgenic mouse described above. If the rOct4 injected is functional, it can bind to the Oct4 promoter and induce higher levels of expression of EGFP as compared to embryos injected with recombinant GST as a control.
In a pilot experiment, one blastomere of the 2-cell stage mouse embryo was injected with rOct4 protein, along with a tracer (Alexa 488 dextran beads 40 kDa) (Fig. 6). Blastocyst formation was not affected by the procedure. The injected blastomere was tracked to see its contribution to the ICM or TE. We examined six blastocysts and observed 4 instances of blastocysts with the green fluorescent tracer in the ICM and 2 instences in the TE (2). Torres-Padilla et al. (51) showed that the pattern of the second cell division (meridian, equatorial) determines the shape of the 4-cell embryo, and according to this shape, the allocation of blastomere in the blastocyst is biased. Therefore, it will be very interesting to track the cleavage timing and geometry of the injected blastomere.
In parallel to this approach, we have also implemented a second strategy, based on microinjection of in vitro transcribed mRNA encoding for Oct4. We have cloned the Oct4 cDNA in the pcDNA3.1-FLAG vector, which allows its in vitro transcription by the bacteriophage T7 polymerase. The resulting mRNA is then in vitro polyadenylated at the 3′ and capped at the 5′ terminus and finally injected in 1- or 2-cell stage embryos. The ability of the produced mRNAs to be properly translated will be tested using the reticulocyte lysate system (Promega) and 1- or 2-cell stage embryos derived from GOF18-(∆PE)-EGFP transgenic mice, as for rOct4. Compared with the previous strategy, this method offers the advantage of being able to deliver mRNAs encoding not only for wild-type Oct4, but potentially for any mutant of interest. In addition, this method offers a higher precision in the amount of mRNA delivered (measured by a Nanodrop® system) as compared to the injection of rOct4 protein, whose amount we roughly estimate using the Bradford method. Lastly, the injection of both rOct4 and of mRNA appears to be advantageous over the use of retro- or lentiviruses because there are no potential issues of gene reactivation at later stages and because the time window in which the exogenous protein will be present in the embryo is short and defined. Although we do not know the half-lives of our rOct4 or Oct4 mRNA in the embryo, experiments can be designed in which various amounts of proteins and mRNA are injected. In any case, the effects of our method are likely to be transient and limited to the very early stages of development, thus allowing us to understand the impact of transiently modulating the levels of Oct4 in the zygote or in the 2-cell stage embryo.
The combination of the above described two methods will hopefully provide exciting insights in the effects of engineering Oct4 levels in the early embryo without affecting its genome. A major question that still needs to be addressed is what the consequences of transiently changing Oct4 levels in a blastomere of a 2-cell stage embryo would be. Will the subsequent cleavage pattern be influenced? Will the fate of the injected blastomere be biased? Our strategy will possibly also allow us to map the region(s) in Oct4 responsible for a given effect by injecting mRNAs encoding for certain point mutants or deletions of Oct4.
What GOF18-ΔPE-EGFP May and May Not Tell Us About Nuclear Reprogramming
We used the GOF18-ΔPE-EGFP transgenic tool to investigate why gene expression of cloned embryos does not resemble that of fertilized embryos. In some cloned embryos, somatic cell-encoded GFP under control of the GOF18-ΔPE promoter is visible at the late 4-cell stage, i.e., 48 h after nuclear transfer. This is rather unexpected based on a previous report where GOF18-GFP and GOF18-ΔPE-EGFP were detected at the 8-cell stage after fertilization (58). In case of reprogramming by fusion with ES cells, somatic GOF18-GFP is reactivated 36–48 h after fusion with ES cells (30). In both cases, nuclear transfer and cell fusion data imply that two to four cell cycles are sufficient to enable significant induction of pluripotency in the somatic genome. However, the level of expression, whether mRNA or protein, is lower than normal while the timing is correct (53). This suggests that erasure of somatic, repressive mechanisms is partial, and only some of the gene regulatory elements can be used by the cellular machinery of the ooplasm. It is well possible that repeated exposure of the somatic genome to reprogramming factors is a key event in the erasure of somatic cell memory. Therefore, it will be interesting to test if preventing cell cycle progression after nuclear transfer allows for the expression of pluripotency-associated genes. Expression of these genes when the cell cycle is blocked would be a strong indication that the ooplasm can actively reprogram the nucleus transplant.
