Abstract
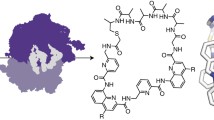
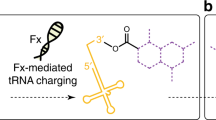
Flexizymes, highly flexible tRNA aminoacylation ribozymes, have enabled charging of virtually any amino acid (including non-proteogenic ones) onto tRNA molecules. Coupling to a custom-made in vitro translation system, namely the flexible in vitro translation (FIT) system, has unveiled the remarkable tolerance of the ribosome toward molecules, remote from what nature has selected to carry out its elaborate functions. Among the very diverse molecules and chemistries that have been ribosomally incorporated, a plethora of entities capable of mediating intramolecular cyclization have revolutionized the design and discovery of macrocyclic peptides. These macrocyclization reactions (which can be spontaneous, chemical, or enzymatic) have all served as tools for the discovery of peptides with natural-like structures and properties. Coupling of the FIT system and mRNA display techniques, known as the random non-standard peptide integrated discovery (RaPID) system, has in turn allowed for the simultaneous screening of trillions of macrocyclic peptides against challenging biological targets. The macrocyclization methodologies are chosen depending on the structural and functional characteristics of the desired molecule. Thus, they can emanate from the peptide’s N-terminus or its side chains, attributing flexibility or rigidity, or even result in the installation of fluorescent probes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Heinis C, Rutherford T, Freund S, Winter G (2009) Phage-encoded combinatorial chemical libraries based on bicyclic peptides. Nat Chem Biol 5(7):502–507. https://doi.org/10.1038/nchembio.184
Chen S, Rentero Rebollo I, Buth SA, Morales-Sanfrutos J, Touati J, Leiman PG, Heinis C (2013) Bicyclic peptide ligands pulled out of cysteine-rich peptide libraries. J Am Chem Soc 135(17):6562–6569. https://doi.org/10.1021/ja400461h
Daly NL, Craik DJ (2011) Bioactive cystine knot proteins. Curr Opin Chem Biol 15(3):362–368. https://doi.org/10.1016/j.cbpa.2011.02.008
Pluckthun A (2012) Ribosome display: a perspective. Methods Mol Biol 805:3–28. https://doi.org/10.1007/978-1-61779-379-0_1
Roberts RW (1999) Totally in vitro protein selection using mRNA-protein fusions and ribosome display. Curr Opin Chem Biol 3(3):268–273. https://doi.org/10.1016/S1367-5931(99)80042-8
Rogers JM, Suga H (2015) Discovering functional, non-proteinogenic amino acid containing, peptides using genetic code reprogramming. Org Biomol Chem 13(36):9353–9363. https://doi.org/10.1039/c5ob01336d
Hoesl MG, Budisa N (2012) Recent advances in genetic code engineering in Escherichia coli. Curr Opin Biotechnol 23(5):751–757. https://doi.org/10.1016/j.copbio.2011.12.027
Terasaka N, Hayashi G, Katoh T, Suga H (2014) An orthogonal ribosome-tRNA pair via engineering of the peptidyl transferase center. Nat Chem Biol 10(7):555–557. https://doi.org/10.1038/nchembio.1549
Goto Y, Katoh T, Suga H (2011) Flexizymes for genetic code reprogramming. Nat Protoc 6(6):779–790. https://doi.org/10.1038/nprot.2011.331
Murakami H, Ohta A, Ashigai H, Suga H (2006) A highly flexible tRNA acylation method for non-natural polypeptide synthesis. Nat Methods 3(5):357–359. https://doi.org/10.1038/nmeth877
Illangasekare M, Sanchez G, Nickles T, Yarus M (1995) Aminoacyl-RNA synthesis catalyzed by an RNA. Science 267(5198):643–647
