Abstract
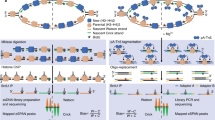
The DNA replication process can be heavily perturbed by several different conditions of genotoxic stress, particularly relevant for cancer onset and therapy. The combination of psoralen crosslinking and electron microscopy has proven instrumental to reveal the fine architecture of in vivo DNA replication intermediates and to uncover their remodeling upon specific conditions of genotoxic stress. The replication structures are stabilized in vivo (by psoralen crosslinking) prior to extraction and enrichment procedures, allowing their visualization at the transmission electron microscope. This chapter outlines the procedures required to visualize and interpret in vivo replication intermediates of eukaryotic genomic DNA, and includes an improved method for enrichment of replication intermediates, compared to previously used BND-cellulose columns.
Similar content being viewed by others
References
Inciarte MR, Salas M, Sogo JM (1980) Structure of replicating DNA molecules of Bacillus subtilis bacteriophage phi 29. J Virol 34:187–199
Lucchini R, Sogo JM (1995) Replication of transcriptionally active chromatin. Nature 374:276–280
Sogo JM, Stahl H, Koller T, Knippers R (1986) Structure of replicating simian virus 40 minichromosomes. The replication fork, core histone segregation and terminal structures. J Mol Biol 189:189–204
Avemann K, Knippers R, Koller T, Sogo JM (1988) Camptothecin, a specific inhibitor of type I DNA topoisomerase, induces DNA breakage at replication forks. Mol Cell Biol 8:3026–3034
Lopes M, Foiani M, Sogo JM (2006) Multiple mechanisms control chromosome integrity after replication fork uncoupling and restart at irreparable UV lesions. Mol Cell 21:15–27
Engels K, Giannattasio M, Muzi-Falconi M et al (2011) 14-3-3 proteins regulate exonuclease 1-dependent processing of stalled replication forks. PLoS Genet 7:e1001367. doi:10.1371/journal.pgen.1001367
Giannattasio M, Follonier C, Tourriere H et al (2010) Exo1 competes with repair synthesis, converts NER intermediates to long ssDNA gaps, and promotes checkpoint activation. Mol Cell 40:50–62. doi:10.1016/j.molcel.2010.09.004
Hashimoto Y, Chaudhuri AR, Lopes M, Costanzo V (2010) Rad51 protects nascent DNA from Mre11-dependent degradation and promotes continuous DNA synthesis. Nat Struct Mol Biol 17:1305–1311. doi:10.1038/nsmb.1927
Sogo J, Lopes M, Foiani M (2002) Fork reversal and ssDNA accumulation at stalled replication forks owing to checkpoint defects. Science (New York, NY) 297:599–602
Berti M, Chaudhuri AR, Thangavel S et al (2013) Human RECQ1 promotes restart of replication forks reversed by DNA topoisomerase I inhibition. Nat Struct Mol Biol 20:347–354. doi:10.1038/nsmb.2501
Follonier C, Oehler J, Herrador R, Lopes M (2013) Friedreich’s ataxia-associated GAA repeats induce replication-fork reversal and unusual molecular junctions. Nat Struct Mol Biol. doi:10.1038/nsmb.2520
Neelsen KJ, Zanini IMY, Mijic S et al (2013) Deregulated origin licensing leads to chromosomal breaks by rereplication of a gapped DNA template. Genes Dev 27:2537–2542. doi:10.1101/gad.226373.113
Neelsen KJ, Zanini IMY, Herrador R, Lopes M (2013) Oncogenes induce genotoxic stress by mitotic processing of unusual replication intermediates. J Cell Biol 200:699–708. doi:10.1128/MCB.24.16.7140-7150.2004
Neelsen KJ, Lopes M (2015) Replication fork reversal in eukaryotes: from dead end to dynamic response. Nat Rev Mol Cell Biol 16:207–220. doi:10.1038/nrm3935
Ray Chaudhuri A, Hashimoto Y, Herrador R et al (2012) Topoisomerase I poisoning results in PARP-mediated replication fork reversal. Nat Struct Mol Biol 19:417–423. doi:10.1038/nsmb.2258
Ray Chaudhuri A, Ahuja AK, Herrador R, Lopes M (2015) Poly(ADP-ribosyl) glycohydrolase prevents the accumulation of unusual replication structures during unperturbed S phase. Mol Cell Biol 35:856–865. doi:10.1128/MCB.01077-14
