Abstract
Autophagy is an evolutionarily conserved biological phenomenon related to protein degradation and organelle turnover. Three types of autophagy have been defined: macroautophagy, microautophagy, and chaperone-mediated autophagy, which differ the way of in the delivery of substrates to the lysosome. In macroautophagy, substrates are wrapped in a double membrane structure, called the autophagosome. The formation of the autophagosome and its fusion with the lysosome are genetically controlled by a series of autophagy molecules and are activated in response to a number of environmental cues. Much has yet to be learned about the signaling pathway and the molecular mechanisms about this process. Autophagy is important to multiple cellular functions, particularly for nutrient and energy balance, and the turnover of cellular substances. The relationship of autophagy with cell death is complicated and may be context-dependent. Whereas the nature of autophagic death has yet to be carefully defined, it seems that autophagy may, in fact, be a key regulator of both apoptosis and necrosis. In this context, the roles of macroautophagy in both prosurvival and prodeath have been identified. Understanding the circumstance in which autophagy affects cell functions and therefore cell viability is critical for the future intervention of this process to control cancer, tissue injury, and other disease processes.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Autophagy
- Macroautophagy
- Cell death
- Apoptosis
- Necrosis
- Atg molecules
- Bcl-2 family proteins
- Starvation
- Metabolic stress
- Mitophagy
- ER stress
- Hypoxia
Introduction
Two major protein degradation systems are present in eukaryotic cells: the proteasome and the lysosome. They differ in their functional significance and the type of substrates they take in for degradation. In the lysosome system, the degradation of extracellular materials is mediated by endocytosis (heterophagy), whereas the degradation of intracellular components is mediated by three types of autophagy: macroautophagy, microautophagy, and chaperone-mediated autophagy (CMA) (1–7), which differ in how the cytoplasmic materials are delivered to the lysosome. In macroautophagy, the content is sequestered in a double-membrane structure called the autophagosome, which subsequently fuses with the lysosome. In microautophagy, the content is directly taken up by the lysosomes through membrane invagination, whereas in CMA, the content binds to Hsc70 and its co-chaperones. The complex then binds to LAMP2a on the surface of the lysosome. The substrate protein is then transported into the lysosome (6). This chapter discusses macroautophagy (hereafter referred to as autophagy) and its relationship with cell death in mammalian cells.
The term autophagy comes from Greek, meaning “self-eating.” Autophagy as a biological phenomenon was first systemically described by de Duve and Wattiaux 40 years ago (1), although it seems that the phenomenon, as a process of bulk segregation of cellular constituents, was reported as early as in 1957 in mammalian cells (8). Unlike the ubiquitin proteasome system, autophagy is responsible for the degradation of long-lived proteins and is the only system that can degrade organelles, such as mitochondria (4, 9). Although autophagy has long been recognized, progression of the study was slow due to a lack of understanding of its molecular mechanisms. The breakthrough came in the 1990 s when the phenomenon was studied in yeast. The powerful yeast genetics allows the identification of multiple genes required for autophagy (10, 11). These works have since revolutionized the field and brought the research of autophagy into the molecular era.
Basic Autophagy Machinery
Autophagy is evolutionarily conserved and operates in plants, yeast, C. elegans, Drosophila, and mammals. A large portion of the molecular machinery of autophagy is conserved in these organisms (11). Currently, 31 autophagy-related genes (ATG) have been identified since the first gene, Atg1, was discovered from a genetic screening in yeast (12, 13). Detailed discussions of these genes, particularly those of yeast, can be found in several recent reviews (14, 15). A brief summary of the mammalian system is given below.
The core autophagy machinery seems to be built around two ubiquitin-like conjugation systems (3) (Fig. 30.1). In one system, the ubiquitin-like protein Atg12 was first activated by Atg7, a ubiquitin-activating enzyme (E1)-like protein, and then transferred by Atg10, a ubiquitin carrier protein (E2)-like protein, to Atg5 through a covalent bond. The Atg5-Atg12 complex interacts with Atg16 to form a multimer complex, which is localized to membranes of early autophagosomes. It seems that the assembly of this system is independent of autophagy activation. Thus, the complex appears to provide the necessary platform for autophagy activation.
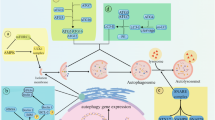
Activation of macroautophagy in mammalian cells. Macroautophagy in mammalian cells could be activated by multiple signals. Some of them are listed at the top of the diagram. The signaling to the core autophagy machinery is only partially revealed, and remains largely unknown to many of the stimuli. The effects of mTOR, Bnip3, calcium, and the unfolded protein response components are among the better-understood mechanisms. The Beclin-1/Atg6 complex, composed of Beclin-1, VPS34 (the class III PI-3 kinase), and its regulatory partner, p150/VPS15, Ambra-1, UVRAG, and its binding partner, Bif-1, seems to act upstream by responding to the stimulation. The activity of this complex can be regulated by a number of other factors, including the Bcl-2 family proteins, and some pharmacological agents that suppress VPS34 enzymatic activity. It is not entirely clear how the Beclin-1 complex positively regulates the two conjugation systems: the Atg12-Atg5-Atg16 and the PE-LC3/Atg8 complexes, which form in the presence of Atg7, an E1-like enzyme, and Atg10 or Atg3, E2-like enzymes. These two complexes are required for the formation of an autophagosome (AV), which is a double-membrane structure. During the formation of AV, the cytosolic materials to be degraded are enclosed within the structure. Autophagosomes are then fused with the lysosomes to form autolysosomes, which degrade the enclosed materials. Both the fusion process and the lysosome activity can be blocked by multiple chemicals as indicated, which can thus block autophagy at a later stage
In another system, the microtubule-associated protein 1 light chain 3 (LC3), or GATE-16 or GABARAP, all mammalian homologues of the yeast Atg8, is first cleaved by Atg4 to expose the conserved Gly120 at its C-terminus. LC3 is then conjugated to phosphatidylethanolamine (PE), also via Atg7, and Atg3, another ubiquitin carrier protein (E2)-like protein (3, 16). The unconjugated form of LC3 (called LC3-I) is in the cytosol, while the conjugated form (called LC3-II) targets to the autophagosomal membrane (16) with the assist of the Atg5-Atg12-Atg16 complex (17). This association of LC3-PE to the autophagosomes is considered important for the membrane extension of the autophagosome and the eventual enclosure of the membrane to form the vacuoles. The Atg5-Atg12-Atg16 complex is recycled, while the LC3 complex stays on the membrane until it is degraded by the lysosome. LC3 is thus widely used as a marker for monitoring the autophagy process.
