Summary
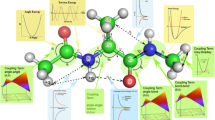
In the context of molecular dynamics simulations of proteins, the term “force field” refers to the combination of a mathematical formula and associated parameters that are used to describe the energy of the protein as a function of its atomic coordinates. In this review, we describe the functional forms and parameterization protocols of the widely used biomolecular force fields Amber, CHARMM, GROMOS, and OPLS-AA. We also summarize the ability of various readily available noncommercial molecular dynamics packages to perform simulations using these force fields, as well as to use modern methods for the generation of constant-temperature, constant-pressure ensembles and to treat long-range interactions. Finally, we finish with a discussion of the ability of these force fields to support the modeling of proteins in conjunction with nucleic acids, lipids, carbohydrates, and/or small molecules.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
1. Cornell, W. D., Cieplak, P., Bayly, C. I., Gould, I. R., Merz Jr., K. M., Ferguson, D. M., Spellmeyer, D. C., Fox, T., Caldwell, J. W., and Kollman, P. A. (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 117, 5179–5197.
2. MacKerell, A. D., Jr., Bashford, D., Bellott, M., Dunbrack, R. L., Evanseck, J. D., Field, M. J., Fischer, S., Gao, J., Guo, H., Ha, S., Joseph-McCarthy, D., Kuchnir, L., Kuczera, K., Lau, F. T. K., Mattos, C., Michnick, S., Ngo, T., Nguyen, D. T., Prodhom, B., Reiher, W. E., Roux, B., Schlenkrich, M., Smith, J. C., Stote, R., Straub, J., Watanabe, M., Wiórkiewicz-Kuczera, J., Yin, D., and Karplus, M. (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616.
3. Oostenbrink, C., Villa, A., Mark, A. E., and Van Gunsteren, W. F. (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: The GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 25, 1656–1676.
4. Jorgensen, W. L., Maxwell, D. S., and Tirado-Rives, J. (1996) Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 118, 11225–11236.
5. Kitson, D. H. and Hagler, A. T. (1988) Theoretical studies of the structure and molecular dynamics of a peptide crystal. Biochemistry 27, 5246–5257.
6. Momany, F. A., Carruthers, L. M., McGuire, R. F., and Scheraga, H. A. (1974) Intermolecular potentials from crystal data. III. Determination of empirical potentials and application to the packing configurations and lattice energies in crystals of hydrocarbons, carboxylic acids, amines, and amides. J. Phys. Chem. 78, 1595–1620.
7. Momany, F. A., McGuire, R. F., Burgess, A. W., and Scheraga, H. A. (1975) Energy parameters in polypeptides. VII. Geometric parameters, partial atomic charges, nonbonded interactions, hydrogen bond interactions, and intrinsic torsional potentials for the naturally occurring amino acids. J. Phys. Chem 79, 2361–2381.
8. Némethy, G., Pottle, M. S., and Scheraga, H. A. (1983) Energy parameters in polypeptides. 9. Updating of geometrical parameters, nonbonded interactions, and hydrogen bond interactions for the naturally occurring amino acids. J. Phys. Chem. 87, 1883–1887.
9. Némethy, G., Gibson, K. D., Palmer, K. A., Yoon, C. N., Paterlini, G., Zagari, A., Rumsey, S., and Scheraga, H. A. (1992) Energy parameters in polypeptides. 10. Improved geometrical parameters and nonbonded interactions for use in the ECEPP/3 algorithm, with application to proline-containing peptides. J. Phys. Chem. 96, 6472–6484.
10. Levitt, M. (1983) Molecular dynamics of native protein: I. Computer simulation of trajectories. J. Mol. Biol. 168, 595–620.
11. Levitt, M., Hirshberg, M., Sharon, R., and Daggett, V. (1995) Potential energy function and parameters for simulations of the molecular dynamics of proteins and nucleic acids in solution. Comput. Phys. Commun. 91, 215–231.
12. Allinger, N. L., Chen, K. S., and Lii, J. H. (1996) An improved force field (MM4) for saturated hydrocarbons. J. Comput. Chem. 17, 642–668.
13. Nevins, N., Chen, K. S., and Allinger, N. L. (1996) Molecular mechanics (MM4) calculations on alkenes. J. Comput. Chem. 17, 669–694.
14. Nevins, N., Lii, J. H., and Allinger, N. L. (1996) Molecular mechanics (MM4) calculations on conjugated hydrocarbons. J. Comput. Chem. 17, 695–729.