The underlying mechanism to reprogramming is not known, but there are clues to it. Pronuclear mouse eggs microinjected with unmethylated GOF18-ΔPE-EGFP plasmid show GFP expression already at the 2-cell stage (71), therefore, the mechanism underlying reprogramming may involve DNA demethylation. This is in accord with our observation that the Oct4 DE is more methylated in cloned morulae when GOF18-ΔPE-EGFP levels are lower (64). Thus it would be sensible to reduce the DNA methylation of the somatic genome prior or immediately after nuclear transfer. Regardless of the agent used, genomic imprints relying on CpG methylation should not be modified, especially if applications in tissue regeneration are envisioned. In fact, Tada et al. (30) showed that when using embryonic germ (EG) cells for fusion, imprints are erased, while they are retained when ES cells are used.
In spite of lower expression levels, Oct4 and GOF18-ΔPE-EGFP are informative of the developmental potential of cloned morulae. GOF18-ΔPE-EGFP level predicts development to blastocyst, and the correlation is higher for cloned than for fertilized embryos (64). Differences in GOF18-ΔPE-EGFP expression are mirrored by the expression of the pluripotency-related transcripts Oct4, Nanog, Sox2, and Cdx2. There are clones that appear indistinguishable from fertilized embryos, but they are far less competent to develop to later stages in vivo than fertilized counterparts. In contrast, ES cell formation is less clear-cut. Although the rate of NT-ES cell derivation was proportional to GFP intensity, no differences could be detected subsequently. Established ES cells might always contribute to organogenesis in vivo after prior selection for in vitro proliferation during derivation. Indeed ES cells derived from cloned blastocysts are reported to be equivalent to those derived from fertilized blastocysts (68, 69). It would be interesting to know, before selection occurs, if precursor ICM cells give rise to NT-ES cells in a similar or different way than ICM cells of normal embryos. The different responses of clones and ICSI embryos to Oct4 level suggest that the dose–effect response does not only apply to ES cells but also to embryo development, albeit in different ways. Although some aspects are conserved between natural and induced pluripotency, other aspects are distinct.
In conclusion, flawed resetting of epigenetic marks may explain why gene expression levels in clones are abnormal. Given the 30,000 genes that make up the mouse genome and the long list of genes that are dysregulated besides Oct4, it seems unlikely that reprogramming is only random and subject to selection, otherwise no embryo would survive. The “antichaos” model proposed by Stuart Kauffman (72) explains how a complex system that is disordered at the beginning spontaneously organizes into a high degree of order. We propose that all embryos are inherently dysregulated at the outset and self-organize out of such chaos. Self-organization needs a certain amount of time, and a somatic nucleus transplant commences this process from a more distant starting point than a gamete. Most cloned embryos form blastocysts in vitro but are doomed in utero, while ES cells derived from those blastocysts appear normal. This suggests that the in vivo environment allows minimal flexibility, whereas the in vitro environment allows substantial flexibility.
References
Tarkowski AK, Ozdzeński W, Czołowska R. How many blastomeres of the 4-cell embryo contribute cells to the mouse body? Int J Dev Biol 2001;45:811–6.
Tarkowski AK, Ozdzenski W, Czolowska R. Identical triplets and twins developed from isolated blastomeres of 8- and 16-cell mouse embryos supported with tetraploid blastomeres. Int J Dev Biol 2005;49:825–32.
Chung Y, Klimanskaya I, Becker S, et alet al. Embryonic and extraembryonic stem cell lines derived from single mouse blastomeres. Nature 2006;439:216–9.
Rossant J. Postimplantation development of blastomeres isolated from 4- and 8-cell mouse eggs. J Embryol Exp Morphol 1976;36:283–90.
Boiani M, Schöler HR. Regulatory networks in embryo-derived pluripotent stem cells. Nat Rev Mol Cell Biol 2005;6:872–84.
Niwa H, Miyazaki J, Smith AG. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet 2000;24:372–6.
Hübner K, Fuhrmann G, Christenson LK, et alet al. Derivation of oocytes from mouse embryonic stem cells. Science 2003;300:1251–6.
Seguin C, Draper JS. Chapter 11. Extraembryonic differentiation of human ES cell. In: Sullivan S, Cowan CA, Eggan K, editors. Human Embryonic Stem Cells: The Practical Handbook. Wiley, West Sussex, England, 2007:404.
