Abstract
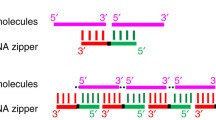
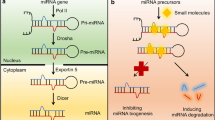
The discovery of microRNAs (miRNAs) has opened an entire new avenue for drug development. These short (15–22 nucleotides) noncoding RNAs, which function in RNA silencing and posttranscriptional regulation of gene expression, have been shown to critically affect numerous pathways in both development and disease progression. Current miRNA drug development focuses on either reintroducing the miRNA into cells through the use of a miRNA mimic or inhibiting its function via use of a synthetic antagomir. Although these methods have shown some success as therapeutics, they face challenges particularly with regard to cellular uptake and for use as systemic reagents. We recently presented a novel mechanism of inhibiting miR-544 by directed inhibition of miRNA biogenesis. We found that inhibition of DICER processing of miR-544 through the use of a small molecule abolished miR-544 function in regulating adaptation of breast cancer cells to hypoxic stress. Herein, we describe a protocol that utilizes bioinformatics to first identify lead small molecules that bind to DICER cleavage sites in pre-miRNAs and then employ an efficient, high-throughput fluorescent-based screening system to determine the inhibitory potential of the lead compounds and their derivatives.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854
Wang J, Lu Z, Wientjes MG, Au JLS (2010) Delivery of siRNA therapeutics: barriers and carriers. AAPS J 12(4):492–503
Burnett John C, Rossi John J (2012) RNA-based therapeutics: current progress and future prospects. Chem Biol 19(1):60–71
Cullen BR (2004) Transcription and processing of human microRNA precursors. Mol Cell 16(6):861–865
Velagapudi SP, Gallo SM, Disney MD (2014) Sequence-based design of bioactive small molecules that target precursor microRNAs. Nat Chem Biol 10(4):291–297
Haga CL et al (2015) Small molecule inhibition of miR-544 biogenesis disrupts adaptive responses to hypoxia by modulating ATM-mTOR signaling. ACS Chem Biol 10(10):2267–2276
Haga CL, Phinney DG (2012) MicroRNAs in the imprinted DLK1-DIO3 region repress the epithelial-to-mesenchymal transition by targeting the TWIST1 protein signaling network. J Biol Chem 287(51):42695–42707
Eddy SR (2004) How do RNA folding algorithms work? Nat Biotechnol 22(11):1457–1458
Lorenz R et al (2011) ViennaRNA Package 2.0. Algorithms Mol Biol 6:26
Mathews DH (2014) RNA secondary structure analysis using RNAstructure. Curr Protoc Bioinformatics Chapter 12:Unit 12.6
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31(13):3406–3415
Seetin MG, Mathews DH (2012) RNA structure prediction: an overview of methods. Methods Mol Biol 905:99–122
Bernhart SH (2011) RNA structure prediction. Methods Mol Biol 760:307–323
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36(Database issue):D154–D158
Milligan JF, Uhlenbeck OC (1989) Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol 180:51–62
Chaires JB (2003) A competition dialysis assay for the study of structure-selective ligand binding to nucleic acids. Curr Protoc Nucleic Acid Chem Chapter 8:Unit 8.3
Tse WC, Boger DL (2005) A fluorescent intercalator displacement assay for establishing DNA binding selectivity and affinity. Curr Protoc Nucleic Acid Chem 8:85
Puglisi JD, Tinoco I Jr (1989) Absorbance melting curves of RNA. Methods Enzymol 180:304–325
Acknowledgement
This work was supported by Department of Defense CDMRP Grant W81XWH-16-1-0029 to D.G.P. and M.D.D.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Haga, C.L., Velagapudi, S.P., Childs-Disney, J.L., Strivelli, J., Disney, M.D., Phinney, D.G. (2017). Rapid Generation of miRNA Inhibitor Leads by Bioinformatics and Efficient High-Throughput Screening Methods. In: Schmidt, M. (eds) Drug Target miRNA. Methods in Molecular Biology, vol 1517. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6563-2_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6563-2_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6561-8
Online ISBN: 978-1-4939-6563-2
eBook Packages: Springer Protocols