15. Nevins, N. and Allinger, N. L. (1996) Molecular mechanics (MM4) vibrational frequency calculations for alkenes and conjugated hydrocarbons. J. Comput. Chem. 17, 730–746.
16. Allinger, N. L., Chen, K. S., Katzenellenbogen, J. A., Wilson, S. R., and Anstead, G. M. (1996) Hyperconjugative effects on carbon-carbon bond lengths in molecular mechanics (MM4). J. Comput. Chem. 17, 747–755.
17. Langley, C. H. and Allinger, N. L. (2002) Molecular mechanics (MM4) calculations on amides. J. Phys. Chem. A 106, 5638–5652.
18. Halgren, T. A. (1996) Merck molecular force field. 1. Basis, form, scope, parameterization, and performance of MMFF94. J. Comput. Chem. 17, 490–519.
19. Halgren, T. A. (1996) Merck molecular force field. 2. MMFF94 van der Waals and electrostatic parameters for intermolecular interactions. J. Comput. Chem. 17, 520–552.
20. Halgren, T. A. (1996) Merck molecular force field. 3. Molecular geometries and vibrational frequencies for MMFF94. J. Comput. Chem. 17, 553–586.
21. Halgren, T. A. and Nachbar, R. B. (1996) Merck molecular force field. 4. Conformational energies and geometries for MMFF94. J. Comput. Chem. 17, 587–615.
22. Halgren, T. A. (1996) Merck molecular force field. 5. Extension of MMFF94 using experimental data, additional computational data, and empirical rules. J. Comput. Chem. 17, 616–641.
23. Halgren, T. A. (1999) MMFF VI. MMFF94s option for energy minimization studies. J. Comput. Chem. 20, 720–729.
24. Halgren, T. A. (1999) MMFF VII. Characterization of MMFF94, MMFF94s, and other widely available force fields for conformational energies and for intermolecular-interaction energies and geometries. J. Comput. Chem. 20, 730–748.
25. Rappé, A. K., Casewit, C. J., Colwell, K. S., Goddard, W. A., and Skiff, W. M. (1992) UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J. Am. Chem. Soc. 114, 10024–10035.
26. Ponder, J. W. and Case, D. A. (2003) Force fields for protein simulations, in Protein Simulations, Vol. 66, Academic Press, San Diego, pp. 27–85.
27. MacKerell, A. D., Jr. (2004) Empirical force fields for biological macromolecules: Overview and issues. J. Comput. Chem. 25, 1584–1604.
28. Daggett, V. (2006) Protein folding-simulation. Chem. Rev. 106, 1898–1916.
29. Oldziej, S., Czaplewski, C., Liwo, A., Chinchio, M., Nanias, M., Vila, J. A., Khalili, M., Arnautova, Y. A., Jagielska, A., Makowski, M., Schafroth, H. D., Kazmierkiewicz, R., Ripoll, D. R., Pillardy, J., Saunders, J. A., Kang, Y. K., Gibson, K. D., and Scheraga, H. A. (2005) Physics-based protein-structure prediction using a hierarchical protocol based on the UNRES force field: Assessment in two blind tests. Proc. Natl. Acad. Sci. U. S. A. 102, 7547–7552.
30. Allen, M. P. and Tildesley, D. J. (1987) Computer simulation of liquids, Oxford University Press, Oxford.
31. Matsuoka, O., Clementi, E., and Yoshimine, M. (1976) CI study of the water dimer potential surface. J. Chem. Phys. 64, 1351–1361.
32. Owicki, J. C. and Scheraga, H. A. (1977) Monte Carlo calculations on the isothermal-isobaric ensemble. 1. Liquid water. J. Am. Chem. Soc. 99, 7403–7412.
Reiher, W. E. (1985), Theoretical studies of hydrogen bonding, Ph.D. thesis, Harvard University.
34. Morozov, A. V., Kortemme, T., Tsemekhman, K., and Baker, D. (2004) Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations. Proc. Natl. Acad. Sci. U. S. A. 101, 6946–6951.
35. Weiner, S. J., Kollman, P. A., Case, D. A., Singh, U. C., Ghio, C., Alagona, G., Profeta, S., and Weiner, P. A. (1984) A new force field for molecular mechanical simulation of nucleic acids and proteins. J. Am. Chem. Soc. 106, 765–784.
36. Mahoney, M. W. and Jorgensen, W. L. (2000) A five-site model for liquid water and the reproduction of the density anomaly by rigid, nonpolarizable potential functions. J. Chem. Phys. 112, 8910–8922.
37. Mahoney, M. W. and Jorgensen, W. L. (2001) Diffusion constant of the TIP5P model of liquid water. J. Chem. Phys. 114, 363–366.