Several other key autophagy genes are important to the initiation and regulation of autophagy. Beclin-1, a mammalian homologue of Atg6, is particularly important. Beclin-1 forms a complex with VPS34, p150/VPS15. VPS34 is a class III PI-3-kinase that is required for autophagy and can be suppressed by 3-methyladenine (3-MA) and also Wortmannin. In yeast, Atg6 also binds to Atg14, which has not been identified in mammalian cells. However, several prosurvival Bcl-2 family proteins (18) and Ambra-1(19) are found in mammalian cells that can also interact with Beclin-1. However, Bcl-2 suppresses, but Ambra-1 promotes, Beclin-1/VPS34 activity, and therefore autophagy. While the exact mechanism is not clear, the recent finding that Beclin-1 contains a BH3-only domain (18, 20, 21) suggests that Bcl-2 may sequester Beclin-1 from its interaction with VPS34. Finally, while UVRAG has not been found to be instrumental in yeast autophagy, it seems important for mammalian autophagy due to its interaction with Beclin-1 (22). Another recently discovered molecule, Bif-1, promotes autophagy by interacting with UVRAG (23). Thus, there are extensive molecular interactions at the early stage of the formation of autophagosomes (Fig. 30.1).
Activation of Autophagy
A key regulatory of autophagy in yeast and mammalian cells is the TOR complex (5, 24). A functional TOR activity would suppress autophagy. Because the TOR pathway is central to the signaling of growth and energy metabolism, autophagy is intimately coupled with growth and energy control (Fig. 30.1). It can be affected by the upstream signaling of the TOR pathway. For example, growth factors, insulin, and the class I PI3-kinase and Akt will all suppress autophagy by activating Tor (5). On the other hand, suppressing TOR function with rapamycin could induce autophagy.
At the moment, it is not clear how TOR suppression could cause autophagy. In yeast, this seems to be coupled with the activity of the Atg1-Atg13 complex. Although the mammalian cells express two Atg1 homologues, ULK1 and ULK2, they do not seem to possess the Atg13 homologue. Furthermore, genetic deletion of ULK1, unlike that of Atg5 or Atg7, does not result in global defects in autophagy, but specifically affects mitochondria autophagy during erythrocyte maturation (25). It is possible, however, that the Beclin-1 complex would be among the first to be activated by the signals from the TOR suppression.
Several other signaling pathways have been defined in mammalian cells, some of which are likely mTOR-independent (Fig. 30.1). One such pathway can be characterized as calcium-calpain-Gsα–mediated (26). In a number of cases, the induction of autophagy in mammalian cells seems to be related to the transcriptional activation of a number of molecules that are either involved in the core process of autophagy or involved in yet-to-be defined manners. Several transcription factors can be involved. In muscle cells, activation of the FoxO3 transcription by the suppression of the IGF-1-PI-3 K-Akt pathway could in turn enhance the transcription of several autophagy genes, including LC3 and Bnip3 (27, 28). Bnip3 could be also transcriptionally activated by HIF-1, which is induced during hypoxia (29, 30). Bnip3 is responsible for the autophagy induced by hypoxia and ischemia. Furthermore, E2F1 can be responsible for the upregulation of Atg1, Atg5, LC3, and DRAM, which can be important for DNA damage-induced autophagy (31). DRAM is a lysosomal membrane protein participating in autophagy that had previously been identified as a p53 transcriptional target (32). Whereas nucleic p53 could promote autophagy via upregulating DRAM, a recent study has also indicated that cytoplasmic p53 can suppress autophagy by unknown mechanisms (33).
Finally, ER stress caused by chemicals and misfolded protein aggregates can induce autophagy, which helps to relieve ER stress (34–39). In this case, it seems that the unfolded protein response (UPR) pathways, PERK/eIF2alpha and IRE-1/JNK, may be responsible for the autophagy induction. The PERK/eIF2alpha pathway could promote Atg12 expression, which may provide a mechanism of autophagy induction in this case (35). ER stress could elevate the intracellular calcium level, which in turn can activate calmodulin-dependent kinase kinase beta to promote the activity of AMPK, leading to mTOR suppression and autophagy (40). Alternatively, calcium can induce the phosphorylation of PCKθ, which seems to specifically participate in ER-stress-induced, but not in amino acid deprivation-induced, autophagy (41). In this case, the activation of PKCθ is independent of either mTOR or the UPR.
Functional Roles of Autophagy in Mammalian Cells
It is now clear that autophagy is activated under and regulated by many physiological and pathological conditions and, in turn, affects these processes (4, 6, 7, 42). Thus, autophagy is inevitably associated with the pathogenesis of many human diseases (6, 43). It is required for normal development and participates in the clearance of apoptotic cells during embryogenesis (42). In adults, autophagy seems to be involved in the extension of life span and in protecting cells from stress response, such as starvation. The autophagic degradation of cellular constituents can efficiently recycle essential nutrients to sustain basic biological processes. Thus, autophagy is important for the regulation of energy and nutrient metabolism (5, 24). Autophagy is commonly known to be activated by amino acid starvation (in mammalian cells) or nitrogen deprivation (in yeast cells). Under these conditions, autophagy is activated to degrade proteins and recycle amino acids to meet the cell’s energy requirement (5, 10, 24, 44). This function of autophagy in meeting the energy and nutritional needs of cells was evolutionarily conserved. Genetic deletion of Atg5 in mice led to perinatal death due to the lack of sufficient nutrients in the cardiac and diaphragm muscles for their vital functions (45).
In addition, the degradation of mitochondria, peroxisomes, endoplasmic reticulum (ER), or ribosomes by autophagy is most likely associated with cellular homeostasis as well as changing metabolic needs (7, 13, 46). The ability of autophagy to degrade misfolded proteins is an important beneficial function in the pathogenesis of conformational diseases (6, 37). Autophagy is also employed as a defense mechanism to clear up intracellular microbes, misfolded proteins, and damaged organelles (7).
The role of autophagy in cancer development and cancer therapy has been an area of intense study in recent years (47, 48). In the first study aiming to understand the function of Beclin-1/Atg6, a mouse stain deficient in this gene was constructed. While Beclin-1-null mice were embryonic lethal, Beclin-1 heterozygous mice were normal at the beginning, but developed multiple tumors later on, suggesting that Beclin-1 is a haplo-insufficiency tumor suppressor (49). The loss of heterozygosity of Beclin-1 is frequently seen in breast and ovarian cancers (50). While the mechanism of how Beclin-1 serves as a tumor suppressor is not known, it is possible that its proautophagy function is important in maintaining cellular homeostasis by removing damaged organelles, such as mitochondria (51). Damaged mitochondria may become a major source of intracellular free radicals that could cause genomic instability and tumors. As a consequence, autophagy may suppression tumorigenesis by preventing genomic instability (48, 52).