Masui S, Nakatake Y, Toyooka Y, et alet al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat Cell Biol 2007;9:625–35.
Boyer LA, Lee TI, Cole MF, et alet al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005;122:947–56.
Babaie Y, Herwig R, Greber B, et alet al. Analysis of OCT4 dependent transcriptional networks regulating self renewal and pluripotency in human embryonic stem cells. Stem Cells 2006;25:500–10.
Greber B, Lehrach H and Adjaye J. FGF2 modulates TGFβ signaling in MEFs and human ES cells to support hESC self-renewal. Stem Cells 2006;25:455–64.
Reményi A, Schöler HR, Wilmanns M. Combinatorial control of gene expression. Nat Struct Mol Biol 2004;11:812–5.
Hatano SY, Tada M, Kimura H, et alet al. Pluripotential competence of cells associated with Nanog activity. Mech Dev 2005;122:67–79.
Darr H, Mayshar Y, Benvenisty N. Overexpression of NANOG in human ES cells enables feeder-free growth while inducing primitive ectoderm features. Development 2006;133: 1193–201.
Tanaka Y, Era T, Nishikawa S, Kawamata S. Forced expression of Nanog in hematopoietic stem cells results in a gamma delta T-cell disorder. Blood 2007;110:107–15.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006;126:663–76.
Schöler HR, Hatzopoulos AK, Balling R, Suzuki N, Gruss P. A family of octamer-specific proteins present during mouse embryogenesis: evidence for germline-specific expression of an Oct factor. EMBO J 1989;8:2543–50.
Okamoto K, Okazawa H, Okuda A, Sakai M, Muramatsu M, Hamada H. A novel octamer binding transcription factor is differentially expressed in mouse embryonic cells. Cell 1990;60:461–72.
Rosner MH, Vigano MA, Ozato K, et alet al. A POU-domain transcription factor in early stem cells and germ cells of the mammalian embryo. Nature 1990;345:686–92.
Lengner CJ, Camargo FD, Hochedlinger K, et alet al. Oct4 expression is not required for mouse somatic stem cell self-renewal. Cell Stem Cell 2007;1:403–15.
Nichols J, Zevnik B, Anastassiadis K, et alet al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 1998;95:379–91.
Zeineddine D, Papadimou E, Chebli K, et alet al. Oct-3/4 dose dependently regulates specification of embryonic stem cells toward a cardiac lineage and early heart development. Dev Cell 2006;11:535–46.
Ramos-Mejía V, Escalante-Alcalde D, Kunath T, et alet al. Phenotypic analyses of mouse embryos with ubiquitous expression of Oct4: effects on mid-hindbrain patterning and gene expression. Dev Dyn 2005;232:180–90.
Hochedlinger K, Yamada Y, Beard C, Jaenisch R. Ectopic expression of Oct-4 blocks progenitor-cell differentiation and causes dysplasia in epithelial tissues. Cell 2005;121:465–77.
Rodriguez RT, Velkey JM, Lutzko C,et alet al. Manipulation of OCT4 levels in human embryonic stem cells results in induction of differential cell types. Exp Biol Med 2007;232:1368–80.
Gidekel S, Pizov G, Bergman Y, Pikarsky E. Oct-3/4 is a dose-dependent oncogenic fate determinant. Cancer Cell 2003;4:361–70.
Camara-Clayette V, Le Pesteur F, Vainchenker W, Sainteny F. Quantitative Oct4 overproduction in mouse embryonic stem cells results in prolonged mesoderm commitment during hematopoietic differentiation in vitro. Stem Cells 2006;24:1937–45.
Wei F, Schöler HR, Atchison ML. Sumoylation of Oct4 enhances its stability, DNA binding, and transactivation. J Biol Chem 2007;282:21551–60.
Tada M, Takahama Y, Abe K, Nakatsuji N, Tada T. Nuclear reprogramming of somatic cells by in vitro hybridization with ES cells. Curr Biol 2001;11:1553–8.
Ng RK, Gurdon JB. Epigenetic memory of active gene transcription is inherited through somatic cell nuclear transfer. Proc Natl Acad Sci U S A 2005;102:1957–62.
