Abstract
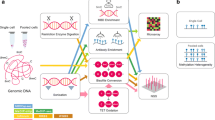
DNA methylation is an epigenetic modification that plays a crucial role in the control of gene expression and chromosome structure in plants and mammalian cells. Multiple types of DNA fingerprinting techniques have been developed and applied to investigate DNA methylation profiles in different experimental settings. One of these techniques, the amplification of intermethylated sites (AIMS) is a simple approach appropriate for genome-wide estimates of DNA methylation and the discovery of specific methylated sequences. AIMS is based on the differential enzymatic digestion of genomic DNA with methylation-sensitive and methylation-insensitive isoschizomers followed by restrained PCR amplification of methylated sequences. This method is appropriate to compare large series of samples and the simultaneous identification of hypo- and hypermethylation events. Applications of AIMS include the study of DNA methylation changes in cancer and aging, and the discovery of DNA methylation in a social insect.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Jones, P. A., Baylin, S. B. (2002) The fundamental role of epigenetic events in cancer. Nat Rev Genet 3, 415–428.
Jaenisch, R., Bird, A. (2003) Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 33 Suppl, 245–254.
Feinberg, A. P., Tycko, B. (2004) The history of cancer epigenetics. Nat Rev Cancer 4, 143–153.
Cottrell, S. E. (2004) Molecular diagnostic applications of DNA methylation technology. Clin Biochem 37, 595–604.
Fazzari, M. J., Greally, J. M. (2004) Epigenomics: beyond CpG islands. Nat Rev Genet 5, 446–455.
Esteller, M. (2007) Cancer epigenomics: DNA methylomes and histone-modification maps. Nat Rev Genet 6, 286–298.
Frigola, J., Ribas, M., Risques, R. A., et al. (2002) Methylome profiling of cancer cells by amplification of inter- methylated sites (AIMS). Nucleic Acids Res 30, e28.
Frigola, J., Mu noz, M., Clark, S. J., et al. (2005) Hypermethylation of the prostacyclin synthase (PTGIS) promoter is a frequent event in colorectal cancer and associated with aneuploidy. Oncogene 24, 7320–7326.
Frigola J, Song, J., Stirzaker, C., et al. (2006) Epigenetic remodeling in colorectal cancer results in coordinate gene suppression across an entire chromosome band. Nat Genet 38, 540–549.
Paz, M. F., Wei, S., Cigudosa, J. C., et al. (2003) Genetic unmasking of epigenetically silenced tumor suppressor genes in colon cancer cells deficient in DNA methyltransferases. Hum Mol Genet 12, 2209–2219.
Sadikovic, B., Haines, T. R., Butcher, D. T., et al. (2004) Chemically induced DNA hypomethylation in breast carcinoma cells detected by the amplification of intermethylated sites. Breast Cancer Res 6, R329-337.
Frigola, J., Sole, X., Paz, M. F., et al. (2005) Differential DNA hypermethylation and hypomethylation signatures in colorectal cancer. Hum Mol Genet 14, 319–326.
Rodriguez, J., Frigola, J., Vendrell, E., et al. (2006) Chromosomal instability correlates with genome-wide DNA demethylation in human primary colorectal cancers. Cancer Res 66, 8462–8468.
Fraga, M. F., Ballestar, E., Paz, M. F., et al. (2005) Epigenetic differences arise during the lifetime of monozygotic twins.Proc Natl Acad Sci USA 102, 10604–10609.
Honeybee Genome Sequence Consortium. (2006) Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949.
Wang, Y., Jorda, M., Jones, P. L., et al. (2006) Functional CpG methylation system in a social insect. Science 314, 645–647.
Perucho, M., Welsh, J., Peinado, M. A., et al. (1995) Fingerprinting of DNA and RNA by arbitrarily primed polymerase chain reaction: applications in cancer research. Methods Enzymol 254, 275–290.
Arribas, R., Tortola, S., Welsh, J., et al. (1997) Analysis of tumor-specific genetic alterations by Arbitrarily Primed PCR, in (Micheli, M. R., and Bova, R., eds.), Fingerprinting Methods Based on Arbitrarily Primed PCR, pp. 177–185, Heidelberg, Springer, Germany.
Acknowledgements
The research was supported by the Spanish Ministry of Education and Science (SAF2006/351 and Consolider-Ingenio 2010 CSD2006-49).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Jordà, M., Rodríguez, J., Frigola, J., Peinado, M.A. (2009). Analysis of DNA Methylation by Amplification of Intermethylated Sites (AIMS). In: Tost, J. (eds) DNA Methylation. Methods in Molecular Biology, vol 507. Humana Press. https://doi.org/10.1007/978-1-59745-522-0_9
Download citation
DOI: https://doi.org/10.1007/978-1-59745-522-0_9
Publisher Name: Humana Press
Print ISBN: 978-1-934115-61-9
Online ISBN: 978-1-59745-522-0
eBook Packages: Springer Protocols