Illangasekare M, Yarus M (1999) Specific, rapid synthesis of Phe-RNA by RNA. Proc Natl Acad Sci U S A 96(10):5470–5475
Lohse PA, Szostak JW (1996) Ribozyme-catalysed amino-acid transfer reactions. Nature 381(6581):442–444. https://doi.org/10.1038/381442a0
Lee N, Bessho Y, Wei K, Szostak JW, Suga H (2000) Ribozyme-catalyzed tRNA aminoacylation. Nat Struct Biol 7(1):28–33. https://doi.org/10.1038/71225
Saito H, Kourouklis D, Suga H (2001) An in vitro evolved precursor tRNA with aminoacylation activity. EMBO J 20(7):1797–1806. https://doi.org/10.1093/emboj/20.7.1797
Morimoto J, Hayashi Y, Iwasaki K, Suga H (2011) Flexizymes: their evolutionary history and the origin of catalytic function. Acc Chem Res 44(12):1359–1368. https://doi.org/10.1021/ar2000953
Murakami H, Kourouklis D, Suga H (2003) Using a solid-phase ribozyme aminoacylation system to reprogram the genetic code. Chem Biol 10(11):1077–1084
Ramaswamy K, Saito H, Murakami H, Shiba K, Suga H (2004) Designer ribozymes: programming the tRNA specificity into flexizyme. J Am Chem Soc 126(37):11454–11455. https://doi.org/10.1021/ja046843y
Saito H, Watanabe K, Suga H (2001) Concurrent molecular recognition of the amino acid and tRNA by a ribozyme. RNA 7(12):1867–1878
Murakami H, Saito H, Suga H (2003) A versatile tRNA aminoacylation catalyst based on RNA. Chem Biol 10(7):655–662
Shimizu Y, Kanamori T, Ueda T (2005) Protein synthesis by pure translation systems. Methods 36(3):299–304. https://doi.org/10.1016/j.ymeth.2005.04.006
Katoh T, Suga H (2018) Ribosomal incorporation of consecutive beta-amino acids. J Am Chem Soc 140(38):12159–12167. https://doi.org/10.1021/jacs.8b07247
Ohshiro Y, Nakajima E, Goto Y, Fuse S, Takahashi T, Doi T, Suga H (2011) Ribosomal synthesis of backbone-macrocyclic peptides containing gamma-amino acids. Chembiochem 12(8):1183–1187. https://doi.org/10.1002/cbic.201100104
Rogers JM, Kwon S, Dawson SJ, Mandal PK, Suga H, Huc I (2018) Ribosomal synthesis and folding of peptide-helical aromatic foldamer hybrids. Nat Chem 10(4):405–412. https://doi.org/10.1038/s41557-018-0007-x
Torikai K, Suga H (2014) Ribosomal synthesis of an amphotericin-B inspired macrocycle. J Am Chem Soc 136(50):17359–17361. https://doi.org/10.1021/ja508648s
Yamagishi Y, Ashigai H, Goto Y, Murakami H, Suga H (2009) Ribosomal synthesis of cyclic peptides with a fluorogenic oxidative coupling reaction. Chembiochem 10(9):1469–1472. https://doi.org/10.1002/cbic.200900021
Atsushi O, Hiroshi M, Hiroaki S, (2008) Polymerization of α-Hydroxy Acids by Ribosomes. Chem Bio Chem 9(17):2773–2778
Yamagishi Y, Shoji I, Miyagawa S, Kawakami T, Katoh T, Goto Y, Suga H (2011) Natural product-like macrocyclic N-methyl-peptide inhibitors against a ubiquitin ligase uncovered from a ribosome-expressed de novo library. Chem Biol 18(12):1562–1570. https://doi.org/10.1016/j.chembiol.2011.09.013
Roberts RW, Szostak JW (1997) RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc Natl Acad Sci U S A 94(23):12297–12302
Nemoto N, Miyamoto-Sato E, Husimi Y, Yanagawa H (1997) In vitro virus: bonding of mRNA bearing puromycin at the 3′-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro. FEBS Lett 414(2):405–408
Kawamura A, Munzel M, Kojima T, Yapp C, Bhushan B, Goto Y, Tumber A, Katoh T, King ON, Passioura T, Walport LJ, Hatch SB, Madden S, Muller S, Brennan PE, Chowdhury R, Hopkinson RJ, Suga H, Schofield CJ (2017) Highly selective inhibition of histone demethylases by de novo macrocyclic peptides. Nat Commun 8:14773. https://doi.org/10.1038/ncomms14773