38. Jorgensen, W. L. and Tirado-Rives, J. (2005) Potential energy functions for atomic-level simulations of water and organic and biomolecular systems. Proc. Natl. Acad. Sci. U. S. A. 102, 6665–6670.
39. Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W., and Klein, M. L. (1983) Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935.
40. Berendsen, H. J. C., Grigera, J. R., and Straatsma, T. P. (1987) The missing term in effective pair potentials. J. Phys. Chem. 91, 6269–6271.
41. Cornell, W. D., Cieplak, P., Bayly, C. E., and Kollman, P. A. (1993) Application of RESP charges to calculate conformational energies, hydrogen bond energies, and free energies of solvation. J. Amer.Chem.Soc. 115, 9620–9631.
42. MacKerell, A. D., Jr., Feig, M., and Brooks, C. L., III, (2004) Improved treatment of the protein backbone in empirical force fields. J. Am. Chem. Soc. 126, 698–699.
43. Hu, H., Elstner, M., and Hermans, J. (2003) Comparison of a QM/MM force field and molecular mechanics force fields in simulations of alanine and glycine “dipeptides” (Ace-Ala-Nme and Ace-Gly-Nme) in water in relation to the problem of modeling the unfolded peptide backbone in solution. Proteins 50, 451–463.
44. Wang, J. M., Cieplak, P., and Kollman, P. A. (2000) How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 21, 1049–1074.
45. Okur, A., Strockbine, B., Hornak, V., and Simmerling, C. (2003) Using PC clusters to evaluate the transferability of molecular mechanics force fields for proteins. J. Comput. Chem. 24, 21–31.
46. Kaminski, G. A., Friesner, R. A., Tirado-Rives, J., and Jorgensen, W. L. (2001) Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J. Phys. Chem. B 105, 6474–6487.
47. Shirley, W. A. and Brooks, C. L., III (1997) Curious structure in “canonical” alanine based peptides. Proteins 28, 59–71.
48. Feig, M., MacKerell, A. D., Jr., and Brooks, C. L., III (2003) Force field influence on the observation of pi-helical protein structures in molecular dynamics simulations. J. Phys. Chem. B 107, 2831–2836.
49. MacKerell, A. D., Jr., Feig, M., and Brooks, C. L., III (2004) Extending the treatment of backbone energetics in protein force fields: Limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415.
50. Jacob, E. J., Thompson, H. B., and Bartell, L. S. (1967) Influence of nonbonded interactions on molecular geometry and energy: Calculations for hydrocarbons based on Urey-Bradley field. J. Chem. Phys. 47, 3736–3753.
51. Lifson, S. and Warshel, A. (1968) Consistent force field for calculations of conformations, vibrational spectra, and enthalpies of cycloalkane and n-alkane molecules. J. Chem. Phys. 49, 5116–5129.
52. Allinger, N. L., Tribble, M. T., Miller, M. A., and Wertz, D. H. (1971) Conformational analysis. LXIX. An improved force field for the calculation of the structures and energies of hydrocarbons. J. Am. Chem. Soc. 93, 1637–1648.
53. Jorgensen, W. L. and Tirado-Rives, J. (1988) The OPLS potential function for proteins. Energy minimizations for crystals of cyclic peptides and crambin. J. Amer. Chem. Soc. 110, 1657–1666.
54. Bayly, C. I., Cieplak, P., Cornell, W. D., and Kollman, P. A. (1993) A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: The RESP model. J. Phys. Chem. 97, 10269–10280.
55. Fox, T. and Kollman, P. A. (1998) Application of the RESP methodology in the parametrization of organic solvents. J. Phys. Chem. B 102, 8070–8079.
56. Cieplak, P., Cornell, W. D., Bayly, C. I., and Kollman, P. K. (1995) Application of the multimolecule and multiconformational RESP methodology to biopolymers: Charge derivation for DNA, RNA, and proteins. J. Comput. Chem. 16, 1357–1377.
57. Neria, E., Fischer, S., and Karplus, M. (1996) Simulation of activation free energies in molecular systems. J. Chem. Phys. 105, 1902–1919.
58. Oostenbrink, C., Soares, T. A., van der Vegt, N. F. A., and van Gunsteren, W. F. (2005) Validation of the 53A6 GROMOS force field. Eur. Biophys. J. Biophys. Lett. 34, 273–284.
59. Shirts, M. R., Pitera, J. W., Swope, W. C., and Pande, V. S. (2003) Extremely precise free energy calculations of amino acid side chain analogs: Comparison of common molecular mechanics force fields for proteins. 119, 5740–5761.