Zellweger R, Dalcher D, Mutreja K et al (2015) Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J Cell Biol 208:563–579. doi:10.1083/jcb.201406099
Thangavel S, Berti M, Levikova M et al (2015) DNA2 drives processing and restart of reversed replication forks in human cells. J Cell Biol 208:545–562. doi:10.1083/jcb.201406100
Fugger K, Mistrik M, Neelsen KJ et al (2015) FBH1 catalyzes regression of stalled replication forks. Cell Rep. doi:10.1016/j.celrep.2015.02.028
Lopes M (2009) Electron microscopy methods for studying in vivo DNA replication intermediates. Methods Mol Biol 521:605–631
Neelsen KJ, Chaudhuri AR, Follonier C et al (2014) Visualization and interpretation of eukaryotic DNA replication intermediates in vivo by electron microscopy. Methods Mol Biol 1094:177–208. doi:10.1007/978-1-62703-706-8_15
Gasser R, Koller T, Sogo JM (1996) The stability of nucleosomes at the replication fork. J Mol Biol 258:224–239
Gruss C, Wu J, Koller T, Sogo JM (1993) Disruption of the nucleosomes at the replication fork. EMBO J 12:4533–4545
Mejlvang J, Feng Y, Alabert C et al (2014) New histone supply regulates replication fork speed and PCNA unloading. J Cell Biol 204:29–43. doi:10.1083/jcb.201305017
Wellinger RE, Lucchini R, Dammann R, Sogo JM (1999) In vivo mapping of nucleosomes using psoralen-DNA crosslinking and primer extension. Methods Mol Biol 119:161–173
Vollenweider HJ, Sogo JM, Koller T (1975) A routine method for protein-free spreading of double- and single-stranded nucleic acid molecules. Proc Natl Acad Sci U S A 72:83–87
Sogo JM, Thoma F (1989) Electron microscopy of chromatin. Methods Enzymol 170:142–165
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning, a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press., Cold Spring Harbor, NY
Trenz K, Errico A, Costanzo V (2008) Plx1 is required for chromosomal DNA replication under stressful conditions. EMBO J 27:876–885. doi:10.1038/emboj.2008.29
Ahuja AK, Jodkowska K, Teloni F et al (2016) A short G1 phase imposes constitutive replication stress and fork remodelling in mouse embryonic stem cells. Nat Commun 7:10660. doi:10.1038/ncomms10660
Sogo JM (2002) Fork reversal and ssDNA accumulation at stalled replication forks owing to checkpoint defects. Science 297:599–602. doi:10.1126/science.1074023
Mojas N, Lopes M, Jiricny J (2007) Mismatch repair-dependent processing of methylation damage gives rise to persistent single-stranded gaps in newly replicated DNA. Genes Dev 21:3342–3355
Acknowledgments
We are grateful to José M. Sogo for his crucial support while learning all technicalities of this approach. We wish to thank the whole team at the ZMB (Center for Microscopy and Image Analysis of the University Zurich) for consistently excellent technical assistance, while running our EM experiments. We are grateful to Arnab Ray Chaudhuri, Yoshitami Hashimoto, Fabio Puddu, and Vincenzo Costanzo for their assistance in optimizing this EM approach on Xenopus egg extracts. We also wish to thank Petr Cejka for suggesting the use of QIAGEN columns as possible valuable alternative to BND cellulose for the enrichment of a subpopulation of DNA molecules based on differential ssDNA content. We are also grateful to Sebastian Ursich for his recent efforts optimizing this approach and for careful reading of the manuscript. Work in the Lopes lab is currently financed by the SNF grant 31003A_169959, the ERC Consolidator grant 617102 (ReStreCa) and the Swiss Cancer League grant KFS-3967-08-2016.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Zellweger, R., Lopes, M. (2018). Dynamic Architecture of Eukaryotic DNA Replication Forks In Vivo, Visualized by Electron Microscopy. In: Muzi-Falconi, M., Brown, G. (eds) Genome Instability. Methods in Molecular Biology, vol 1672. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7306-4_19
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7306-4_19
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7305-7
Online ISBN: 978-1-4939-7306-4
eBook Packages: Springer Protocols