Autophagy in the Regulation of Cell Death
The relationship between autophagy and cell death has been hotly debated in recent years. There are ample observations indicating that the two processes are intimately connected. Evidence for the role of autophagy in promoting cell survival or in promoting cell death is compelling in both cases (53, 54). In addition, it seems that the regulation of autophagy, apoptosis, and necrosis can be coupled so that one type of mechanism may activate or inactivate the other.
Programmed cell death was initially classified into several categories, primarily based on the ultrastructural morphology of the dying cells (55). While one category of PCD demonstrates features of apoptosis, another category shows the accumulation of autophagosomes (56). The latter was classified as autophagic death. However, considerable controversy exists regarding whether the autophagy process actually promotes cell death or instead is a reactive process that may actually provide protection (53–56). These concerns were not just for the developmental biology process, but also for many pathological processes where autophagy is induced significantly in response to cytotoxic or metabolic stress.
With the understanding of the molecular machinery of autophagy, it is now easier to address these issues. Thus, by inhibiting the key autophagy genes through genetic deletion, RNAi-mediated knockdown, or pharmacological interventions, one may determine whether cell death is suppressed, enhanced, or not changed at all. In this way, the influence of autophagy on cell death could be determined. Indeed, depending on the circumstances, both pro- and antideath functions of autophagy could thus be identified.
Autophagy Promotes Cell Survival
There are numerous conditions in which autophagy clearly plays a prosurvival role. In neonatal mice, autophagy is required for the endogenous generation of nutrients in such energy-dependent organs as the heart and diaphragm as the newborn adapts to taking in nutrients from an exogenous source, i.e., milk (45, 57). Autophagy deficiency due to the deletion of key autophagy genes, such as Atg5, can thus lead to the premature death of newborn mice. A similar dependency of survival on autophagy during starvation has been demonstrated in Drosophila, which occurs primarily in the nutrient-sensing organ, the fat body (58, 59). At the cellular level, the importance of autophagy in survival during nutrient or growth factor deprivation can also be shown in mammalian cells (44, 60) and yeast cells (61).
Autophagy is also important for cellular survival under other stressful conditions. In mammalian cells, autophagy could be activated in response to metabolic stress, ischemia, or hypoxia (48, 62). The suppression of autophagy can result in increased cell death. Cytotoxic agents, including many chemotherapeutic agents, such as proteasome inhibitors (39), ER stressors (34–36, 39), DNA-damaging agents (31, 32, 63), and histone deacetylase inhibitor (64), can all activate autophagy, likely in response to the damage caused by these agents. Under pathological conditions, such as the accumulation of misfolded proteins, autophagy is required for the cellular clearance of these proteins and survival (37). In C. elegans, limited food, high temperature, a highly dense population, and mutation in the insulin-like growth receptor (daf-2) could all cause development arrest in the form of dauer diapause, which is specialized for survival under these adverse conditions. Autophagy is required for dauer entry and therefore for the survival of the worm (65).
Autophagy promotes cellular survival through its basic function of degrading intracellular components. In the nutrient/growth factor depletion/deficiency condition, autophagic degradation recycles the cellular proteins and glycogen to provide amino acids and glucose for ATP generation (44). In cells under DNA damage or metabolic stress, autophagy may play an important role in removing damaged organelles, such as mitochondria, to reduce the cellular ROS level and maintain genomic stability (48, 62). The clearance of misfolded proteins resulting from ER stress, proteasome inhibition, or genetic mutation is another important function of autophagy in maintaining cellular viability (37).
The mechanisms of autophagy induction under these different conditions are not all well defined. Growth factor deprivation seems to be linked to the downregulation of the Akt signaling, which leads, on one hand, to the suppression of the Tor signaling and, on the other hand, to the activation of FoxO3, a transcription factor that can cause the upregulation of several autophagy genes (discussed earlier). Both events can lead to autophagy activation.
Autophagy induced by the deprivation of nitrogen in yeast is also critically related to the suppression of Tor signaling (61). However, amino acid deprivation in the mammalian cells does not seem to be completely dependent on the mTOR pathway (5). The PERK/eIF-2α signaling, part of the UPR initiated at the ER, can also contribute (66). This pathway, together with another UPR pathway orchestrated by IRE-1, is also involved in autophagy initiated by the misfolded proteins and ER stress (35, 36, 39). However, in response to proteasome inhibitors, which also cause ER stress and the accumulation of misfolded proteins, only the IRE-1 pathway is required (39). ER stress can lead to calcium release, which in turn can activate calmodulin kinase kinase beta and AMPK to suppress mTOR (40). Interestingly, in these cases, cytoplasmic p53 can suppress the induction of autophagy, thereby ensuring its rapid degradation following the induction (67).
Autophagy-mediated prosurvival function could suppress either apoptosis or necrosis. In many apoptosis-competent cells, autophagy could co-exist with apoptosis, and the suppression of autophagy increases apoptosis (35, 36, 39, 44, 60, 63, 64). However, in apoptosis-incompetent cells, caused by the deletion of key proapoptosis genes, such as Bax and Bak, or the overexpression of antiapoptosis genes, such as Bcl-2 or Bcl-xL, or the use of caspase inhibitors, the inhibition of autophagy often leads to necrosis (48, 68, 69). Many agents could induce both apoptosis and necrosis, although apoptosis can be a dominant type of death, and autophagy can suppress both types. The latter suggests that autophagy acts at the upstream level, where the death stimulation is derived, so it can mitigate the cause for both apoptosis and necrosis. This thinking is consistent with the idea that the clearance function of autophagy is responsible for its prosurvival function by removing the “damaged” cellular content.
Autophagy Can Participate in Cell Death
Although early studies largely employed morphological criteria to define autophagic death, which is subject to verification, more recent works are based on molecular evidence to substantiate the role of autophagy in promoting cell death.
A typical example for autophagy to participate in developmentally regulated programmed cell death is the degradation of the salivary glands during Drosophila’s pupal stage, which is triggered by the steroid hormone ecdysone. Cell death is accompanied with both autophagy and apoptosis features, and determining whether or not autophagy promotes cell death had previously been confusing. A recent study using autophagy gene mutants now indicates that cell death can be significantly inhibited, suggesting that autophagy can promote cell death (70). Furthermore, autophagy is activated by growth arrest and can induce cell death in the absence of caspase activation. In this case, it seems that autophagy and apoptosis independently contribute to the death. Indeed, a combined inhibition of both pathways increased the suppression of the gland degradation (70).