Goto Y, Suga H (2009) Translation initiation with initiator tRNA charged with exotic peptides. J Am Chem Soc 131(14):5040–5041. https://doi.org/10.1021/ja900597d
Roberts KD, Lambert JN, Ede NJ, Bray AM (1998) Efficient synthesis of thioether-based cyclic peptide libraries. Tetrahedron Lett 39(45):8357–8360. https://doi.org/10.1016/S0040-4039(98)01843-7
Lung F-D, King CR, Roller PP (1999) Development of non-phosphorylated cyclic thioether peptide to the Grb2-SH2 domain. Lett Pept Sci 6(1):45–49. https://doi.org/10.1007/BF02443617
Yu L, Lai Y, Wade JV, Coutts SM (1998) A simple and efficient method for the syntheses of thioether cyclic peptides. Tetrahedron Lett 39(37):6633–6636. https://doi.org/10.1016/S0040-4039(98)01397-5
Iwasaki K, Goto Y, Katoh T, Suga H (2012) Selective thioether macrocyclization of peptides having the N-terminal 2-chloroacetyl group and competing two or three cysteine residues in translation. Org Biomol Chem 10(30):5783–5786. https://doi.org/10.1039/C2OB25306B
Ezure T, Nanatani K, Sato Y, Suzuki S, Aizawa K, Souma S, Ito M, Hohsaka T, von Heijine G, Utsumi T, Abe K, Ando E, Uozumi N (2014) A cell-free translocation system using extracts of cultured insect cells to yield functional membrane proteins. PLoS One 9(12):e112874. https://doi.org/10.1371/journal.pone.0112874
Kawakami T, Ohta A, Ohuchi M, Ashigai H, Murakami H, Suga H (2009) Diverse backbone-cyclized peptides via codon reprogramming. Nat Chem Biol 5(12):888–890. https://doi.org/10.1038/nchembio.259
Sako Y, Goto Y, Murakami H, Suga H (2008) Ribosomal synthesis of peptidase-resistant peptides closed by a nonreducible inter-side-chain bond. ACS Chem Biol 3(4):241–249. https://doi.org/10.1021/cb800010p
Berkowitz DB, McFadden JM, Sloss MK (2000) Engineering acyclic stereocontrol in the alkylation of vinylglycine-derived dianions: asymmetric synthesis of higher alpha-vinyl amino acids. J Org Chem 65(10):2907–2918
Bretschneider T, Miltz W, Münster P, Steglich W (1988) New syntheses of α-amino acids based on n-acylimino acetates. Tetrahedron 44(17):5403–5414. https://doi.org/10.1016/S0040-4020(01)86046-4
Goto Y, Iwasaki K, Torikai K, Murakami H, Suga H (2009) Ribosomal synthesis of dehydrobutyrine- and methyllanthionine-containing peptides. Chem Commun (Camb) (23):3419–3421. https://doi.org/10.1039/b904314d
Nakajima E, Goto Y, Sako Y, Murakami H, Suga H (2009) Ribosomal synthesis of peptides with C-terminal lactams, thiolactones, and alkylamides. Chembiochem 10(7):1186–1192. https://doi.org/10.1002/cbic.200900058
Bashiruddin NK, Nagano M, Suga H (2015) Synthesis of fused tricyclic peptides using a reprogrammed translation system and chemical modification. Bioorg Chem 61:45–50. https://doi.org/10.1016/j.bioorg.2015.06.002
Sako Y, Morimoto J, Murakami H, Suga H (2008) Ribosomal synthesis of bicyclic peptides via two orthogonal inter-side-chain reactions. J Am Chem Soc 130(23):7232–7234. https://doi.org/10.1021/ja800953c
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Tsiamantas, C., Otero-Ramirez, M.E., Suga, H. (2019). Discovery of Functional Macrocyclic Peptides by Means of the RaPID System. In: Goetz, G. (eds) Cyclic Peptide Design. Methods in Molecular Biology, vol 2001. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9504-2_14
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9504-2_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9503-5
Online ISBN: 978-1-4939-9504-2
eBook Packages: Springer Protocols