Abstract
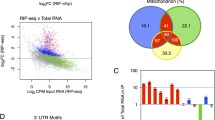
Recent progress in the technology of transcriptome-wide high-throughput sequencing has revealed that nonsense-mediated mRNA decay (NMD) targets ~10% of physiologic transcripts for the purpose of tuning gene expression in response to various environmental conditions. Regardless of the eukaryote studied, NMD requires the ATP-dependent RNA helicase upframeshift 1 (UPF1). It was initially thought that cellular NMD targets could be defined by their binding to steady-state UPF1, which is largely hypophosphorylated. However, the propensity for steady-state UPF1 to bind RNA nonspecifically, coupled with regulated phosphorylation of UPF1 on an NMD target serving as the trigger for NMD, made it clear that it is phosphorylated UPF1 (p-UPF1), rather than steady-state UPF1, that can be used to distinguish cellular NMD targets from cellular RNAs that are not. Here, we describe the immunoprecipitation of p-UPF1 followed by RNA sequencing (p-UPF1 RIP−seq) as a transcriptome-wide approach to define physiologic NMD targets.
Similar content being viewed by others
References
Lewis BP, Green RE, Brenner SE (2003) Evidence for the widespread coupling of alternative splicing and nonsense-mediated mRNA decay in humans. Proc Natl Acad Sci U S A 100:189–192. https://doi.org/10.1073/pnas.0136770100
Pan Q, Saltzman AL, Kim YK et al (2006) Quantitative microarray profiling provides evidence against widespread coupling of alternative splicing with nonsense-mediated mRNA decay to control gene expression. Genes Dev 20:153–158. https://doi.org/10.1101/gad.1382806
He F, Li X, Spatrick P et al (2003) Genome-wide analysis of mRNAs regulated by the nonsense-mediated and 5′ to 3′ mRNA decay pathways in yeast. Mol Cell 12:1439–1452. https://doi.org/10.1016/S1097-2765(03)00446-5
Kurosaki T, Maquat LE (2016) Nonsense-mediated mRNA decay in humans at a glance. J Cell Sci:461–467. https://doi.org/10.1242/jcs.181008
Mendell JT, Sharifi NA, Meyers JL et al (2004) Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat Genet 36:1073–1078. https://doi.org/10.1038/ng1429
Gregersen LH, Schueler M, Munschauer M et al (2014) MOV10 is a 5′ to 3′ RNA helicase contributing to UPF1 mRNA target degradation by translocation along 3′ UTRs. Mol Cell 22:573–585. https://doi.org/10.1016/j.molcel.2014.03.017
Hogg JR, Goff SP (2010) Upf1 senses 3′UTR length to potentiate mRNA decay. Cell 143:379–389. https://doi.org/10.1016/j.cell.2010.10.005
Hurt JA, Robertson AD, Burge CB (2013) Global analyses of UPF1 binding and function reveal expanded scope of nonsense-mediated mRNA decay. Genome Res 23:1636–1650. https://doi.org/10.1101/gr.157354.113
Kurosaki T, Maquat LE (2013) Rules that govern UPF1 binding to mRNA 3′ UTRs. Proc Natl Acad Sci U S A 110:3357–3362. https://doi.org/10.1073/pnas.1219908110
Lee SR, Pratt GA, Martinez FJ et al (2015) Target discrimination in nonsense-mediated mRNA decay requires Upf1 ATPase activity. Mol Cell 59:413–425. https://doi.org/10.1016/j.molcel.2015.06.036
Zünd D, Gruber AR, Zavolan M et al (2013) Translation-dependent displacement of UPF1 from coding sequences causes its enrichment in 3′ UTRs. Nat Struct Mol Biol 20:936–943. https://doi.org/10.1038/nsmb.2635
Isken O, Kim YK, Hosoda N et al (2008) Upf1 phosphorylation triggers translational repression during nonsense-mediated mRNA decay. Cell 133:314–327. https://doi.org/10.1016/j.cell.2008.02.030
Cho H, Kim KM, Kim YK (2009) Human proline-rich nuclear receptor coregulatory protein 2 mediates an interaction between mRNA surveillance machinery and decapping complex. Mol Cell 33:75–86. https://doi.org/10.1016/j.molcel.2008.11.022
Kashima I, Yamashita A, Izumi N et al (2006) Binding of a novel SMG-1-Upf1-eRF1-eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev 20:355–367. https://doi.org/10.1101/gad.1389006