In mammalian cells, autophagy can contribute to cell death under several stressful conditions. Thus, autophagy can be induced in response to certain chemotherapeutic drugs (71, 72), radiation (73), hypoxia (74), ischemia in the brain (75), cytokines such as INF-γ (76), and ligands such as HIV-1-encoded envelope glycoproteins (77). In these cases, the deletion or RNAi-mediated knockdown of key autophagy genes can significantly reduce cell death, while overexpressing these genes can promote it.
How autophagy promotes cell death is not entirely clear. Although it may be tempting to assume that excessive self-digestion could lead to the depletion of key molecules or organelles essential to the process, the mechanism of killing may be as diverse as the stress signals that induce autophagy in the first place. The autophagic machinery may be interfaced with the apoptotic machinery or the necrotic mechanism to promote apoptosis or necrosis. For example, it was found that in INF-γ-induced autophagic death in HeLa cells, Atg5 can bind to FADD and activate caspase-8 and downstream caspases as if there were a death receptor engagement (76). In another case, when Atg5 was overexpressed, it could be cut by calpains. The 24-Kda cleaved Atg5 N-terminal fragment (aa 1-193) is then translocated to the mitochondria, where it binds to Bcl-xL and inactivates it, resulting in cytochrome c release and cell death (71). In these cases, autophagy seems to link to the classical apoptosis pathway; therefore, the death is actually mediated by the apoptosis machinery.
However, in other cases, autophagy-induced cell death is activated by the loss of the ability to mount an effective apoptosis. Thus, in apoptosis-deficient fibroblasts lacking both Bax and Bak, treatment with etopside (72) or prolonged ER stress (78) caused autophagy. Similarly, radiation-induced cell death would switch from apoptosis to autophagic death if caspase activity could be blocked (73). The molecular mechanism is not clear in this type of autophagy-mediated nonapoptotic death, although the role of JNK has been implicated (72). It is likely that cell death in these circumstances is necrotic (78). Similarly, ROS-induced autophagic death can also be necrotic (79). Autophagy-promoted necrosis has also been reported in C. elegans (80).
Interestingly, a number of BH3-only Bcl-2 family proteins or their binding partners, when overexpressed, can directly induce cell death with contributions from the autophagic process. Bnip3 is a BH3-only Bcl-2 family protein that has been shown to induce autophagic death in a number of cases. A more detailed discussion of this molecule is given in the following section, since autophagy induced by this molecule can also be prosurvival in other cases (81). Other BH3-only molecules that can induce autophagic death include Apolipoprotein L1 (ApoL1) (82) and Bik (83). Finally, hSpin1, a human homologue of the Drosophila spinster (spin) gene, can induce nonapoptotic death, which can be inhibited by Bcl-xL (84). The latter effect may not be related to apoptosis regulation but to a direct physical interaction. This type of nonapoptotic autophagy-mediated death is necrotic (84).
One potential mechanism by which BH3-only molecules can induce autophagy is through competitive binding to Bcl-2 or Bcl-xL to disrupt the interaction of the latter with Beclin-1 (Fig. 30.1). Beclin-1 is an important autophagy molecule and interacts with multiple other molecules, such as the class III PI-3 kinase VPS34, UVRAG, and Ambra-1, to promote autophagy (85). However, Beclin-1 also possesses a conserved BH3 domain of the Bcl-2 family proteins and, in fact, can interact with multiple antideath Bcl-2 family members, such as Bcl-2 and Bcl-xL (85), which suppress the function of Beclin-1 as a proautophagy molecule. However, this interaction can be disrupted by the phosphorylation of Bcl-2 or by other BH3-only molecules, such as Bad, or BH3 mimetics, such as ABT737 (see Chapter 2) (85). As a result, Beclin-1 can be desuppressed and can in turn promote autophagy. Indeed, in a human leukemia cell line, HL60, simply knocking down Bcl-2 can cause autophagy, which seems to contribute to the accompanying cell death (86). In the case of Bik-induced autophagic death, the loss of Bcl-2 is a prerequisite (83). Likewise, ApoL1-induced autophagy depends on its BH3 domain, which may thus be involved in the competitive binding with Bcl-2 to release Beclin-1.
Factors That May Affect Whether Autophagy Presents a Prosurvival or Prodeath Effect
The relationship between autophagy and cell death can be quite complicated and may be affected by many factors. Autophagy likely evolves as a physiological process but can be diverted to a prodeath pathway under pathological conditions. A very unique example of autophagy in promoting cellular injury has recently been reported in Atg5-deficient mice (87). Autophagy has been found to be required for the activation of trypsinogen to trypsin under normal conditions. The enzyme is harmful to tissue if its activation and release are not properly controlled, which can be a cause of pancreatitis. Notably, because of autophagy’s role in promoting enzyme maturation, it participates both in normal pancreatic functions, essentially of a “prosurvival” nature, and in pancreatitis in pathological conditions, essentially of a “prodeath” nature.
The level of autophagy could determine the outcome. In C. elegans, it has been suggested that only the physiological level of autophagy during starvation is prosurvival and that excessive autophagy could be prodeath (88). Excessive autophagy can cause cellular atrophy and the deficiency of vital cellular components.
The presence of compensatory mechanisms, such as chaperone-mediated autophagy (CMA), may also determine whether the inhibition of macroautophagy renders cells to be more sensitive or more resistant to certain stressful signals. Thus, murine fibroblasts prepared from Atg5-deficient embryos were more sensitive to death receptor-initiated death, but were more resistant to menadione- and UV radiation-induced death due to increased CMA (89).
In a more general way, whether autophagy can be prosurvival or prodeath can also be dependent on both the agents used for stimulation and the status of the cells. Thus, autophagy induced by ER stress (38) or proteasome inhibitors (Ding and Yin, unpublished observation) is protective in tumor cells but indifferent or detrimental in nontransformed cells. In contrast, ROS-induced autophagy is prodeath in cancer cells but may not be so in nontransformed cells (79).