60. Deng, Y. and Roux, B. (2004) Hydration of amino acid side chains: Non-polar and electrostatic contributions calculated from staged molecular dynamics free energy simulations with explicit water molecules. J. Phys. Chem. B 108, 16567–16576.
61. Metropolis, N., Rosenbluth, A. W., Rosenbluth, M. N., Teller, A. H., and Teller, E. (1953) Equation of state calculations by fast computing machines. J. Chem. Phys. 21, 1087–1092.
62. Wood, W. W. (1968) Monte Carlo calculations for hard disks in the isothermal-isobaric ensemble. J. Chem. Phys. 48, 415–434.
63. McDonald, I. R. (1969) Monte Carlo calculations for one- and two-component fluids in the isothermal-isobaric ensemble. Chem. Phys. Lett. 3, 241–243.
64. Maurits, N. M., Zvelindovsky, A. V., Sevink, G. J. A., van Vlimmeren, B. A. C., and Fraaije, J. (1998) Hydrodynamic effects in three-dimensional microphase separation of block copolymers: Dynamic mean-field density functional approach. J. Chem. Phys. 108, 9150–9154.
65. Groot, R. D., Madden, T. J., and Tildesley, D. J. (1999) On the role of hydrodynamic interactions in block copolymer microphase separation. J. Chem. Phys. 110, 9739–9749.
66. Shelley, J. C., Shelley, M. Y., Reeder, R. C., Bandyopadhyay, S., Moore, P. B., and Klein, M. L. (2001) Simulations of phospholipids using a coarse grain model. J. Phys. Chem. B 105, 9785–9792.
67. Andersen, H. C. (1980) Molecular dynamics simulations at constant pressure and/or temperature. J. Chem. Phys. 72, 2384–2393.
68. Parrinello, M. and Rahman, A. (1981) Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 52, 7182–7190.
69. Nosé, S. (1984) A molecular dynamics method for simulations in the canonical ensemble. Mol. Phys. 52, 255–268.
70. Hoover, W. G. (1985) Canonical dynamics: Equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697.
71. Feller, S. E., Zhang, Y. H., Pastor, R. W., and Brooks, B. R. (1995) Constant pressure molecular dynamics simulation: The Langevin piston method. J. Chem. Phys. 103, 4613–4621.
72. Tuckerman, M. E. and Martyna, G. J. (2000) Understanding modern molecular dynamics: Techniques and applications. J. Phys. Chem. B 104, 159–178.
73. Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., Di Nola, A., and Haak, J. R. (1984) Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690.
74. Steinbach, P. J. and Brooks, B. R. (1994) New spherical-cutoff methods for long-range forces in macromolecular simulation. J. Comput. Chem. 15, 667–683.
75. Ewald, P. P. (1921) Die Berechnung optischer und elektrostatischer Gitterpotentiale. Ann. Phys. 64, 253–287.
76. Hockney, R. W. and Eastwood, J. W. (1981) Computer simulations using particles, McGraw Hill, New York.
77. Darden, T., York, D., and Pedersen, L. (1993) Particle mesh Ewald: An N•log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092.
78. Essmann, U., Perera, L., Berkowitz, M. L., Darden, T. A., Lee, H., and Pedersen, L. G. (1995) A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593.
79. Darden, T., Toukmaji, A., and Pedersen, L. (1997) Long-range electrostatic effects in biomolecular systems. J. Chem. Phys. 94, 1346–1364.
80. Lagüe, P., Pastor, R. W., and Brooks, B. R. (2004) Pressure-based long-range correction for Lennard-Jones interactions in molecular dynamics simulations: Application to alkanes and interfaces. J. Phys. Chem. B 108, 363–368
81. Brooks, C. L., III, and Karplus, M. (1983) Deformable stochastic boundaries in molecular dynamics. J. Chem. Phys. 79, 6312–6325.
82. Brünger, A. T., Brooks, C. L., III, and Karplus, M. (1984) Stochastic boundary conditions for molecular dynamics simulations of ST2 water. Chem. Phys. Lett. 105, 495.
83. Im, W., Bernéche, S., and Roux, B. (2001) Generalized solvent boundary potential for computer simulations. J. Chem. Phys. 114, 2924–2937.
84. Still, W. C., Tempczyk, A., Hawley, R. C., and Hendrickson, T. (1990) Semianalytical treatment of solvation for molecular mechanics and dynamics. J. Am. Chem. Soc. 112, 6127–6129.
