Abstract
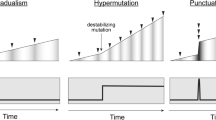
Genome chaos, or karyotype chaos, represents a powerful survival strategy for somatic cells under high levels of stress/selection. Since the genome context, not the gene content, encodes the genomic blueprint of the cell, stress-induced rapid and massive reorganization of genome topology functions as a very important mechanism for genome (karyotype) evolution. In recent years, the phenomenon of genome chaos has been confirmed by various sequencing efforts, and many different terms have been coined to describe different subtypes of the chaotic genome including “chromothripsis,” “chromoplexy,” and “structural mutations.” To advance this exciting field, we need an effective experimental system to induce and characterize the karyotype reorganization process. In this chapter, an experimental protocol to induce chaotic genomes is described, following a brief discussion of the mechanism and implication of genome chaos in cancer evolution.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Heng HH (2015) Debating cancer: the paradox in cancer research. World Scientific, Singapore
Heng HH, Stevens JB, Liu G et al (2004) Imaging genome abnormalities in cancer research. Cell Chromosome 3:1
Heng HH, Stevens JB, Liu G et al (2006) Stochastic cancer progression driven by nonclonal chromosome aberrations. J Cell Physiol 208:461–472
Heng HH, Stevens JB, Bremer SW et al (2011) Evolutionary mechanisms and diversity in cancer. Adv Cancer Res 112:217–253
Heng HH, Liu G, Stevens JB et al (2011) Decoding the genome beyond sequencing: the new phase of genomic research. Genomics 98(4):242–252
Heng HH, Bremer SW, Stevens JB et al (2013) Chromosomal instability (CIN): what it is and why it is crucial to cancer evolution. Cancer Metastasis Rev 32(3–4):325–340
Heng HH, Liu G, Stevens JB et al (2013) Karyotype heterogeneity and unclassified chromosomal abnormalities. Cytogenet Genome Res 139(3):144–157
Stephens PJ, Greenman CD, Fu B et al (2011) Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144(1):27–40
Horne SD, Heng HH (2014) Genome chaos, chromothripsis and cancer evolution. J Cancer Stud Ther 1:1–6
Heng HH (2017) Cancer genomic landscape. In: Ujvari B, Roche B, Thomas F (eds) Ecology and evolution of cancer. Academic Press, Elsevier, pp 69–86
Baca SC, Prandi D, Lawrence MS et al (2013) Punctuated evolution of prostate cancer genomes. Cell 153(3):666–677
Crasta K, Ganem NJ, Dagher R et al (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482(7383):53–58
Forment JV, Kaidi A, Jackson SP (2012) Chromothripsis and cancer: causes and consequences of chromosome shattering. Nat Rev Cancer 12(10):663–670
Holland AJ, Cleveland DW (2012) Chromoanagenesis and cancer: mechanisms and consequences of localized, complex chromosomal rearrangements. Nat Med 18(11):1630–1638
Inaki K, Liu ET (2012) Structural mutations in cancer: mechanistic and functional insights. Trends Genet 28(11):550–559
Jones MJ, Jallepalli PV (2012) Chromothripsis: chromosomes in crisis. Dev Cell 23(5):908–917
Liu P, Erez A, Nagamani SC et al (2011) Chromosome catastrophes involve replication mechanisms generating complex genomic rearrangements. Cell 146(6):889–903
Malhotra A, Lindberg M, Faust GG et al (2013) Breakpoint profiling of 64 cancer genomes reveals numerous complex rearrangements spawned by homology-independent mechanisms. Genome Res 23:762–776
Righolt C, Mai S (2012) Shattered and stitched chromosomes-chromothripsis and chromoanasynthesis-manifestations of a new chromosome crisis? Genes Chromosomes Cancer 51(11):975–981