Cell death and autophagy can be mediated by the same molecule, but through separate mechanisms. Thus, while autophagy and cell death may seem to be coupled by the same molecule, their relationship can be more complicated than a simple interpretation. This point may be best illustrated in the case of Bnip3, a BH3-only Bcl-2 family protein originally identified as an E1B19K and Bcl-2 interacting molecule (90) (see Chapter 2). Bnip3 is usually localized at the mitochondrial outer membrane. Unlike other BH3-only molecules, its transmembrane (TM) domain, but not its BH3-domain, is required for its activity and membrane targeting (91, 92). The expression of Bnip3 is often low in normal conditions, but it can be rapidly induced in adverse conditions, in particular, in hypoxia or ischemia (29, 30, 93, 94). Bnip3 can be transcriptionally activated by HIF-1 (29, 30) or FoxO3 (27) but repressed by NF-κB (95) or Rb/E2F (96).
The increased expression of Bnip3 is often accompanied by apoptotic or necrotic cell death, and autophagy, in cell lines subjected to hypoxia (29, 74, 96), or treated with ceramide or arsenic trioxide (97, 98), or in cardiac myocytes subjected to ischemia-reperfusion injury (99, 100). While both cell death and autophagy can be attributed to Bnip3, in most studies, direct evidence that autophagy contributes to cell death is limited, e.g., with tumor cells under hypoxia (74). In contrast, in a cardiac myocyte line under simulated ischemia-reperfusion conditions (99), or in murine embryonic fibroblasts subjected to hypoxia (101), autophagy was found to be important for cell survival. In both cases, it was found that this protective effect is due to the Bnip3-promoted autophagic removal of damaged mitochondria (mitophagy) (99, 101), which is necessary to prevent increased levels of reactive oxygen species (101). Interestingly, a close homologue of Bnip3, Bnip3L/Nix, can also be induced by hypoxia and has been shown to be required for mitophagy during erythrocyte maturation (102, 103).
Could the different roles of Bnip3-mediated autophagy be related to different cell types under different treatments or be related to other factors? The mechanism by which Bnip3 induces autophagy has not been completely elucidated, although one study indicated that Bnip3 could bind to and inactivate Rheb, therefore inactivating mTOR during hypoxia (104). As mTOR is a major factor of autophagy suppression, the negative effect of Bnip 3 on mTOR can be expected to induce autophagy. Bnip3 is thus considered to be required for both hypoxia-induced autophagy and hypoxia-induced mTOR repression. Alternatively, Bnip3 may compete with Bcl-2 for binding to Beclin-1 and thus can cause the derepression of Beclin-1, which in turn promotes autophagy (101) (Fig. 30.1).
It is not completely clear whether cell death and autophagy are mechanistically coupled in the case of Bnip3. It is possible that Bnip3-induced cell death may not be related to autophagy but instead goes through a separate mechanism. Some studies suggested that cell death is related to mitochondria permeabilization by its dimerized TM domain with or without the participation of Bax and Bak (105, 106). It has been postulated that Bnip3’s killing activity may be secondary to prolonged hypoxia, ischemia-reperfusion injury, and anaerobic glycolysis, which results in acidosis (81). Notably, acidosis has been found to stabilize Bnip3 and to increase its association with the mitochondria and its killing ability (107). Furthermore, hypoxia or ischemia does not induce cell death in cardiac myocytes in the absence of acidosis (108). Thus, it is possible that Bnip3 may primarily induce autophagy, as a protective mechanism at the early stage of hypoxia/ischemia, but induce cell death at the later stage when acidosis occurs, which promotes Bnip3 interaction with the mitochondria and membrane permeabilization.
Finally, when considering the cell death’s relationship with autophagy, one may need to be aware that sometimes the same phenomenon may be subjected to different interpretations, as the mechanisms of the action are not always clear, particularly at the beginning. A pan-caspase inhibitor, z-VAD, is often used to suppress caspases and, therefore, caspase-mediated apoptosis. However, this chemical could inhibit other types of proteases, notably, the lysosomal cathepsins (69). The application of z-VAD to several types of cells resulted in the increased accumulation of autophagic markers and necrotic cell death that seemed to be attributable to autophagy (109). However, further investigation indicated that the increased autophagosomes are not due to an increase in the induction, but to a reduction in the degradation of the autophagosome because of the lysosomal inhibition by z-VAD (69). Thus, the increased cell death is related to reduced, rather than increased, autophagy, which was confirmed by a subsequent study indicating that autophagy was prosurvival rather than prodeath following z-VAD treatment.
The Potential Clinical Significance of Modulating Autophagy to Control Cell Death
Understanding the various conditions under which autophagy may be prosurvival or prodeath can have practical benefits in controlling the disease process. This topic is particularly attractive for cancer therapy, where the goal is to eliminate tumor cells by promoting cell death. Both properties of autophagy in regulating cell death have been explored for this purpose. Thus, numerous reports have indicated that suppressing autophagy during treatment with chemotherapeutic agents such as alkylating drugs, proteasome inhibitors, and histone deactylase inhibitors (39, 63, 64) could enhance apoptosis in various types of tumor lines. Conversely, promoting autophagy by the combined use of rapamycin has been found to enhance radiation therapy (73). The future challenge is to define the conditions under which autophagy plays a specific role in cell death, promotion, or inhibition. It is possible to take advantage of this dichotomic characteristic of autophagy for maximal benefits. For example, ER-stress inducers (38) and proteasome inhibitors (Ding and Yin, Molecular Cancer Therapeutics, inpress, 2009) can induce the prosurvival function of autophagy in the tumor cells, but they induce the prodeath function in normal cells. Thus, the suppression of autophagy in vivo may specifically enhance death in the tumor cells, but reduce death in the normal cells under these circumstances.
The prodeath activity of autophagy during ischemic injury in the brain (75) may be suppressed to reduce organ damage. Other than directly targeting the autophagy genes, it may be possible to act on specific targets if the mechanism of autophagic death is known, such as the inactivation of Bcl-xL by the cleaved Atg5 (71). On the other hand, autophagic activity can be enhanced for the removal of misfolded proteins, such as the mutant huntingtin, as seen in Huntington’s disease, and the alpha-1 antitrypsin Z mutant (26, 37, 110). The prosurvival function of autophagy could thus be explored in these conformational diseases.
Conclusion
Autophagy is an evolutionarily conserved physiological process that degrades various cellular contents. Its functions are thus interfaced with cellular survival and cell death. Depending on the context, autophagy activity could contribute to either cell survival or cell death. Only in very limited cases are the mechanisms by which autophagy modulates cell death understood. But both apoptosis and necrosis can be modulated by autophagy, and excessive digestion by autophagy can also lead to cellular atrophy and direct death. Future work should focus on understanding individual cases so that this character of autophagy can be explored for cancer therapy, control of tissue injury, and treatment of conformational diseases.