85. Hawkins, G. D., Cramer, C. J., and Truhlar, D. G. (1995) Pairwise solute descreening of solute charges from a dielectric medium. Chem. Phys. Lett. 246, 122–129.
86. Qui, D., Shenkin, P. S., Hollinger, F. P., and Still, W. C. (1997) The GB/SA continuum model for solvation. A fast analytical method for the calculation of approximate Born radii. J. Phys. Chem. A 101, 3005–3014.
87. Ghosh, A., Rapp, C. S., and Friesner, R. A. (1998) Generalized born model based on a surface integral formulation. J. Phys. Chem. B 102, 10983–10990.
88. Dominy, B. N. and Brooks, C. L., III (1999) Development of a generalized born model parametrization for proteins and nucleic acids. J. Phys. Chem. B 103, 3765–3773.
89. Onufriev, A., Bashford, D., and Case, D. A. (2000) Modification of the generalized Born model suitable for macromolecules. J. Phys. Chem. B 104, 3712–3720.
90. Lee, M. S., Salsbury, F. R., and Brooks, C. L., III (2002) Novel generalized Born methods. J. Chem. Phys. 116, 10606–10614.
91. Im, W. P., Lee, M. S., and Brooks, C. L., III (2003) Generalized born model with a simple smoothing function. J. Comput. Chem. 24, 1691–1702.
92. Guvench, O., Shenkin, P., Kolossvary, I., and Still, W. C. (2002) Application of the frozen atom approximation to the GB/SA continuum model for solvation free energy. J. Comput. Chem. 23, 214–221.
93. Im, W., Feig, M., and Brooks, C. L., III (2003) An implicit membrane generalized born theory for the study of structure, stability, and interactions of membrane proteins. Biophys. J. 85, 2900–2918.
Tanizaki, S. and Feig, M. (2005) A generalized Born formalism for heterogeneous dielectric environments: Application to the implicit modeling of biological membranes. J. Chem. Phys. 122.
95. Gallicchio, E., Zhang, L. Y., and Levy, R. M. (2002) The SGB/NP hydration free energy model based on the surface generalized born solvent reaction field and novel nonpolar hydration free energy estimators. J. Comput. Chem. 23, 517–529.
96. Gallicchio, E. and Levy, R. M. (2004) AGBNP: An analytic implicit solvent model suitable for molecular dynamics simulations and high-resolution modeling. J. Comput. Chem. 25, 479–499.
97. Weiner, P. K. and Kollman, P. A. (1981) AMBER: Assisted model building with energy refinement. A general program for modeling molecules and their interactions. J. Comput. Chem. 2, 287–303.
98. Pearlman, D. A., Case, D. A., Caldwell, J. W., Ross, W. S., Cheatham, T. E., Debolt, S., Ferguson, D., Seibel, G., and Kollman, P. (1995) AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comput. Phys. Commun. 91, 1–41.
99. Case, D. A., Cheatham, T. E., Darden, T., Gohlke, H., Luo, R., Merz, K. M., Onufriev, A., Simmerling, C., Wang, B., and Woods, R. J. (2005) The Amber biomolecular simulation programs. J. Comput. Chem. 26, 1668–1688.
100. Brooks, B. R., Bruccoleri, R. E., Olafson, B. D., States, D. J., Swaminathan, S., and Karplus, M. (1983) CHARMM: A program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 4, 187–217.
101. Scott, W. R. P., Hünenberger, P. H., Tironi, I. G., Mark, A. E., Billeter, S. R., Fennen, J., Torda, A. E., Huber, T., Krüger, P., and van Gunsteren, W. F. (1999) The GROMOS biomolecular simulation program package. J. Phys. Chem. A 103, 3596–3607.
102. Christen, T., Hünenberger, P. H., Bakowies, D., Baron, R., Burgi, R., Geerke, D. P., Heinz, T. N., Kastenholz, M. A., Krautler, V., Oostenbrink, C., Peter, C., Trzesniak, D., and Van Gunsteren, W. F. (2005) The GROMOS software for biomolecular simulation: GROMOS05. J. Comput. Chem. 26, 1719–1751.
103. Jorgensen, W. L. (1998) BOSS — Biochemical and organic simulation system, in The Encyclopedia of Computational Chemistry (Schleyer, P. v. R., ed.), Vol. 5, John Wiley and Sons, Ltd., Athens, GA.
104. Jorgensen, W. L. and Tirado-Rives, J. (2005) Molecular modeling of organic and biomolecular systems using BOSS and MCPRO. J. Comput. Chem. 26, 1689–1700.