Setlur SR, Lee C (2012) Tumor archaeology reveals that mutations love company. Cell 149(9):959–961
Tubio JM, Estivill X (2011) Cancer: when catastrophe strikes a cell. Nature 470(7335):476–477
Heng HH (2007) Karyotypic chaos, a form of non-clonal chromosome aberrations, plays a key role for cancer progression and drug resistance. FASEB: Nuclear Structure and Cancer. Vermont Academy, Saxtons River, VT
Duesberg P (2007) Chromosomal chaos and cancer. Sci Am 296(5):52–59
Zhang CZ, Spektor A, Cornils H et al (2015) Chromothripsis from DNA damage in micronuclei. Nature 522(7555):179–184
Maciejowski J, Li Y, Bosco N, Campbell PJ, de Lange T (2015) Chromothripsis and Kataegis induced by telomere crisis. Cell 163(7):1641–1654
Morishita M, Muramatsu T, Suto Y et al (2016) Chromothripsis-like chromosomal rearrangements induced by ionizing radiation using proton microbeam irradiation system. Oncotarget 7(9):10182–10192
Mardin BR, Drainas AP, Waszak SM et al (2015) A cell-based model system links chromothripsis with hyperploidy. Mol Syst Biol 11(9):828
Stevens JB, Liu G, Bremer SW et al (2007) Mitotic cell death by chromosome fragmentation. Cancer Res 67(16):7686–7694
Stevens JB, Abdallah BY, Liu G et al (2011) Diverse system stresses: common mechanisms of chromosome fragmentation. Cell Death Dis 2:e178
Stevens JB, Abdallah BY, Liu G et al (2013) Heterogeneity of cell death. Cytogenet Genome Res 139(3):164–173
Ye CJ, Stevens JB, Liu G et al (2009) Genome based cell population heterogeneity promotes tumorigenicity: the evolutionary mechanism of cancer. J Cell Physiol 219(2):288–300
Horne SD, Chowdhury SK, Heng HH (2014) Stress, genomic adaptation, and the evolutionary trade-off. Front Genet 5:92
Heng HH, Regan S, Ye CJ (2016) Genotype, environment, and evolutionary mechanism of diseases. Environ Dis 1:14–23
Heng HH (2007) Cancer genome sequencing: the challenges ahead. BioEssays 29(8):783–794
Heng HH (2009) The genome-centric concept: resynthesis of evolutionary theory. BioEssays 31(5):512–525
Horne SD, Ye CJ, Heng HH (2015) Chromosomal instability (CIN) in cancer. eLS:1–9
Horne SD, Pollick SA, Heng HH (2015) Evolutionary mechanism unifies the hallmarks of cancer. Int J Cancer 136(9):2012–2021
Liu G, Stevens JB, Horne SD et al (2014) Genome chaos: survival strategy during crisis. Cell Cycle 13(4):528–537
Manolio TA, Collins FS, Cox NJ et al (2009) Finding the missing heritability of complex diseases. Nature 461(7265):747–753
Heng HH (2011) Missing heritability and stochastic genome alterations. Nat Rev Genet 11(11):813
Stevens JB, Liu G, Abdallah BY et al (2014) Unstable genomes elevate transcriptome dynamics. Int J Cancer 134(9):2074–2087
Stepanenko AA, Kavsan VM (2012) Evolutionary karyotypic theory of cancer versus conventional cancer gene mutation theory. Biopolym Cell 28(4):267–280
Duesberg P, McCormack A (2013) Immortality of cancers: a consequence of inherent karyotypic variations and selections for autonomy. Cell Cycle 12(5):783–802
Poot M, Haaf T (2015) Mechanisms of origin, phenotypic effects and diagnostic implications of complex chromosome rearrangements. Mol Syndromol 6:109–133
Li P, Cui C (2016) A broader view of cancer cytogenetics: from nuclear aberrations to cytogenetic abnormalities. J Mol Genet Med 10:e108
Poot M (2017) Of simple and complex genome rearrangements, chromothripsis, chromoanasynthesis, and chromosome chaos. Mol Syndromol 8:115–117
Heng HH, Regan SM, Liu G et al (2016) Why is it crucial to analyze non clonal chromosomal aberrations or NCCAs? Mol Cytogenet 9:15