Kurosaki T, Li W, Hoque M et al (2014) A post-translational regulatory switch on UPF1 controls targeted mRNA degradation. Genes Dev 28:1900–1916. https://doi.org/10.1101/gad.245506.114
Loh B, Jonas S, Izaurralde E (2013) The SMG5-SMG7 heterodimer directly recruits the CCR4-NOT deadenylase complex to mRNAs containing nonsense codons via interaction with POP2. Genes Dev 27:2125–2138. https://doi.org/10.1101/gad.226951.113
Okada-Katsuhata Y, Yamashita A, Kutsuzawa K et al (2012) N- and C-terminal Upf1 phosphorylations create binding platforms for SMG-6 and SMG-5:SMG-7 during NMD. Nucleic Acids Res 40:1251–1266. https://doi.org/10.1093/nar/gkr791
Anders KR, Grimson A, Anderson P (2003) SMG-5, required for C. elegans nonsense-mediated mRNA decay, associates with SMG-2 and protein phosphatase 2A. EMBO J 22:641–650. https://doi.org/10.1093/emboj/cdg056
Chiu S, Serin G, Ohara O et al (2003) Characterization of human Smg5/7a: a protein with similarities to Caenorhabditis elegans SMG5 and SMG7 that functions in the dephosphorylation of Upf1. RNA 9:77–87. https://doi.org/10.1261/rna.2137903
Ohnishi T, Yamashita A, Kashima I et al (2003) Phosphorylation of hUPF1 induces formation of mRNA surveillance complexes containing hSMG-5 and hSMG-7. Mol Cell 12:1187–1200. https://doi.org/10.1016/S1097-2765(03)00443-X
Durand S, Franks TM, Lykke-Andersen J (2016) Hyperphosphorylation amplifies UPF1 activity to resolve stalls in nonsense-mediated mRNA decay. Nat Commun 7:12434. https://doi.org/10.1038/nsmb.2575
Yamashita A, Ohnishi T, Kashima I et al (2001) Human SMG-1, a novel phosphatidylinositol 3-kinase-related protein kinase, associates with components of the mRNA surveillance complex and is involved in the regulation of nonsense-mediated mRNA decay. Genes Dev 15:2215–2228. https://doi.org/10.1101/gad.913001
Cohen P, Holmes CFB, Tsukitani Y (1990) Okadaic acid: a new probe for the study of cellular regulation. Trends Biochem Sci 15:98–102. https://doi.org/10.1016/0968-0004(90)90192-E
Nicholson P, Josi C, Kurosawa H et al (2014) A novel phosphorylation-independent interaction between SMG6 and UPF1 is essential for human NMD. Nucleic Acids Res 42:9217–9235. https://doi.org/10.1093/nar/gku645
Jayaprakash AD, Jabado O, Brown BD et al (2011) Identification and remediation of biases in the activity of RNA ligases in small-RNA deep sequencing. Nucleic Acids Res 39:e141. https://doi.org/10.1093/nar/gkr693
Zhuang F, Fuchs RT, Sun Z, Zheng Y, Robb GB (2012) Structural bias in T4 RNA ligase-mediated 3′-adapter ligation. Nucleic Acids Res 40:e54. https://doi.org/10.1093/nar/gkr1263
Mitrovich QM, Anderson P (2000) Unproductively spliced ribosomal protein mRNAs are natural targets of mRNA surveillance in C. elegans. Genes Dev 14:2173–2184. https://doi.org/10.1101/gad.819900
Takei S, Togo-Ohno M, Suzuki Y et al (2016) Evolutionarily conserved autoregulation of alternative pre-mRNA splicing by ribosomal protein L10a. Nucleic Acids Res 44:5585–5596. https://doi.org/10.1093/nar/gkw152
Acknowledgments
We thank Keita Miyoshi for comments on the manuscript. Work on NMD in the Maquat lab is supported by NIH R01 GM059614 to L.E.M. T.K. was partially supported by a postdoctoral fellowship from FRAXA Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Kurosaki, T., Hoque, M., Maquat, L.E. (2018). Identifying Cellular Nonsense-Mediated mRNA Decay (NMD) Targets: Immunoprecipitation of Phosphorylated UPF1 Followed by RNA Sequencing (p-UPF1 RIP−Seq). In: Lamandé, S. (eds) mRNA Decay. Methods in Molecular Biology, vol 1720. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7540-2_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7540-2_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7539-6
Online ISBN: 978-1-4939-7540-2
eBook Packages: Springer Protocols