105. Berendsen, H. J. C., van der Spoel, D., and van Drunen, R. (1995) GROMACS—A message-passing parallel molecular-dynamics implementation. Comput. Phys. Commun. 91, 43–56.
106. Lindahl, E., Hess, B., and van der Spoel, D. (2001) GROMACS 3.0: A package for molecular simulation and trajectory analysis. J. Mol. Model. 7, 306–317.
107. van der Spoel, D., Lindahl, E., Hess, B., Groenhof, G., Mark, A. E., and Berendsen, H. J. C. (2005) GROMACS: Fast, flexible, and free. J. Comput. Chem. 26, 1701–1718.
108. Nelson, M., Humphrey, W., Kufrin, R., Gursoy, A., Dalke, A., Kale, L., Skeel, R., and Schulten, K. (1995) MDScope—A visual computing environment for structural biology. Comput. Phys. Commun. 91, 111–133.
109. Nelson, M. T., Humphrey, W., Gursoy, A., Dalke, A., Kale, L. V., Skeel, R. D., and Schulten, K. (1996) NAMD: A parallel, object oriented molecular dynamics program. Int. J. Supercomput. Appl. High Perform. Comput. 10, 251–268.
110. Kale, L., Skeel, R., Bhandarkar, M., Brunner, R., Gursoy, A., Krawetz, N., Phillips, J., Shinozaki, A., Varadarajan, K., and Schulten, K. (1999) NAMD2: Greater scalability for parallel molecular dynamics. J. Comput. Phys. 151, 283–312.
111. Phillips, J. C., Braun, R., Wang, W., Gumbart, J., Tajkhorshid, E., Villa, E., Chipot, C., Skeel, R. D., Kale, L., and Schulten, K. (2005) Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802.
112. Banks, J. L., Beard, H. S., Cao, Y. X., Cho, A. E., Damm, W., Farid, R., Felts, A. K., Halgren, T. A., Mainz, D. T., Maple, J. R., Murphy, R., Philipp, D. M., Repasky, M. P., Zhang, L. Y., Berne, B. J., Friesner, R. A., Gallicchio, E., and Levy, R. M. (2005) Integrated modeling program, applied chemical theory (IMPACT). J. Comput. Chem. 26, 1752–1780.
113. Karplus, M., and McCammon, J. A. (2002) Molecular dynamics simulations of biomolecules. Nat. Struct. Biol. 9, 646–652.
114. MacKerell, A. D., Jr., Wiórkiewicz-Kuczera, J., and Karplus, M. (1995) An all-atom empirical energy function for the simulation of nucleic acids. J. Am. Chem. Soc. 117, 11946–11975.
115. Ponomarev, S. Y., Thayer, K. M., and Beveridge, D. L. (2004) Ion motions in molecular dynamics simulations on DNA. Proc. Natl. Acad. Sci. U. S. A. 101, 14771–14775.
116. Zhang, Q. and Schlick, T. (2006) Stereochemistry and position-dependent effects of carcinogens on TATA/TBP binding. Biophys. J. 90, 1865–1877.
117. Spackova, N. and Sponer, J. (2006) Molecular dynamics simulations of sarcin-ricin rRNA motif. Nucleic Acids Res. 34, 697–708.
118. Trobro, S. and Aqvist, J. (2006) Analysis of predictions for the catalytic mechanism of ribosomal peptidyl transfer. Biochemistry 45, 7049–7056.
119. Pastor, N. (2005) The B- to A-DNA transition and the reorganization of solvent at the DNA surface. Biophys. J. 88, 3262–3275.
120. Huang, N., Banavali, N. K., and MacKerell, A. D., Jr. (2003) Protein-facilitated base flipping in DNA by cytosine-5-methyltransferase. Proc. Natl. Acad. Sci. U. S. A. 100, 68–73.
121. Soares, T. A., Hünenberger, P. H., Kastenholz, M. A., Krautler, V., Lenz, T., Lins, R. D., Oostenbrink, C., and Van Gunsteren, W. F. (2005) An improved nucleic acid parameter set for the GROMOS force field. J. Comput. Chem. 26, 725–737.
122. Schlenkrich, M., Brinkman, J., MacKerell, A. D., Jr., and Karplus, M. (1996) An empirical potential energy function for phospholipids: Criteria for parameter optimization and applications, in Membrane Structure and Dynamics (Merz, K. M., and Roux, B., eds.), Birkhauser, Boston, pp. 31–81.
123. Feller, S. E., Yin, D. X., Pastor, R. W., and MacKerell, A. D., Jr. (1997) Molecular dynamics simulation of unsaturated lipid bilayers at low hydration: Parameterization and comparison with diffraction studies. Biophys. J. 73, 2269–2279.