References
De Duve C, Wattiaux R. Functions of lysosomes. Annu Rev Physiol 1966;28(1):435–92.
Seglen PO, Bohley P. Autophagy and other vacuolar protein degradation mechanisms. Experientia 1992;48(2):158–72.
Ohsumi Y. Molecular dissection of autophagy: Two ubiquitin-like systems. Nat Rev Mol Cell Biol 2001;2(3):211–6.
Levine B, Klionsky DJ. Development by self-digestion: Molecular mechanisms and biological functions of autophagy. Dev Cell 2004;6(4):463–77.
Meijer AJ, Codogno P. Regulation and role of autophagy in mammalian cells. Int J Biochem Cell Biol 2004;36(12):2445–62.
Cuervo AM. Autophagy: In sickness and in health. Trends Cell Biol 2004;14(2):70–7.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ. Autophagy fights disease through cellular self-digestion. Nature 2008;451(7182):1069–75.
Clark SL, Jr. Cellular differentiation in the kidneys of newborn mice studied with the electron microscope. J Cell Biol 1957;3(3):349–62.
Lum JJ, DeBerardinis RJ, Thompson CB. Autophagy in metazoans: Cell survival in the land of plenty. Nat Rev Mol Cell Biol 2005;6(6):439–48.
Mizushima N, Ohsumi Y, Yoshimori T. Autophagosome formation in mammalian cells. Cell Struct Funct 2002;27(6):421–9.
Klionsky DJ, Cregg JM, Dunn WA, Jr., et al. A unified nomenclature for yeast autophagy-related genes. Dev Cell 2003;5(4):539–45.
Tsukada M, Ohsumi Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett 1993;333(1–2):169–74.
Klionsky DJ. Autophagy: From phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol2007;8(11):931–7.
Suzuki K, Ohsumi Y. Molecular machinery of autophagosome formation in yeast, Saccharomyces cerevisiae. FEBS Lett 2007;581(11):2156–61.
Xie Z, Klionsky DJ. Autophagosome formation: Core machinery and adaptations. Nat Cell Biol 2007;9(10):1102–9.
Kabeya Y, Mizushima N, Ueno T, et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000;19(21):5720–8.
Fujita N, Itoh T, Fukuda M, Noda T, Yoshimori T. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell 2008;19(5):2092–100.
Pattingre S, Tassa A, Qu X, et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005;122(6):927–39.
Maria Fimia G, Stoykova A, Romagnoli A, et al. Ambra1 regulates autophagy and development of the nervous system. Nature 2007;447(7148):1121–5.
Maiuri MC, Le Toumelin G, Criollo A, et al. Functional and physical interaction between Bcl-X(L) and a BH3-like domain in Beclin-1. EMBO J 2007;26(10):2527–39.
Oberstein A, Jeffrey PD, Shi Y. Crystal structure of the Bcl-XL-Beclin 1 peptide complex: Beclin 1 is a novel BH3-only protein. J Biol Chem 2007;282(17):13123–32.
Liang C, Feng P, Ku B, et al. Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat Cell Biol 2006;8(7):688–99.
Takahashi Y, Coppola D, Matsushita N, et al. Bif-1 interacts with Beclin 1 through UVRAG and regulates autophagy and tumorigenesis. Nat Cell Biol 2007;9(10):1142–51.
Kamada Y, Funakoshi T, Shintani T, Nagano K, Ohsumi M, Ohsumi Y. Tor-mediated induction of autophagy via an Apg1 protein kinase complex. J Cell Biol 2000;150(6):1507–13.
Kundu M, Lindsten T, Yang CY, et al. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood 2008;112(4):1493–502.
Sarkar S, Ravikumar B, Floto RA, Rubinsztein DC. Rapamycin and mTOR-independent autophagy inducers ameliorate toxicity of polyglutamine-expanded huntingtin and related proteinopathies. Cell Death Differ 2009;16(1):46–56.
Mammucari C, Milan G, Romanello V, et al. FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab 2007;6(6):458–71.
Zhao J, Brault JJ, Schild A, et al. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell Metab 2007;6(6):472–83.
Guo K, Searfoss G, Krolikowski D, et al. Hypoxia induces the expression of the pro-apoptotic gene BNIP3. Cell Death Differ 2001;8(4):367–76.
Bruick RK. Expression of the gene encoding the proapoptotic Nip3 protein is induced by hypoxia. Proc Natl Acad Sci USA 2000;97(16):9082–7.
Polager S, Ofir M, Ginsberg D. E2F1 regulates autophagy and the transcription of autophagy genes. Oncogene 2008;27(35):4860–4.
Crighton D, Wilkinson S, O'Prey J, et al. DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell 2006;126(1):121–34.
Tasdemir E, Chiara Maiuri M, Morselli E, et al. A dual role of p53 in the control of autophagy. Autophagy 2008;4(6).
Yorimitsu T, Klionsky DJ. Eating the endoplasmic reticulum: Quality control by autophagy. Trends Cell Biol 2007;17(6):279–85.
Kouroku Y, Fujita E, Tanida I, et al. ER stress (PERK/eIF2alpha phosphorylation) mediates the polyglutamine-induced LC3 conversion, an essential step for autophagy formation. Cell Death Differ 2007;14(2):230–9.
Ogata M, Hino S-i, Saito A, et al. Autophagy is activated for cell survival after endoplasmic reticulum stress. Mol Cell Biol 2006;26(24):9220–31.
Ding WX, Yin XM. Sorting, recognition and activation of the misfolded protein degradation pathways through macroautophagy and the proteasome. Autophagy 2008;4(2):141–50.
Ding WX, Ni HM, Gao W, et al. Differential effects of endoplasmic reticulum stress-induced autophagy on cell survival. J Biol Chem 2007;282(7):4702–10.
Ding WX, Ni HM, Gao W, et al. Linking of autophagy to ubiquitin-proteasome system is important for the regulation of endoplasmic reticulum stress and cell viability. Am J Pathol 2007;171(2):513–24.
Hoyer-Hansen M, Bastholm L, Szyniarowski P, et al. Control of macroautophagy by calcium, calmodulin-dependent kinase kinase-beta, and Bcl-2. Mol Cell 2007;25(2):193–205.
Sakaki K, Wu J, Kaufman RJ. Protein kinase Ctheta is required for autophagy in response to stress in the endoplasmic reticulum. J Biol Chem 2008;283(22):15370–80.
Qu X, Zou Z, Sun Q, et al. Autophagy gene-dependent clearance of apoptotic cells during embryonic development. Cell 2007;128(5):931–46.