Eggan K, Akutsu H, Hochedlinger K, Rideout W3rd, Yanagimachi R, Jaenisch R. X-chromosome inactivation in cloned mouse embryos. Science 2000;290:1578–81.
Kimura H, Tada M, Hatano S, Yamazaki M, Nakatsuji N, Tada T. Chromatin reprogramming of male somatic cell-derived XIST and TSIX in ES hybrid cells. Cytogenet Genome Res 2002;99:106–14.
Egli D, Rosains J, Birkhoff G, Eggan K. Developmental reprogramming after chromosome transfer into mitotic mouse zygotes. Nature 2007;447:679–85.
Do JT, Schöler HR. Cell–cell fusion as a means to establish pluripotency. Ernst Schering Res Found Workshop 2006;60:35–45.
Matsumura H, Tada M, Otsuji T, et alet al. Targeted chromosome elimination from ES-somatic hybrid cells. Nat Methods 2007;4:23–5.
Takahashi K, Tanabe K, Ohnuki M, et alet al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007;31:861–72.
Yu J, Vodyanik MA, Smuga-Otto K, et alet al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007;318:1917–20.
Nakagawa M, Koyanagi M, Tanabe K, et alet al. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat Biotechnol 2008;26:101–6.
Taranger CK, Noer A, Sørensen AL, Håkelien AM, Boquest AC, Collas P. Induction of dedifferentiation, genome wide transcriptional programming, and epigenetic reprogramming by extracts of carcinoma and embryonic stem cells. Mol Biol Cell 2005;16:5719–35.
Ding S, Schultz PG. Small molecules and future regenerative medicine. Curr Top Med Chem 2005;5:383–95.
Patsch C, Edenhofer F. Conditional mutagenesis by cell-permeable proteins: potential, limitations and prospects. Handbook Exp Pharmacol 2007;178:203–32.
Fittipaldi A, Giacca M. Transcellular protein transduction using the Tat protein of HIV-1. Adv Drug Deliv Rev 2005;57:597–608.
Cavaleri F, Gentile L, Schöler HR, Boiani M. Recombinant human albumin supports development of somatic cell nuclear transfer embryos in mice: toward the establishment of a chemically defined cloning protocol. Cloning Stem Cells 2006;8:24–40.
Kues WA, Niemann H. The contribution of farm animals to human health. Trends Biotechnol 2004;22:286–94.
Fulka JJr, Miyashita N, Nagai T, Ogura A. Do cloned mammals skip a reprogramming step. Nat Biotechnol 2004;22:25–6.
Bortvin A, Eggan K, Skaletsky H, et alet al. Incomplete reactivation of Oct4-related genes in mouse embryos cloned from somatic nuclei. Development 2003;130:1673–80.
Li X, Kato Y, Tsunoda Y. Comparative analysis of development-related gene expression in mouse preimplantation embryos with different developmental potential. Mol Reprod Dev 2005;72:152–60.
Li X, Amarnath D, Kato Y, Tsunoda Y. Analysis of development-related gene expression in cloned bovine blastocysts with different developmental potential. Cloning Stem Cells 2006;8:41–50.
Balbach ST, Jauch A, Böhm-Steuer B, Cavaleri FM, Han YM, Boiani M. Chromosome stability differs in cloned mouse embryos and derivative ES cells. Dev Biol 2007;308:309–21.
Torres-Padilla ME, Parfitt DE, Kouzarides T, Zernicka-Goetz M. Histone arginine methylation regulates pluripotency in the early mouse embryo. Nature 2007;445:214–8.
Yadav J, Lee J, Kim J, et alet al. Bedford. Specific protein methylation defects and gene expression perturbations in coactivator-associated arginine methyltransferase 1-deficient mice, Proc Natl Acad Sci U S A 2003;100:6464–8.
Sebastiano V, Gentile L, Garagna S, Redi CA, Zuccotti M. Cloned pre-implantation mouse embryos show correct timing but altered levels of gene expression. Mol Reprod Dev 2005;70:146–54.
Hadjantonakis AK, Papaioannou VE. Dynamic in vivo imaging and cell tracking using a histone fluorescent protein fusion in mice. BMC Biotechnol 2004;4:33.
Hadjantonakis AK, Gertsenstein M, Ikawa M, Okabe M, Nagy A. Non-invasive sexing of preimplantation stage mammalian embryos. Nat Genet 1998;19:220–2.
