Abstract
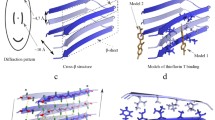
Many proteins and peptides are able to self-assemble in solution in vitro and in vivo to form amyloid-like fibrils. These fibrils share common structural characteristics. In order for a fibril to be characterized as amyloid, it is expected to fit certain criteria including the composition of cross-β. Here we describe how the formation of amyloid fibrils can be characterized in vitro using a variety of methods including circular dichroism and intrinsic tyrosine/tryptophan fluoresence to follow conformational changes; Thioflavin and/or ThS assembly to monitor nucleation and growth; transmission electron microscopy to visualize fibrillar morphology and X-ray fiber diffraction to examine cross-β structure.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sipe JD, Benson MD, Buxbaum JN, Ikeda S, Merlini G, Saraiva MJ, Westermark P (2014) Nomenclature 2014: amyloid fibril proteins and clinical classification of the amyloidosis. Amyloid 21(4):221–224. https://doi.org/10.3109/13506129.2014.964858
Puchtler H, Sweat F (1965) Congo red as a stain for fluorescence microscopy of amyloid. J Histochem Cytochem 13(8):693–694
LeVine H 3rd (1999) Quantification of beta-sheet amyloid fibril structures with thioflavin T. Methods Enzymol 309:274–284
Glabe CG (2004) Conformation-dependent antibodies target diseases of protein misfolding. Trends Biochem Sci 29(10):542–547
Lambert MP, Velasco PT, Chang L, Viola KL, Fernandez S, Lacor PN, Khuon D, Gong Y, Bigio EH, Shaw P, De Felice FG, Krafft GA, Klein WL (2007) Monoclonal antibodies that target pathological assemblies of Abeta. J Neurochem 100(1):23–35. https://doi.org/10.1111/j.1471-4159.2006.04157.x JNC4157 [pii]
Marshall KE, Hicks MR, Williams TL, Hoffmann SV, Rodger A, Dafforn TR, Serpell LC (2010) Characterizing the assembly of the Sup35 yeast prion fragment, GNNQQNY: structural changes accompany a fiber-to-crystal switch. Biophys J 98(2):330–338. https://doi.org/10.1016/j.bpj.2009.10.020 S0006-3495(09)01627-0 [pii]
Sawaya MR, Sambashivan S, Nelson R, Ivanova MI, Sievers SA, Apostol MI, Thompson MJ, Balbirnie M, Wiltzius JJ, McFarlane HT, Madsen AO, Riekel C, Eisenberg D (2007) Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447(7143):453–457
Tycko R (2001) Biomolecular solid state NMR: advances in structural methodology and applications to peptide and protein fibrils. Annu Rev Phys Chem 52:575–606
Fitzpatrick AWP, Falcon B, He S, Murzin AG, Murshudov G, Garringer HJ, Crowther RA, Ghetti B, Goedert M, Scheres SHW (2017) Cryo-EM structures of tau filaments from Alzheimer's disease. Nature 547(7662):185–190. https://doi.org/10.1038/nature23002
van der Wel PC, Lewandowski JR, Griffin RG (2007) Solid-state NMR study of amyloid nanocrystals and fibrils formed by the peptide GNNQQNY from yeast prion protein Sup35p. J Am Chem Soc 129(16):5117–5130
White HE, Hodgkinson JL, Jahn TR, Cohen-Krausz S, Gosal WS, Muller S, Orlova EV, Radford SE, Saibil HR (2009) Globular tetramers of beta(2)-microglobulin assemble into elaborate amyloid fibrils. J Mol Biol 389(1):48–57
Xue WF, Radford SE (2013) An imaging and systems modeling approach to fibril breakage enables prediction of amyloid behavior. Biophys J 105(12):2811–2819. https://doi.org/10.1016/j.bpj.2013.10.034