124. Yin, D. X. and MacKerell, A. D., Jr. (1998) Combined ab initio empirical approach for optimization of Lennard-Jones parameters. J. Comput. Chem. 19, 334–348.
125. Feller, S. E. and MacKerell, A. D., Jr. (2000) An improved empirical potential energy function for molecular simulations of phospholipids. J. Phys. Chem. B 104, 7510–7515.
126. Feller, S. E., Gawrisch, K., and MacKerell, A. D., Jr. (2002) Polyunsaturated fatty acids in lipid bilayers: Intrinsic and environmental contributions to their unique physical properties. J. Am. Chem. Soc. 124, 318–326.
127. Spijker, P., Vaidehi, N., Freddolino, P. L., Hilbers, P. A. J., and Goddard, W. A. (2006) Dynamic behavior of fully solvated beta 2-adrenergic receptor, embedded in the membrane with bound agonist or antagonist. Proc. Natl. Acad. Sci. U. S. A. 103, 4882–4887.
128. Lagüe, P., Roux, B., and Pastor, R. W. (2005) Molecular dynamics simulations of the influenza hemagglutinin fusion peptide in micelles and bilayers: Conformational analysis of peptide and lipids. J. Mol. Biol. 354, 1129–1141.
129. Tajkhorshid, E., Nollert, P., Jensen, M. O., Miercke, L. J. W., O'Connell, J., Stroud, R. M., and Schulten, K. (2002) Control of the selectivity of the aquaporin water channel family by global orientational tuning. Science 296, 525–530.
130. Berger, O., Edholm, O., and Jahnig, F. (1997) Molecular dynamics simulations of a fluid bilayer of dipalmitoylphosphatidylcholine at full hydration, constant pressure, and constant temperature. Biophys. J. 72, 2002–2013.
131. Chandrasekhar, I., Kastenholz, M., Lins, R. D., Oostenbrink, C., Schuler, L. D., Tieleman, D. P., and van Gunsteren, W. F. (2003) A consistent potential energy parameter set for lipids: Dipalmitoylphosphatidylcholine as a benchmark of the GROMOS96 45A3 force field. Eur. Biophys. J. Biophys. Lett. 32, 67–77.
132. Glattli, A., Chandrasekhar, I., and van Gunsteren, W. F. (2006) A molecular dynamics study of the bee venom melittin in aqueous solution, in methanol, and inserted in a phospholipid bilayer. Eur. Biophys. J. Biophys. Lett. 35, 255–267.
133. Leontiadou, H., Mark, A. E., and Marrink, S. J. (2006) Antimicrobial peptides in action. J. Am. Chem. Soc. 128, 12156–12161.
134. Grottesi, A., Domene, C., Hall, B., and Sansom, M. S. P. (2005) Conformational dynamics of M2 helices in KirBac channels: Helix flexibility in relation to gating via molecular dynamics simulations. Biochemistry 44, 14586–14594.
135. Moore, P. B., Lopez, C. F., and Klein, M. L. (2001) Dynamical properties of a hydrated lipid bilayer from a multinanosecond molecular dynamics simulation. Biophys. J. 81, 2484–2494.
136. Liepina, I., Czaplewski, C., Janmey, P., and Liwo, A. (2003) Molecular dynamics study of a gelsolin-derived peptide binding to a lipid bilayer containing phosphatidylinositol 4,5-bisphosphate. Biopolymers 71, 49–70.
137. Bastug, T. and Kuyucak, S. (2005) Test of molecular dynamics force fields in gramicidin A. Eur. Biophys. J. Biophys. Lett. 34, 377–382.
138. Woods, R. J., Dwek, R. A., Edge, C. J., and Fraserreid, B. (1995) Molecular mechanical and molecular dynamical simulations of glycoproteins and oligosaccharides. 1. GLYCAM 93 parameter development. J. Phys. Chem. 99, 3832–3846.
139. Woods, R. J. and Chappelle, R. (2000) Restrained electrostatic potential atomic partial charges for condensed-phase simulations of carbohydrates. Theochem—J. Mol. Struct. 527, 149–156.
140. Basma, M., Sundara, S., Calgan, D., Vernali, T., and Woods, R. J. (2001) Solvated ensemble averaging in the calculation of partial atomic charges. J. Comput. Chem. 22, 1125–1137.
141. Kawatkar, S. P., Kuntz, D. A., Woods, R. J., Rose, D. R., and Boons, G. J. (2006) Structural basis of the inhibition of Golgi alpha-mannosidase II by mannostatin A and the role of the thiomethyl moiety in ligand-protein interactions. J. Am. Chem. Soc. 128, 8310–8319.