Shintani T, Klionsky DJ. Autophagy in health and disease: A double-edged sword. Science 2004;306(5698):990–5.
Lum JJ, Bauer DE, Kong M, et al. Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell 2005;120(2):237–48.
Kuma A, Hatano M, Matsui M, et al. The role of autophagy during the early neonatal starvation period. Nature 2004;432(7020):1032–6.
Mizushima N. Autophagy: Process and function. Genes Dev 2007;21(22):2861–73.
Levine B. Cell biology: Autophagy and cancer. Nature 2007;446(7137):745–7.
Mathew R, Karantza-Wadsworth V, White E. Role of autophagy in cancer. Nat Rev Cancer 2007;7(12):961–7.
Qu X, Yu J, Bhagat G, et al. Promotion of tumorigenesis by heterozygous disruption of the Beclin 1 autophagy gene. J Clin Invest 2003;112(12):1809–20.
Liang XH, Jackson S, Seaman M, et al. Induction of autophagy and inhibition of tumorigenesis by Beclin 1. Nature 1999;402(6762):672–6.
Zhang Y, Qi H, Taylor R, Xu W, Liu LF, Jin S. The role of autophagy in mitochondria maintenance: Characterization of mitochondrial functions in autophagy-deficient S. cerevisiae strains. Autophagy 2007;3(4):337–46.
Karantza-Wadsworth V, Patel S, Kravchuk O, et al. Autophagy mitigates metabolic stress and genome damage in mammary tumorigenesis. Genes Dev 2007;21(13):1621–35.
Debnath J, Baehrecke EH, Kroemer G. Does autophagy contribute to cell death? Autophagy 2005;1(2):66–74.
Baehrecke EH. Autophagy: Dual roles in life and death? Nat Rev Mol Cell Biol 2005;6(6):505–10.
Kroemer G, El-Deiry WS, Golstein P, et al. Classification of cell death: Recommendations of the nomenclature committee on cell death. Cell Death Differ 2005;12(Suppl 2):1463–7.
Bursch W. The autophagosomal-lysosomal compartment in programmed cell death. Cell Death Differ 2001;8(6):569–81.
Schiaffino S, Mammucari C, Sandri M. The role of autophagy in neonatal tissues: Just a response to amino acid starvation? Autophagy 2008;4(5).
Scott RC, Schuldiner O, Neufeld TP. Role and regulation of starvation-induced autophagy in the Drosophila fat body. Dev Cell 2004;7(2):167–78.
Rusten TE, Lindmo K, Juhasz G, et al. Programmed autophagy in the Drosophila fat body is induced by ecdysone through regulation of the PI3K pathway. Dev Cell 2004;7(2):179–92.
Boya P, Gonzalez-Polo RA, Casares N, et al. Inhibition of macroautophagy triggers apoptosis. Mol Cell Biol 2005;25(3):1025–40.
Noda T, Ohsumi Y. Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast. J Biol Chem 1998;273(7):3963–6.
Jin S, DiPaola RS, Mathew R, White E. Metabolic catastrophe as a means to cancer cell death. J Cell Sci 2007;120(Pt 3):379–83.
Amaravadi RK, Yu D, Lum JJ, et al. Autophagy inhibition enhances therapy-induced apoptosis in a Myc-induced model of lymphoma. J Clin Invest 2007;117(2):326–36.
Carew JS, Nawrocki ST, Kahue CN, et al. Targeting autophagy augments the anticancer activity of the histone deacetylase inhibitor SAHA to overcome Bcr-Abl-mediated drug resistance. Blood 2007;110(1):313–22.
Melendez A, Talloczy Z, Seaman M, Eskelinen EL, Hall DH, Levine B. Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 2003;301(5638):1387–91.
Talloczy Z, Jiang W, Virgin HW, et al. Regulation of starvation- and virus-induced autophagy by the eIF2alpha kinase signaling pathway. Proc Natl Acad Sci USA 2002;99(1):190–5.
Tasdemir E, Maiuri MC, Galluzzi L, et al. Regulation of autophagy by cytoplasmic p53. Nat Cell Biol 2008;10(6):676–87.
Degenhardt K, Mathew R, Beaudoin B, et al. Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 2006;10(1):51–64.
Wu YT, Tan HL, Huang Q, et al. Autophagy plays a protective role during zVAD-induced necrotic cell death. Autophagy 2008;4(4):457–66.
Berry DL, Baehrecke EH. Growth arrest and autophagy are required for salivary gland cell degradation in Drosophila. Cell 2007;131(6):1137–48.
Yousefi S, Perozzo R, Schmid I, et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol 2006;8(10):1124–32.
Shimizu S, Kanaseki T, Mizushima N, et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat Cell Biol 2004;6(12):1221–8.
Moretti L, Attia A, Kim KW, Lu B. Crosstalk between Bak/Bax and mTOR signaling regulates radiation-induced autophagy. Autophagy 2007;3(2):142–4.
Azad MB, Chen Y, Henson ES, et al. Hypoxia induces autophagic cell death in apoptosis-competent cells through a mechanism involving BNIP3. Autophagy 2008;4(2):195–204.
Koike M, Shibata M, Tadakoshi M, et al. Inhibition of autophagy prevents hippocampal pyramidal neuron death after hypoxic-ischemic injury. Am J Pathol 2008;172(2):454–69.
Pyo JO, Jang MH, Kwon YK, et al. Essential roles of Atg5 and FADD in autophagic cell death: Dissection of autophagic cell death into vacuole formation and cell death. J Biol Chem 2005;280(21):20722–9.
Espert L, Denizot M, Grimaldi M, et al. Autophagy is involved in T cell death after binding of HIV-1 envelope proteins to CXCR4. J Clin Invest 2006;116(8):2161–72.
Ullman E, Fan Y, Stawowczyk M, Chen HM, Yue Z, Zong WX. Autophagy promotes necrosis in apoptosis-deficient cells in response to ER stress. Cell Death Differ 2008;15(2):422–5.
Chen Y, McMillan-Ward E, Kong J, Israels SJ, Gibson SB. Oxidative stress induces autophagic cell death independent of apoptosis in transformed and cancer cells. Cell Death Differ 2008;15(1):171–82.
Samara C, Syntichaki P, Tavernarakis N. Autophagy is required for necrotic cell death in Caenorhabditis elegans. Cell Death Differ 2008;15(1):105–12.
Tracy K, Macleod KF. Regulation of mitochondrial integrity, autophagy and cell survival by BNIP3. Autophagy 2007;3(6):616–9.