Goldsbury C, Kistler J, Aebi U, Arvinte T, Cooper GJ (1999) Watching amyloid fibrils grow by time-lapse atomic force microscopy. J Mol Biol 285(1):33–39
CCP4 (1994) The CCP4 suite programs for crystallography. Acta Cryst D50:760–763
Hammersley AP (1998) FIT2D V9.129 Reference Manual V3.1. ESRF Internal Report ESRF98HA01T
Makin OS, Sikorski P, Serpell LC (2007) CLEARER: a new tool for the analysis of X-ray fibre diffraction patterns and diffraction simulation from atomic structural models. Appl Cryst 40(5):966–972
Broersen K, Jonckheere W, Rozenski J, Vandersteen A, Pauwels K, Pastore A, Rousseau F, Schymkowitz J (2011) A standardized and biocompatible preparation of aggregate-free amyloid beta peptide for biophysical and biological studies of Alzheimer's disease. Protein Eng Des Sel 24(9):743–750. https://doi.org/10.1093/protein/gzr020
Bitan G, Frandinger EA, Spring S, Teplow D (2005) Neurotoxic protein oligomers- what you see is not always what you get. Amyloid 2(Jun):88–95
Fezoui Y, Hartley DM, Harper JD, Khurana R, Walsh DM, Condron MM, Selkoe DJ, Lansbury PT Jr, Fink AL, Teplow DB (2000) An improved method of preparing the amyloid beta-protein for fibrillogenesis and neurotoxicity experiments. Amyloid 7(3):166–178
Kelly SM, Jess TJ, Price NC (2005) How to study proteins by circular dichroism. Biochim Biophys Acta 1751(2):119–139. https://doi.org/10.1016/j.bbapap.2005.06.005
Morris KL, Rodger A, Hicks MR, Debulpaep M, Schymkowitz J, Rousseau F, Serpell LC (2013) Exploring the sequence-structure relationship for amyloid peptides. Biochem J 450(2):275–283. https://doi.org/10.1042/BJ20121773
Dafforn TR, Rajendra J, Halsall DJ, Serpell LC, Rodger A (2004) Protein fiber linear dichroism for structure determination and kinetics in a low-volume, low-wavelength couette flow cell. Biophys J 86(1 Pt 1):404–410
Rodger A, Marrington R, Geeves MA, Hicks M, de Alwis L, Halsall DJ, Dafforn TR (2006) Looking at long molecules in solution: what happens when they are subjected to Couette flow? Phys Chem Chem Phys 8(27):3161–3171
Rodger A, Norden B (1997) Circular Dichroism and linear Dichroism. Oxford University Press, Oxford
Morris KL, Serpell LC (2012) X-ray fibre diffraction studies of amyloid fibrils. Methods Mol Biol 849:121–135. https://doi.org/10.1007/978-1-61779-551-0_9
Winn MD (2003) An overview of the CCP4 project in protein crystallography: an example of a collaborative project. J Synchrotron Radiat 10. (Pt 1:23–25
Makin OS, Serpell LC (2005) X-ray diffraction studies of amyloid structure. In: Sigurdsson EM (ed) Amyloid proteins: methods and protocols. Humana press, Totowa, NJ, pp 67–80
Al-Hilaly YK, Biasetti L, Blakeman BJ, Pollack SJ, Zibaee S, Abdul-Sada A, Thorpe JR, Xue WF, Serpell LC (2016) The involvement of dityrosine crosslinking in alpha-synuclein assembly and deposition in Lewy bodies in Parkinson's disease. Sci Rep 6:39171. https://doi.org/10.1038/srep39171
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Vadukul, D.M., Al-Hilaly, Y.K., Serpell, L.C. (2019). Methods for Structural Analysis of Amyloid Fibrils in Misfolding Diseases. In: Gomes, C. (eds) Protein Misfolding Diseases. Methods in Molecular Biology, vol 1873. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8820-4_7
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8820-4_7
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8819-8
Online ISBN: 978-1-4939-8820-4
eBook Packages: Springer Protocols