142. Lins, R. D., Pereira, C. S., and Hünenberger, P. H. (2004) Trehalose-protein interaction in aqueous solution. Proteins 55, 177–186.
143. Bosques, C. J., Tschampel, S. M., Woods, R. J., and Imperiali, B. (2004) Effects of glycosylation on peptide conformation: A synergistic experimental and computational study. J. Am. Chem. Soc. 126, 8421–8425.
144. Ford, M. G., Weimar, T., Kohli, T., and Woods, R. J. (2003) Molecular dynamics simulations of galectin-1-oligosaccharide complexes reveal the molecular basis for ligand diversity. Proteins 53, 229–240.
145. Kirschner, K. N. and Woods, R. J. (2001) Solvent interactions determine carbohydrate conformation. Proc. Natl. Acad. Sci. U. S. A. 98, 10541–10545.
146. Damm, W., Frontera, A., TiradoRives, J., and Jorgensen, W. L. (1997) OPLS all-atom force field for carbohydrates. J. Comput. Chem. 18, 1955–1970.
147. Margulis, C. J. (2005) Computational study of the dynamics of mannose disaccharides free in solution and bound to the potent anti-HIV virucidal protein cyanovirin. J. Phys. Chem. B 109, 3639–3647.
148. Kony, D., Damm, W., Stoll, S., and van Gunsteren, W. F. (2002) An improved OPLS-AA force field for carbohydrates. J. Comput. Chem. 23, 1416–1429.
149. Kuttel, M., Brady, J. W., and Naidoo, K. J. (2002) Carbohydrate solution simulations: Producing a force field with experimentally consistent primary alcohol rotational frequencies and populations. J. Comput. Chem. 23, 1236–1243.
150. Lins, R. D. and Hünenberger, P. H. (2005) A new GROMOS force field for hexopyranose-based carbohydrates. J. Comput. Chem. 26, 1400–1412.
151. Daura, X., Mark, A. E., and van Gunsteren, W. F. (1998) Parametrization of aliphatic CHn united atoms of GROMOS96 force field. J. Comput. Chem. 19, 535–547.
152. Schuler, L. D., Daura, X., and Van Gunsteren, W. F. (2001) An improved GROMOS96 force field for aliphatic hydrocarbons in the condensed phase. J. Comput. Chem. 22, 1205–1218.
153. Wang, J. M., Wolf, R. M., Caldwell, J. W., Kollman, P. A., and Case, D. A. (2004) Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174.
154. Ren, P. Y. and Ponder, J. W. (2002) Consistent treatment of inter- and intramolecular polarization in molecular mechanics calculations. J. Comput. Chem. 23, 1497–1506.
155. Patel, S., MacKerell, A. D., Jr., and Brooks, C. L. III (2004) CHARMM fluctuating charge force field for proteins: II—Protein/solvent properties from molecular dynamics simulations using a nonadditive electrostatic model. J. Comput. Chem. 25, 1504–1514.
156. Kim, B. C., Young, T., Harder, E., Friesner, R. A., and Berne, B. J. (2005) Structure and dynamics of the solvation of bovine pancreatic trypsin inhibitor in explicit water: A comparative study of the effects of solvent and protein polarizability. J. Phys. Chem. B 109, 16529–16538.
157. Anisimov, V. M., Lamoureux, G., Vorobyov, I. V., Huang, N., Roux, B., and MacKerell, A. D., Jr. (2005) Determination of electrostatic parameters for a polarizable force field based on the classical Drude oscillator. J. Chem. Theory Comput. 1, 153–168.
Acknowledgments
Financial support from the National Institutes of Health (NIH; R01GM051501 and R01GM070855 to ADM, and F32CA119771 to OG) and from the University of Maryland Computer-Aided Drug Design Center is acknowledged.
Author information
Authors and Affiliations
Editor information
Rights and permissions
Copyright information
© 2008 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Guvench, O., MacKerell, A.D. (2008). Comparison of Protein Force Fields for Molecular Dynamics Simulations. In: Kukol, A. (eds) Molecular Modeling of Proteins. Methods Molecular Biology™, vol 443. Humana Press. https://doi.org/10.1007/978-1-59745-177-2_4
Download citation
DOI: https://doi.org/10.1007/978-1-59745-177-2_4
Publisher Name: Humana Press
Print ISBN: 978-1-58829-864-5
Online ISBN: 978-1-59745-177-2
eBook Packages: Springer Protocols