Rangel N, Forero-Castro M, Rondón-Lagos M (2017) New insights in the cytogenetic practice: karyotypic chaos, non-clonal chromosomal alterations and chromosomal instability in human cancer and therapy response. Genes (Basel) 8(6):E155
Heng HH, Horne SD, Chaudhry S et al (2018) A postgenomic perspective on molecular cytogenetics. Current Genomics (in press)
Heng HH, Squire J, Tsui LC (1992) High-resolution mapping of mammalian genes by in situ hybridization to free chromatin. Proc Natl Acad Sci U S A 89(20):9509–9513
Heng HH, Tsui LC, Moens PB (1994) Organization of heterologous DNA inserts on the mouse meiotic chromosome core. Chromosoma 103(6):401–407
Ye CJ, Heng HH (2017) High resolution fiber-fluorescence in situ hybridization. Methods Mol Biol 1541:151–166
Ye CJ, Liu G, Heng HH (2016) Simultaneous fluorescence immunostaining and FISH. In: Liehr T (ed) Fluorescence in situ hybridization (FISH). Springer, New York, pp 301–325
Heng HH, Ye CJ, Yang F et al (2003) Analysis of marker or complex chromosomal rearrangements present in pre- and post-natal karyotypes utilizing a combination of G-banding, spectral karyotyping and fluorescence in situ hybridization. Clin Genet 63(5):358–367
Ye CJ, Lu W, Liu G et al (2001) The combination of SKY and specific loci detection with FISH or immunostaining. Cytogenet Cell Genet 93(3–4):195–202
Heng HH (2018) Genome chaos: rethinking genomics, evolution and molecular medicine. Academic Press, Elsevier, (in press)
Gorelick R, Heng HH (2011) Sex reduces genetic variation: a multidisciplinary review. Evolution 65(4):1088–1098
Heng HH, Horne SD, Stevens SB et al (2016) Heterogeneity mediated system complexity: the ultimate challenge for studying common and complex diseases. In: Sturmberg JP (ed) The value of systems and complexity sciences for healthcare. Springer, New York, pp 107–120
Heng HH (2017) Heterogeneity-mediated cellular adaptation and its trade-off: searching for the general principles of diseases. J Eval Clin Pract 23(1):233–237
Pellestor F, Gatinois V, Puechberty J et al (2014) Chromothripsis: potential origin in gametogenesis and preimplantation cell divisions. A review. Fertil Steril 102(6):1785–1796
Furgason JM, Koncar RF, Michelhaugh SK et al (2015) Whole genome sequence analysis links chromothripsis to EGFR, MDM2, MDM4, and CDK4 amplification in glioblastoma. Oncoscience 2(7):618–628. eCollection
de Pagter MS, van Roosmalen MJ, Baas AF et al (2015) Chromothripsis in healthy individuals affects multiple protein-coding genes and can result in severe congenital abnormalities in offspring. Am J Hum Genet 96(4):651–656
Pellestor F (2014) Chromothripsis: how does such a catastrophic event impact human reproduction? Hum Reprod 29(3):388–393
McDermott DH, Gao JL, Liu Q et al (2015) Chromothriptic cure of WHIM syndrome. Cell 160(4):686–699
Acknowledgments
This manuscript is part of our series of publications on the subject of “the mechanisms of cancer and organismal evolution.” This work was also partially supported by the start-up fund for Christine J. Ye from the Department of Internal Medicine, Division of Hematology/Oncology, University of Michigan. Thanks to Julie Heng for editing.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ye, C.J., Liu, G., Heng, H.H. (2018). Experimental Induction of Genome Chaos. In: Pellestor, F. (eds) Chromothripsis. Methods in Molecular Biology, vol 1769. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7780-2_21
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7780-2_21
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7779-6
Online ISBN: 978-1-4939-7780-2
eBook Packages: Springer Protocols