Abstract
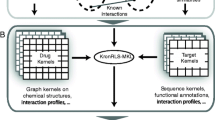
Drug-target networks have an important role in pharmaceutical innovation, drug lead discovery, and recent drug repositioning tasks. Many different in silico approaches for the identification of new drug-target interactions have been proposed, many of them based on a particular class of machine learning algorithms called kernel methods. These pattern classification algorithms are able to incorporate previous knowledge in the form of similarity functions, i.e., a kernel, and they have been successful in a wide range of supervised learning problems. The selection of the right kernel function and its respective parameters can have a large influence on the performance of the classifier. Recently, multiple kernel learning algorithms have been introduced to address this problem, enabling one to combine multiple kernels into large drug-target interaction spaces in order to integrate multiple sources of biological information simultaneously. The Kronecker regularized least squares with multiple kernel learning (KronRLS-MKL) is a machine learning algorithm that aims at integrating heterogeneous information sources into a single chemogenomic space to predict new drug-target interactions. This chapter describes how to obtain data from heterogeneous sources and how to implement and use KronRLS-MKL to predict new interactions.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Csermely P, Korcsmáros T, Kiss HJM et al (2013) Structure and dynamics of molecular networks: a novel paradigm of drug discovery: a comprehensive review. Pharmacol Ther 138:333–408
Ding H, Takigawa I, Mamitsuka H et al (2014) Similarity-based machine learning methods for predicting drug-target interactions: a brief review. Brief Bioinform 15:734–747
Rogers DJ, Tanimoto TT (1960) A computer program for classifying plants. Science 132:1115–1118
Ralaivola L, Swamidass SJ, Saigo H et al (2005) Graph kernels for chemical informatics. Neural Netw 18:1093–1110
Smith TF, Waterman MS (1981) Identification of common molecular subsequences. J Mol Biol 147:195–197
Yamanishi Y, Kotera M, Kanehisa M et al (2010) Drug-target interaction prediction from chemical, genomic and pharmacological data in an integrated framework. Bioinformatics 26:i246–i254
Nascimento ACA, Prudêncio RBC, Costa IG (2016) A multiple kernel learning algorithm for drug-target interaction prediction. BMC Bioinformatics 17:46
Schölkopf B, Smola AJ (2002) Learning with kernels: support vector machines, regularization, optimization, and beyond. MIT Press
Schölkopf B, Tsuda K, Vert J-P (2004) Kernel methods in computational biology. MIT Press
Kanehisa M, Araki M, Goto S et al (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–D484
Gunther S, Kuhn M, Dunkel M et al (2007) SuperTarget and matador: resources for exploring drug-target relationships. Nucleic Acids Res 36:D919–D922
Gilson MK, Liu T, Baitaluk M et al (2016) BindingDB in 2015: a public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res 44:D1045–D1053
Kuhn M, Szklarczyk D, Pletscher-Frankild S et al (2013) STITCH 4: integration of protein–chemical interactions with user data. Nucleic Acids Res 42:D401–D407
Wishart DS, Knox C, Guo AC et al (2008) DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res 36:D901–D906
Pavlidis P, Weston J, Cai J et al (2001) Gene functional classification from heterogeneous data. In: Proceedings of the fifth annual international conference on computational biology—RECOMB ‘01,
Ben-Hur A, Noble WS (2005) Kernel methods for predicting protein-protein interactions. Bioinformatics 21(Suppl 1):i38–i46
van Laarhoven T, Nabuurs SB, Marchiori E (2011) Gaussian interaction profile kernels for predicting drug-target interaction. Bioinformatics 27:3036–3043
Hao M, Bryant SH, Wang Y (2018) Open-source chemogenomic data-driven algorithms for predicting drug–target interactions. Brief Bioinform
Wang Y, Xiao J, Suzek TO et al (2009) PubChem: a public information system for analyzing bioactivities of small molecules. Nucleic Acids Res 37:W623–W633
Rupp M, Schneider G (2010) Graph kernels for molecular similarity. Mol Inform 29:266–273
Klambauer G, Wischenbart M, Mahr M et al (2015) Rchemcpp: a web service for structural analoging in ChEMBL, Drugbank and the connectivity map. Bioinformatics 31:3392–3394
Kuhn M, Letunic I, Jensen LJ et al (2016) The SIDER database of drugs and side effects. Nucleic Acids Res 44:D1075–D1079
Lamb J (2006) The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313:1929–1935
Edgar R (2002) Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Perlman L, Gottlieb A, Atias N et al (2011) Combining drug and gene similarity measures for drug-target elucidation. J Comput Biol 18:133–145
Wang K, Sun J, Zhou S et al (2013) Prediction of drug-target interactions for drug repositioning only based on genomic expression similarity. PLoS Comput Biol 9:e1003315
Wang Y-C, Zhang C-H, Deng N-Y et al (2011) Kernel-based data fusion improves the drug-protein interaction prediction. Comput Biol Chem 35:353–362
Palme J, Hochreiter S, Bodenhofer U (2015) KeBABS: an R package for kernel-based analysis of biological sequences: Fig. 1. Bioinformatics 31:2574–2576
Perrimon N, Friedman A, Mathey-Prevot B et al (2007) Drug-target identification in Drosophila cells: combining high-throughout RNAi and small-molecule screens. Drug Discov Today 12:28–33
Smedley D, Haider S, Durinck S et al (2015) The BioMart community portal: an innovative alternative to large, centralized data repositories. Nucleic Acids Res 43:W589–W598
Ovaska K, Laakso M, Hautaniemi S (2008) Fast gene ontology based clustering for microarray experiments. BioData Min 1:11
Byrd RH, Hribar ME, Nocedal J (1999) An interior point algorithm for large-scale nonlinear programming. SIAM J Optim 9:877–900
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Nascimento, A.C.A., Prudêncio, R.B.C., Costa, I.G. (2019). A Drug-Target Network-Based Supervised Machine Learning Repurposing Method Allowing the Use of Multiple Heterogeneous Information Sources. In: Vanhaelen, Q. (eds) Computational Methods for Drug Repurposing. Methods in Molecular Biology, vol 1903. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8955-3_17
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8955-3_17
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8954-6
Online ISBN: 978-1-4939-8955-3
eBook Packages: Springer Protocols