Wan G, Zhaorigetu S, Liu Z, Kaini R, Jiang Z, Hu CA. Apolipoprotein l1, a novel BH3-only lipid binding protein, induces autophagic cell death. J Biol Chem 2008;283(31):21540–9.
Rashmi R, Pillai SG, Vijayalingam S, Ryerse J, Chinnadurai G. BH3-only protein BIK induces caspase-independent cell death with autophagic features in Bcl-2 null cells. Oncogene 2008;27(10):1366–75.
Yanagisawa H, Miyashita T, Nakano Y, Yamamoto D. HSpin1, a transmembrane protein interacting with Bcl-2/Bcl-xL, induces a caspase-independent autophagic cell death. Cell Death Differ 2003;10(7):798–807.
Levine B, Sinha S, Kroemer G. Bcl-2 family members: Dual regulators of apoptosis and autophagy. Autophagy 2008;4(5):600–6.
Saeki K, Yuo A, Okuma E, et al. Bcl-2 down-regulation causes autophagy in a caspase-independent manner in human leukemic HL60 cells. Cell Death Differ 2000;7(12):1263–9.
Hashimoto D, Ohmuraya M, Hirota M, et al. Involvement of autophagy in trypsinogen activation within the pancreatic acinar cells. J Cell Biol 2008;181(7):1065–72.
Kang C, Avery L. To be or not to be, the level of autophagy is the question: Dual roles of autophagy in the survival response to starvation. Autophagy 2008;4(1):82–4.
Wang Y, Singh R, Massey AC, et al. Loss of macroautophagy promotes or prevents fibroblast apoptosis depending on the death stimulus. J Biol Chem 2008;283(8):4766–77.
Boyd JM, Malstrom S, Subramanian T, et al. Adenovirus E1B 19 kDa and Bcl-2 proteins interact with a common set of cellular proteins. Cell 1994;79(2):341–51.
Chen G, Ray R, Dubik D, et al. The E1B 19 K/Bcl-2-binding protein Nip3 is a dimeric mitochondrial protein that activates apoptosis. J Exp Med 1997;186(12):1975–83.
Ray R, Chen G, Vande Velde C, et al. BNIP3 heterodimerizes with Bcl-2/Bcl-X(L) and induces cell death independent of a Bcl-2 homology 3 (BH3) domain at both mitochondrial and nonmitochondrial sites. J Biol Chem 2000;275(2):1439–48.
Graham RM, Thompson JW, Wei J, Bishopric NH, Webster KA. Regulation of Bnip3 death pathways by calcium, phosphorylation, and hypoxia-reoxygenation. Antioxid Redox Signal 2007;9(9):1309–15.
Lee H, Paik SG. Regulation of BNIP3 in normal and cancer cells. Mol Cells 2006;21(1):1–6.
Baetz D, Regula KM, Ens K, et al. Nuclear factor-kappaB-mediated cell survival involves transcriptional silencing of the mitochondrial death gene BNIP3 in ventricular myocytes. Circulation 2005;112(24):3777–85.
Tracy K, Dibling BC, Spike BT, Knabb JR, Schumacker P, Macleod KF. BNIP3 is an RB/E2F target gene required for hypoxia-induced autophagy. Mol Cell Biol 2007;27(17):6229–42.
Daido S, Kanzawa T, Yamamoto A, Takeuchi H, Kondo Y, Kondo S. Pivotal role of the cell death factor BNIP3 in ceramide-induced autophagic cell death in malignant glioma cells. Cancer Res 2004;64(12):4286–93.
Kanzawa T, Zhang L, Xiao L, Germano IM, Kondo Y, Kondo S. Arsenic trioxide induces autophagic cell death in malignant glioma cells by upregulation of mitochondrial cell death protein BNIP3. Oncogene 2005;24(6):980–91.
Hamacher-Brady A, Brady NR, Logue SE, et al. Response to myocardial ischemia/reperfusion injury involves Bnip3 and autophagy. Cell Death Differ 2007;14(1):146–57.
Diwan A, Krenz M, Syed FM, et al. Inhibition of ischemic cardiomyocyte apoptosis through targeted ablation of Bnip3 restrains postinfarction remodeling in mice. J Clin Invest 2007;117(10):2825–33.
Zhang H, Bosch-Marce M, Shimoda LA, et al. Mitochondrial autophagy is a HIF-1-dependent adaptive metabolic response to hypoxia. J Biol Chem 2008;282(16):10892–903.
Schweers RL, Zhang J, Randall MS, et al. NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc Natl Acad Sci USA 2007;104(49):19500–5.
Sandoval H, Thiagarajan P, Dasgupta SK, et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature 2008;454(7201):232–5.
Li Y, Wang Y, Kim E, et al. Bnip3 mediates the hypoxia-induced inhibition on mammalian target of rapamycin by interacting with Rheb. J Biol Chem 2007;282(49):35803–13.
Bocharov EV, Pustovalova YE, Pavlov KV, et al. Unique dimeric structure of BNip3 transmembrane domain suggests membrane permeabilization as a cell death trigger. J Biol Chem 2007;282(22):16256–66.
Kubli DA, Ycaza JE, Gustafsson AB. Bnip3 mediates mitochondrial dysfunction and cell death through Bax and Bak. Biochem J 2007;405(3):407–15.
Kubasiak LA, Hernandez OM, Bishopric NH, Webster KA. Hypoxia and acidosis activate cardiac myocyte death through the Bcl-2 family protein BNIP3. Proc Natl Acad Sci USA 2002;99(20):12825–30.
Webster KA, Discher DJ, Kaiser S, Hernandez O, Sato B, Bishopric NH. Hypoxia-activated apoptosis of cardiac myocytes requires reoxygenation or a pH shift and is independent of p53. J Clin Invest 1999;104(3):239–52.
Yu L, Alva A, Su H, et al. Regulation of an ATG7-Beclin 1 program of autophagic cell death by caspase-8. Science 2004;304(5676):1500–2.
Perlmutter DH. The role of autophagy in alpha-1-antitrypsin deficiency: A specific cellular response in genetic diseases associated with aggregation-prone proteins. Autophagy 2006;2(4):258–63.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Humana Press, a part of Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Gao, W., Kang, JH., Liao, Y., Li, M., Yin, XM. (2009). Autophagy and Cell Death. In: Dong, Z., Yin, XM. (eds) Essentials of Apoptosis. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-60327-381-7_30
Download citation
DOI: https://doi.org/10.1007/978-1-60327-381-7_30
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-60327-380-0
Online ISBN: 978-1-60327-381-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)