Godwin AR, Stadler HS, Nakamura K, Capecchi MR. Detection of targeted GFP-Hox gene fusions during mouse embryogenesis. Proc Natl Acad Sci U S A 1998;95:13042–7.
Yeom YI, Fuhrmann G, Ovitt CE, et alet al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 1996;122:881–94.
Yoshimizu T, Sugiyama N, De Felice M, et alet al. Germline-specific expression of the Oct-4/green fluorescent protein (GFP) transgene in mice. Dev Growth Differ 1999;41:675–84.
Nordhoff V, Hübner K, Bauer A, Orlova I, Malapetsa A, Schöler HR. Comparative analysis of human, bovine, and murine Oct-4 upstream promoter sequences. Mamm Genome 2001;12:309–17.
Szabo PE, Hübner K, Schöler H, Mann JR. Allele-specific expression of imprinted genes in mouse migratory primordial germ cells. Mech Dev 2002;115:157–60.
Anderson R, Fässler R, Georges-Labouesse E, et alet al. Mouse primordial germ cells lacking β1 integrins enter the germline but fail to migrate normally to the gonads. Development 1999;126:1655–64.
Molyneaux KA, Stallock J, Schaible K, Wylie C. Time-lapse analysis of living mouse germ cell migration. Dev Biol 2001;240:488–98.
Takeuchi A, Mishina Y, Miyaishi O, Kojima E, Hasegawa T, Isobe K. Heterozygosity with respect to Zfp148 causes complete loss of fetal germ cells during mouse embryogenesis. Nat Genet 2003;33:172–6.
Cavaleri F, Balbach ST, Gentile L, et alet al. Subsets of cloned mouse embryos and their non-random relationship to development and nuclear reprogramming. Mech Dev 2008;25:153.
Boiani M, Eckardt S, Schöler HR, McLaughlin KJ. Oct4 distribution and level in mouse clones: consequences for pluripotency. Genes Dev 2002;16:1209–19.
Ross PJ, Perez GI, Ko T, Yoo MS, Cibelli JB. Full developmental potential of mammalian preimplantation embryos is maintained after imaging using a spinning-disk confocal microscope. Biotechniques 2006;41:741–50.
Abramoff MD, Magelhaes PJ, Ram SJ. Image processing with ImageJ. Biophoton Int 2004;11:36–42.
Brambrink T, Hochedlinger K, Bell G, Jaenisch R. ES cells derived from cloned and fertilized blastocysts are transcriptionally and functionally indistinguishable. Proc Natl Acad Sci U S A 2006;103:933–8.
Wakayama S, Jakt ML, Suzuki M, et alet al. Equivalency of nuclear transfer-derived embryonic stem cells to those derived from fertilized mouse blastocysts. Stem Cells 2006;24:2023–33.
Hikichi T, Wakayama S, Mizutani E, et alet al. Differentiation potential of parthenogenetic embryonic stem cells is improved by nuclear transfer. Stem Cells 2007;25:46–53.
Kirchhof N, Carnwath JW, Lemme E, Anastassiadis K, Schöler H, Niemann H. Expression pattern of Oct-4 in preimplantation embryos of different species. Biol Reprod 2000;63:1698–705.
Kauffman SA. Antichaos and adaptation. Sci Am 1991;265:78–84.
Acknowledgments
We are grateful to Dr. Konstantinos Anastassiadis (Technical University Dresden) for sharing unpublished data on GOF18-ΔPE-EGFP; to Dr. Yong-Mahn Han (KAIST, Korea) for contributing to the article of Cavaleri et al., 2008 (64) which provides the groundwork for this chapter; and to Prof. Ivan Dikic (Goethe University, Frankfurt am Main) for supporting the quantitative analysis of Oct4 protein in preimplantation mouse embryos and the synthesis of recombinant Oct4 for microinjection.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Humana Press, a part of Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Balbach, S.T. et al. (2009). Observing and Manipulating Pluripotency in Normal and Cloned Mouse Embryos. In: Baharvand, H. (eds) Trends in Stem Cell Biology and Technology. Humana Press. https://doi.org/10.1007/978-1-60327-905-5_7
Download citation
DOI: https://doi.org/10.1007/978-1-60327-905-5_7
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-60327-904-8
Online ISBN: 978-1-60327-905-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)