Abstract
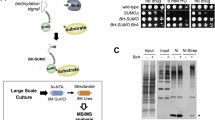
Ni-NTA affinity chromatography under denaturing conditions has proven to be a powerful method for the isolation of SUMO conjugates from total cell extracts, as it minimizes deconjugation and excludes noncovalent interactions. This chapter describes the use of both His-tagged SUMO and a His-tagged target protein for the characterization of the sumoylation process in the budding yeast Saccharomyces cerevisiae. Two well-studied model substrates, the septin Cdc3 and the replication clamp protein PCNA, are used as examples, but the protocol can easily be adapted to other targets and organisms.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Denison, C., Rudner, A. D., Gerber, S. A., Bakalarski, C. E., Moazed, D., and Gygi, S. P. (2005) A proteomic strategy for gaining insights into protein sumoylation in yeast. Mol. Cell. Proteomics 4, 246–254.
Ganesan, A. K., Kho, Y., Kim, S. C., Chen, Y., Zhao, Y., and White, M. A. (2007) Broad spectrum identification of SUMO substrates in melanoma cells. Proteomics 7, 2216–2221.
Hannich, J. T., Lewis, A., Kroetz, M. B., Li, S. J., Heide, H., Emili, A., and Hoch-strasser, M. (2005) Defining the SUMO-modified proteome by multiple approaches in Saccharomyces cerevisiae. J. Biol. Chem. 280, 4102–4110.
Manza, L. L., Codreanu, S. G., Stamer, S. L., Smith, D. L., Wells, K. S., Roberts, R. L., and Liebler, D. C. (2004) Global shifts in protein sumoylation in response to elec-trophile and oxidative stress. Chem. Res. Toxicol. 17, 1706–1715.
Panse, V. G., Hardeland, U., Werner, T., Kuster, B., and Hurt, E. (2004) A proteome-wide approach identifies sumoylated substrate proteins in yeast. J. Biol. Chem. 279, 41346–41351.
Rosas-Acosta, G., Russell, W. K., Deyrieux, A., Russell, D. H., and Wilson, V. G. (2005) A universal strategy for proteomic studies of SUMO and other ubiquitin-like modifiers. Mol. Cell. Proteomics 4, 56–72.
Vertegaal, A. C., Ogg, S. C., Jaffray, E., Rodriguez, M. S., Hay, R. T., Andersen, J. S., Mann, M., and Lamond, A. I. (2004) A proteomic study of SUMO-2 target proteins. J. Biol. Chem. 279, 33791–33798.
Wohlschlegel, J. A., Johnson, E. S., Reed, S. I., and Yates, J. R. 3rd. (2004) Global analysis of protein sumoylation in Saccharomyces cerevisiae. J. Biol. Chem. 279, 45662–45668.
Zhao, Y., Kwon, S. W., Anselmo, A., Kaur, K., and White, M. A. (2004) Broad spectrum identification of cellular small ubiq-uitin-related modifier (SUMO) substrate proteins. J. Biol. Chem. 279, 20999–21002.
Zhou, W., Ryan, J. J., and Zhou, H. (2004) Global analyses of sumoylated proteins in Saccharomyces cerevisiae. Induction of protein sumoylation by cellular stresses. J. Biol. Chem. 279, 32262–32268.
Johnson, E. S. and Blobel, G. (1999) Cell cycle-regulated attachment of the ubiquitin-related protein SUMO to the yeast septins. J. Cell Biol. 147, 981–994.
Johnson, E. S. and Gupta, A. A. (2001) An E3-like factor that promotes SUMO conjugation to the yeast septins. Cell 106, 735–744.
Takahashi, Y., Kahyo, T., Toh-E, A., Yasuda, H., and Kikuchi, Y. (2001) Yeast Ull1/Siz1 is a novel SUMO1/Smt3 ligase for septin components and functions as an adaptor between conjugating enzyme and substrates. J. Biol. Chem. 276, 48973–48977.
Jónsson, Z. O. and Hübscher, U. (1997) Proliferating cell nuclear antigen: more than a clamp for DNA polymerases. BioEssays 19, 967–975.
Hoege, C., Pfander, B., Moldovan, G. L., Pyrowolakis, G., and Jentsch, S. (2002) RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141.
Stelter, P. and Ulrich, H. D. (2003) Control of spontaneous and damage-induced muta-genesis by SUMO and ubiquitin conjugation. Nature 425, 188–191.
Papouli, E., Chen, S., Davies, A. A., Hut-tner, D., Krejci, L., Sung, P., and Ulrich, H. D. (2005) Crosstalk between SUMO and ubiquitin on PCNA is mediated by recruitment of the helicase Srs2p. Mol. Cell 19, 123–133.
Pfander, B., Moldovan, G. L., Sacher, M., Hoege, C., and Jentsch, S. (2005) SUMO-modified PCNA recruits Srs2 to prevent recombination during S phase. Nature 436, 428–433.
Robert, T., Dervins, D., Fabre, F., and Gan-gloff, S. (2006) Mrc1 and Srs2 are major actors in the regulation of spontaneous crossover. EMBO J. 25, 2837–2846.
Windecker, H. and Ulrich, H. D. (2008) Architecture and assembly of poly-SUMO chains on PCNA in Saccharomyces cerevisiae. J. Mol. Biol. 376, 221–231.
Guthrie, C. and Fink, G. R. (1991) Guide to Yeast Genetics and Molecular Biology. Academic Press, San Diego, CA.
Gietz, R. D. and Sugino, A. (1988) New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74, 527–534.
Knop, M., Siegers, K., Pereira, G., Zach-ariae, W., Winsor, B., Nasmyth, K., and Schiebel, E. (1999) Epitope tagging of yeast genes using a PCR-based strategy: more tags and improved practical routines. Yeast 15, 963–972.
Ayyagari, R., Impellizzeri, K. J., Yoder, B. L., Gary, S. L., and Burgers, P. M. (1995) A mutational analysis of the yeast proliferating cell nuclear antigen indicates distinct roles in DNA replication and DNA repair. Mol. Cell. Biol. 15, 4420–4429.
Longtine, M. S., McKenzie, A., 3rd, Demarini, D. J., Shah, N. G., Wach, A., Brachat, A., Philippsen, P., and Pringle, J. R. (1998) Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961.
Sambrook, J. and Russell, D. W. (2001) Molecular Cloning: A Laboratory Manual. 3rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Acknowledgments
The authors would like to thank E. Johnson for providing the strain CDC3 HA, P. Burgers for plasmid pBL243, and M. Knop for plasmid pYM3. Work in this lab is supported by Cancer Research UK.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Ulrich, H.D., Davies, A.A. (2009). In Vivo Detection and Characterization of Sumoylation Targets in Saccharomyces cerevisiae . In: Ulrich, H.D. (eds) SUMO Protocols. METHODS IN MOLECULAR BIOLOGY™, vol 497. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-59745-566-4_6
Download citation
DOI: https://doi.org/10.1007/978-1-59745-566-4_6
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-934115-80-0
Online ISBN: 978-1-59745-566-4
eBook Packages: Springer Protocols